†These authors contributed equally.
Academic Editor: Marcello Iriti
Introduction: Chronic obstructive pulmonary disease (COPD) is an inflammatory disease caused by increasing breathing passage obstruction which completely disrupts human homeostasis. Some patients require lung transplantation or long-term oxygen therapy. COPD is one of the noxious diseases and its fourth leading cause of death around the globe. There is an immediate need for potential drug development to tackle this serious disease. Folk medicines are used to combat complex diseases that have shown effectiveness in the treatment of breathing diseases. Vitex negundo L. is an ethnobotanically important medicinal plant used for various ailments and modulates human cellular events. This shrub has diverse specialized metabolites and is being used as complementary medicine in various countries. Though systems-level understanding is there on the mode of action, the multi-target treatment strategy for COPD is still a bottleneck. Methods: In this investigation, systems pharmacology, cheminformatics, and molecular docking analyses were performed to unravel the multi-targeted mechanisms of V. negundo L. potential bioactives to combat COPD. Results: Cheminformatics analysis combined with the target mining process identified 86 specialized metabolites and their corresponding 1300 direct human receptors, which were further imputed and validated systematically. Furthermore, molecular docking approaches were employed to evaluate the potential activity of identified potential compounds. In addition, pharmacological features of these bioactives were compared with available COPD drugs to recognize potential compounds that were found to be more efficacious with higher bioactive scores. Conclusions: The present study unravels the druggable targets and identifies the bioactive compounds present in V. negundo L., that may be utilized for potential treatment against COPD. However, further in vivo analyses and clinical trials of these molecules are essential to deciphering their efficacy.
Chronic Obstructive Pulmonary Disease (COPD) is one of the complex progressive disorders that affect the respiratory system [1] and causes chronic inflammation of the airways [2]. COPD is accountable for more than 3 million deaths annually, making it the third leading cause of mortality [3]. There are several causes for COPD, which provoke the Endoplasmic Reticulum (ER) stress including smoking, free radicals, carcinogens, and reactive oxygen species [4]. Since ER strives to reimpose the cellular homeostasis through unfolded protein response (UPR), it is evident that ER stress and UPR activation occur in patients with COPD [5]. The other major cause for COPD is breathing of polluted air and also the result of subjection to inhaled irritants [1, 6]. COPD is frequently associated with non-small cell lung cancer (comorbidity) [7]. Factors such as age, bacterial colonization, cardiovascular diseases, usage of antibiotics and steroids, poor quality of life, and airway obstruction severity intensify the risk for COPD [8, 9]. One of the uncharacterized risk factors of COPD is the longtime occupational exposure to irritants like organic and inorganic dust, chemical agents, and fumes [1]. These risk factors are being changed over time with new forms of smoking and also the location with the difference in air pollution [10]. COPD is commonly accompanied by exacerbations, which are marked with increased cough, excess sputum production, and phlegm [11]. It is deemed that among the exacerbations, 70–80% are prompted by bacterial or viral respiratory infections and the remaining 20–30% are due to environmental pollution exposure or unknown causation or origination [12]. In general, COPD is an escalating disease of the airways, the microvasculature, and the alveoli and is strongly associated with cardiovascular diseases [13]. Although COPD represents an increased burden to the Healthcare system, it is not easy to diagnose and curate the disease due to its heterogenicity and complexity. Currently, this disease is treated by both Pharmacological and non-pharmacological approaches. The pharmacological intervention includes inhaled bronchodilators, antibiotics, corticosteroids, Oxygen therapy, and antioxidants. Based on the clinical conditions of the patient, adjunct therapies are also performed. Non-pharmacological intervention includes Chest percussion therapy, which is believed to improve sputum clearance. Ventilatory support should be the primary goal to reduce morbidity and mortality [12]. Currently, the prognosis of COPD is achieved by post-bronchodilator spirometry and by Pulse oximetry [1].
The Indian medical system is one of the ancient traditional health care systems in the globe and this system is predominantly based on herbal plants. Thus, repurposing Indian traditional practices can be employed in mining new options to treat deadly diseases. Among numerous plants mentioned in the Indian medical system, Vitex negundo L. commonly known as Nirgundi belongs to the family Lamiaceae [14] is one of the significant plants that was being used in the treatment for various disorders.
The plant contains various bioactive compounds extracted and concentrated from roots, leaves, and seeds in the form of iridoids, terpenes, volatile oils, lignans, steroids, and flavonoids. These different bioactive compounds exhibit various pharmacological properties including anti-inflammatory, antimicrobial, antioxidant, anti-ulcer, anti-diabetic properties, hepatoprotective properties [15]. The leaves have been used as a sedative, vermifuge, astringent, tonic, febrifuge, and also used in imparting the joint swellings from Acute Rheumatism. The dried fruit can be used as a Quinacrine and is used in the treatment of cough, cold, heart diseases, coronary thrombosis, rheumatic difficulties, etc. [16]. Jianpiyifei II granules (JPYF II) is one of the herbal medications used for COPD in China which comprises V. negundo L. is an important component [5, 15]. Since we have a limited number of drugs and disease-modifying therapies to treat COPD, the unraveling of the genetic determinants of COPD provides the unbiased identification of molecular determinants which would pave the way to derive new insights into the pathogenesis of COPD leading to novel therapeutic interventions and preventive strategies.
Despite the significance of the Indian traditional treatment system, the functions of phytocompounds, their potential human targets, and their mode of actions are still indistinct. Several studies have shown the specific activity of these compounds, but the mode of action and multiple potentialities of these compounds are yet to be fully discovered. Hence, the present study aims to highlight the advances in the discovery of druggable targets of phytomolecules in association with COPD through systems pharmacology, cheminformatics, and molecular docking approaches. This study tries to address the primary queries regarding:
(i) The regulatory aspects of phytocompounds against COPD.
(ii) The biological processes mediated by plant molecules through human targets.
Cheminformatics approach was employed to segregate the phytochemicals with significant curative properties through target-compound interactions and molecular docking was performed to evaluate the interactions. The derived COPD immunological receptors were analyzed using bioinformatic databases to identify the pathways and mechanisms in which the bioactive compounds act. So, it is expected that these in silico systems approaches can aid in the investigation of the immunological significance of V. negundo L. derived specialized molecules and will significantly promote the development of new drugs for the treatment of COPD and other respiratory diseases in mere future.
Scrutinization of literature and web sources [17, 18] revealed the presence of various bioactive compounds in V. negundo L. and were enlisted in Table 1. In total, 86 phytocompounds were procured and their Canonical SMILES were retrieved from the PubChem database [19].
| S.No. | Compounds | PubChem ID | Abb |
| 1 | Negundoside | 9935561 | NS |
| 2 | Agnuside | 442416 | AS |
| 3 | Thujene | 520384 | TJ |
| 4 | 6654 | ||
| 5 | Camphene | 6616 | CP |
| 6 | 80048 | ||
| 7 | δ- Elemene | 12309449 | δ-E |
| 8 | Sabinene | 18818 | SN |
| 9 | Friedelin | 91472 | FD |
| 10 | Vitamin-C | 54670067 | VC |
| 11 | Carotene | 6419725 | CT |
| 12 | Casticin | 5315263 | CC |
| 13 | Artemetin | 5320351 | ART |
| 14 | Terpinen-4-ol | 11230 | T-4-ol |
| 15 | Spathulenol | 92231 | STL |
| 16 | Caryophyllene epoxide | 14350 | CPE |
| 17 | Caryophyllenol | 61125 | CPL |
| 18 | Farnesol | 445070 | FN |
| 19 | 14896 | ||
| 20 | Stearic acid | 5281 | STA |
| 21 | Behenic acid | 8215 | BHA |
| 22 | Myrcene | 31253 | MC |
| 23 | ∆ |
26049 | ∆ |
| 24 | Limonene | 22311 | LMN |
| 25 | 11142 | ||
| 26 | 7461 | ||
| 27 | Dihydromyrcenol | 29096 | DHC |
| 28 | Sabinene hydrate | 62367 | SBH |
| 29 | Linalool | 6549 | LNL |
| 30 | Amyl isovalerate | 95978 | AIV |
| 31 | Nonanol | 8914 | NNL |
| 32 | 4-Terpineol | 11230 | 4-TP |
| 33 | 17100 | ||
| 34 | Carveol | 7438 | CV |
| 35 | Eugenol | 3314 | EUG |
| 36 | 5281515 | ||
| 37 | Isocaryophyllene | 5281522 | ICP |
| 38 | Humulene | 5281520 | HML |
| 39 | Aromadendrene | 91354 | ADD |
| 40 | Viridiflorene | 10910653 | VDFE |
| 41 | δ-Cadinene | 441005 | δ-C |
| 42 | 4,4ʹʹ-Dimethoxy-trans-stilbene | 20828 | 4,4ʹʹ- DM-T-SB |
| 43 | Elemol | 92138 | EL |
| 44 | Caryophyllene oxide | 1742210 | CPO |
| 45 | Vitexicarpin | 5315263 | VTC |
| 46 | Terpinen-4-ol | 11230 | TP-4-ol |
| 47 | Viridiflorol | 11996452 | VDF |
| 48 | 442355 | ||
| 49 | Camphor | 2537 | CMR |
| 50 | 1,8-Cineol | 2758 | 1,8- C |
| 51 | 5317844 | ||
| 52 | Neral | 643779 | NL |
| 53 | Geranial | 638011 | GRN |
| 54 | Bornyl acetate | 6448 | BA |
| 55 | Nerolidol | 5284507 | NLD |
| 56 | 6918391 | ||
| 57 | n-Tritriacontane | 12411 | n-T |
| 58 | n-Hentriacontanol | 68345 | n-H |
| 59 | Epifriedelinol | 119242 | EFDL |
| 60 | Oleanolic acid | 10494 | OLNA |
| 61 | n-Nonacosane | 12409 | n-N |
| 62 | Vitedoin A | 21574226 | VDA |
| 63 | Vitedoamine A | 11348702 | VAA |
| 64 | Negundin A | 10043572 | NA |
| 65 | Negundin B | 10473569 | NB |
| 66 | Vitedoin B | 11771639 | VDB |
| 67 | 222284 | ||
| 68 | 10856614 | ||
| 69 | Germacren-4-ol | 6429375 | GMC-4-ol |
| 70 | 91457 | ||
| 71 | Acetyl oleanolic acid | 151202 | AOA |
| 72 | Sitosterol | 222284 | SS |
| 73 | (E)-Nerolidol | 5281525 | E- N |
| 74 | 28237 | ||
| 75 | 608041 | ||
| 76 | Germacrene D | 5317570 | GD |
| 77 | Hexadecanoic acid | 985 | HDA |
| 78 | p-Cymene | 7463 | P-C |
| 79 | Valencene | 9855795 | VLC |
| 80 | 12300146 | ||
| 81 | Cedrol | 65575 | CD |
| 82 | 6432005 | ||
| 83 | Squalene | 638072 | SQL |
| 84 | Vitexin | 5280441 | VTN |
| 85 | Aucubin | 91458 | AUC |
| 86 | Isovitexin | 162350 | IVT |
Canonical SMILES of these compounds were employed to identify the human targets by using the SwissTargetPrediction tool (http://www.swisstargetprediction.ch/). Thus, identified human targets were submitted to the Expression Atlas database (https://www.ebi.ac.uk/gxa/home), and their features such as UniProt ID, orthologs, and Chromosome number were collated.
Gene target identifiers were subjected to Network Analyst (https://www.networkanalyst.ca/) [20] to acquire the information about Gene Ontology (GO) classified as molecular functions and biological processes against Homo sapiens. In addition, Gene enrichment networks were also procured through the Network Analyst plugin.
Cytoscape v3.8.2 [21] was used for constructing Compound-Target-Network, which plays an important role in identifying and understanding the mechanism of compound activity. Additionally, the interaction between the target genes was visualized using gene mania (https://genemania.org/) [22]. Based on these studies, potential targets were identified.
The protein-protein interaction (PPI) of these potential proteins was performed using STRING v10.5 (https://string-db.org/) with a high confidence score of 0.7. This interactome was utilized to understand the regulatory aspects of potential targets.
Canonical SMILES of the corresponding phytocompounds were subjected to the Molinspiration tool (https://www.molinspiration.com/) to predict the properties such as the number of violations (nVio), GPCR ligand activity (GPCR), Enzymes and nuclear receptors (Ncr), Kinase inhibitory activity (Ki), Enzyme inhibitory activity (Ei) and Protease inhibitory activity (Pi).
Molecular Docking (MD) was performed for the pharmacologically active compounds against the COPD responsible human targets to evaluate the potentials of pharmacologically active compounds.
Commercial drugs available for COPD were retrieved from various sources and their bioactive properties including nVio, GPCR, Ncr, Ki, Ei, and Pi were compared with plant-derived molecules to identify the pharmacologically active compounds.
The canonical SMILES of all the 86 phytocompounds were procured from the PubChem database, which is further employed for systems pharmacological analysis.
Results of SwissTargetPrediction reveal the Human receptors, targeted by phytocompounds. On whole, 86 phytocompounds targeting 1300 targets were identified in this study. A list of compounds along with their target receptors was enlisted in Supplementary Table 1. In addition, the UniProt ID, chromosome number, and orthologous information were also retrieved and tabulated (Table 2). This information can be employed for deeper analyses of molecular functions.
| S.No. | Compound | Target | UniProt ID | Chr. No. | Orthologs |
| 1 | Negundoside | HSP90AA1 | H0YJF5 | 14 | HSP90AA1 (Equus caballus) |
| 2 | Agnuside | HSP90AA1 | H0YJF5 | 14 | HSP90AA1 (Equus caballus) |
| 3 | Thujene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 4 | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) | |
| 5 | Camphene | HSD11B1 | P28845 | 1 | Hsd11b1 (Mus musculus) |
| 6 | ADORA1 | A0A087X173 | 1 | Adora1 (Rattus norvegicus) | |
| 7 | δ-Elemene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 8 | Sabinene | HSD11B1 | P28845 | 1 | Hsd11b1 (Mus musculus) |
| 9 | Friedelin | CYP19A1 | H0YLS2 | 15 | Cyp19a1 (Rattus norvegicus) |
| 10 | Vitamin-C | GSK3B | A0A3B3ITW1 | 3 | GSK3B (Sus scrofa) |
| 11 | Carotene | ADORA1 | A0A087X173 | 1 | Adora1 (Rattus norvegicus) |
| 12 | Casticin | AKR1B1 | P15121 | 7 | AAD14 (Saccharomyces cerevisiae) |
| 13 | Artemetin | ABCG2 | Q9UNQ0 | 4 | SNQ2 (Saccharomyces cerevisiae) |
| 14 | Terpinen-4-ol | AR | E9PEG3 | X | AR (Papio anubis) |
| 15 | Spathulenol | UGT2B7 | A0A087X084 | 4 | Ugt49C1 (Drosophila melanogaster) |
| 16 | Caryophyllene epoxide | SQLE | Q14534 | 8 | Sqle (Rattus rattus) |
| 17 | Caryophyllenol | UGT2B7 | A0A087X084 | 4 | Ugt49C1 (Drosophila melanogaster) |
| 18 | Farnesol | SQLE | Q14534 | 8 | Sqle (Rattus rattus) |
| 19 | SLC5A7 | Q9GZV3 | 2 | Slc5a7 (Mus musculus) | |
| 20 | Stearic acid | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 21 | Behenic acid | FABP4 | P15090 | 8 | Fabp4 (Mus musculus) |
| 22 | Myrcene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 23 | ∆ |
PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 24 | Limonene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 25 | AR | E9PEG3 | X | AR (Papio anubis) | |
| 26 | TRPV1 | I3L1R6 | 17 | TRPV1 (Gallus gallus) | |
| 27 | Dihydromyrcenol | HSD11B1 | P28845 | 1 | Hsd11b1 (Mus musculus) |
| 28 | Sabinene hydrate | TRPM8 | Q7Z2W7 | 2 | Trpm8 (Mus musculus) |
| 29 | Linalool | TRPM8 | Q7Z2W7 | 2 | Trpm8 (Mus musculus) |
| 30 | Amyl isovalerate | CA1 | E5RHP7 | 8 | CA1 (Pan troglodytes) |
| 31 | Nonanol | TRPM8 | Q7Z2W7 | 2 | Trpm8 (Mus musculus) |
| 32 | 4-Terpineol | AR | E9PEG3 | X | AR (Papio anubis) |
| 33 | AR | E9PEG3 | X | AR (Papio anubis) | |
| 34 | Carveol | AR | E9PEG3 | X | AR (Papio anubis) |
| 35 | Eugenol | ADORA1 | A0A087X173 | 1 | Adora1 (Rattus norvegicus) |
| 36 | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) | |
| 37 | Isocaryophyllene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 38 | Humulene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 39 | Aromadendrene | HSD11B1 | P28845 | 1 | Hsd11b1 (Mus musculus) |
| 40 | Viridiflorene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 41 | δ-Cadinene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) |
| 42 | 4,4ʹʹ-Dimethoxy-trans-stilbene | RELA | E9PSE4 | 11 | RELA (Pan troglodytes) |
| 43 | Elemol | UGT2B7 | A0A087X084 | 4 | Ugt49C1 (Drosophila melanogaster) |
| 44 | Caryophyllene oxide | SQLE | Q14534 | 8 | Sqle(Rattus rattus) |
| 45 | Vitexicarpin | AKR1B1 | P15121 | 7 | AAD14 (Saccharomyces cerevisiae) |
| 46 | Terpinen-4-ol | AR | E9PEG3 | X | AR (Papio anubis) |
| 47 | Viridiflorol | UGT2B7 | A0A087X084 | 4 | Ugt49C1 (Drosophila melanogaster) |
| 48 | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) | |
| 49 | Camphor | NR1I3 | Q6GZ72 | 1 | NR1I3 (Sus scrofa) |
| 50 | 1,8-Cineol | CYP19A1 | H0YLS2 | 15 | Cyp19a1 (Rattus norvegicus) |
| 51 | PPARA | B0QYX1 | 22 | Ppara (Mus musculus) | |
| 52 | Neral | ALDH1A1 | P00352 | 9 | Aldh1a1 (Mus musculus) |
| 53 | Geranial | ALDH1A1 | P00352 | 9 | Aldh1a1 (Mus musculus) |
| 54 | Bornyl acetate | ACHE | F8WD34 | 7 | ACHE (Chlorocebus sabaeus) |
| 55 | Nerolidol | SQLE | Q14534 | 8 | Sqle(Rattus rattus) |
| 56 | CXCR3 | P49682 | X | CXCR3 (Bos taurus) | |
| 57 | n-Tritriacontane | SHBG | B0FWH6 | 17 | SHBG (Ovis aries) |
| 58 | n-Hentriacontanol | TRPM8 | Q7Z2W7 | 2 | Trpm8 (Mus musculus) |
| 59 | Epifriedelinol | TRPM8 | Q7Z2W7 | 2 | Trpm8 (Mus musculus) |
| 60 | Oleanolic acid | PTPN1 | B4DSN5 | 20 | Ptpn1 (Mus musculus) |
| 61 | n-Nonacosane | SHBG | B0FWH6 | 17 | SHBG (Ovis aries) |
| 62 | Vitedoin A | SHBG | B0FWH6 | 17 | SHBG (Ovis aries) |
| 63 | Vitedoamine A | NQO2 | Q5TD07 | 6 | NQO2 (Pan troglodytes) |
| 64 | Negundin A | CYP19A1 | H0YLS2 | 15 | Cyp19a1 (Rattus norvegicus) |
| 65 | Negundin B | FLT3 | E7ER61 | 13 | FLT3 (Equus caballus) |
| 66 | Vitedoin B | PTPN1 | B4DSN5 | 20 | Ptpn1 (Mus musculus) |
| 67 | AR | E9PEG3 | X | AR (Papio anubis) | |
| 68 | CYP19A1 | H0YLS2 | 15 | Cyp19a1 (Rattus norvegicus) | |
| 69 | Germacren-4-ol | UGT2B7 | A0A087X084 | 4 | Ugt49C1 (Drosophila melanogaster) |
| 70 | UGT2B7 | A0A087X084 | 4 | Ugt49C1 (Drosophila melanogaster) | |
| 71 | Acetyl oleanolic acid | PTPN1 | B4DSN5 | 20 | Ptpn1 (Mus musculus) |
| 72 | Sitosterol | AR | E9PEG3 | X | AR (Papio anubis) |
| 73 | (E)-Nerolidol | SQLE | Q14534 | 8 | Sqle(Rattus rattus) |
| 74 | CYP19A1 | H0YLS2 | 15 | Cyp19a1 (Rattus norvegicus) | |
| 75 | CYP19A1 | H0YLS2 | 15 | Cyp19a1 (Rattus norvegicus) | |
| 76 | Germacrene D | PPARA | B0QYX1 | 22 | Ppara (Mus musculus)3 |
| 77 | Hexadecanoic acid | PPARA | B0QYX1 | 22 | Ppara (Mus musculus)3 |
| 78 | p-Cymene | CYP2A6 | M0R2Z4 | 19 | Cyp2a12 (Mus musculus) |
| 79 | Valencene | PPARA | B0QYX1 | 22 | Ppara (Mus musculus)3 |
| 80 | AR | E9PEG3 | X | AR (Papio anubis) | |
| 81 | Cedrol | UGT2B7 | A0A087X084 | 4 | Ugt49C1 (Drosophila melanogaster) |
| 82 | CYP19A1 | H0YLS2 | 15 | Cyp19a1 (Rattus norvegicus) | |
| 83 | Squalene | CNR2 | P34972 | 1 | CNR2 (Macaca mulatta) |
| 84 | Vitexin | AKR1B1 | P15121 | 7 | AAD14 (Saccharomyces cerevisiae) |
| 85 | Aucubin | CDA | P32320 | 1 | CDA (Oryctolagus cuniculus) |
| 86 | Isovitexin | AKR1B1 | P15121 | 7 | AAD14 (Saccharomyces cerevisiae) |
Gene Ontology analysis using Network Analyst revealed the involvement of significant proteins in various biological processes including Response to hypoxia, Hemopoiesis, Cell proliferation, Wound and Inflammatory responses (Fig. 1), and molecular functions such as Enzyme binding, Neurotransmitter receptor activity, Growth Factor activity and cytokine receptor activity (Fig. 2). In addition, gene enrichment networking predicts the activity of these genes involved in various disorders like Inflammatory bowel disease (IBD), Leishmaniasis, Malaria, Amoebiasis, and also in pathways which include MAPK signaling pathway, HIF-1 signaling pathway, FoxO signaling pathway, and TGF-beta signaling pathway (Fig. 3). The activity of these phytocompounds on its targets may reduce the risk of COPD and its associated disorders.
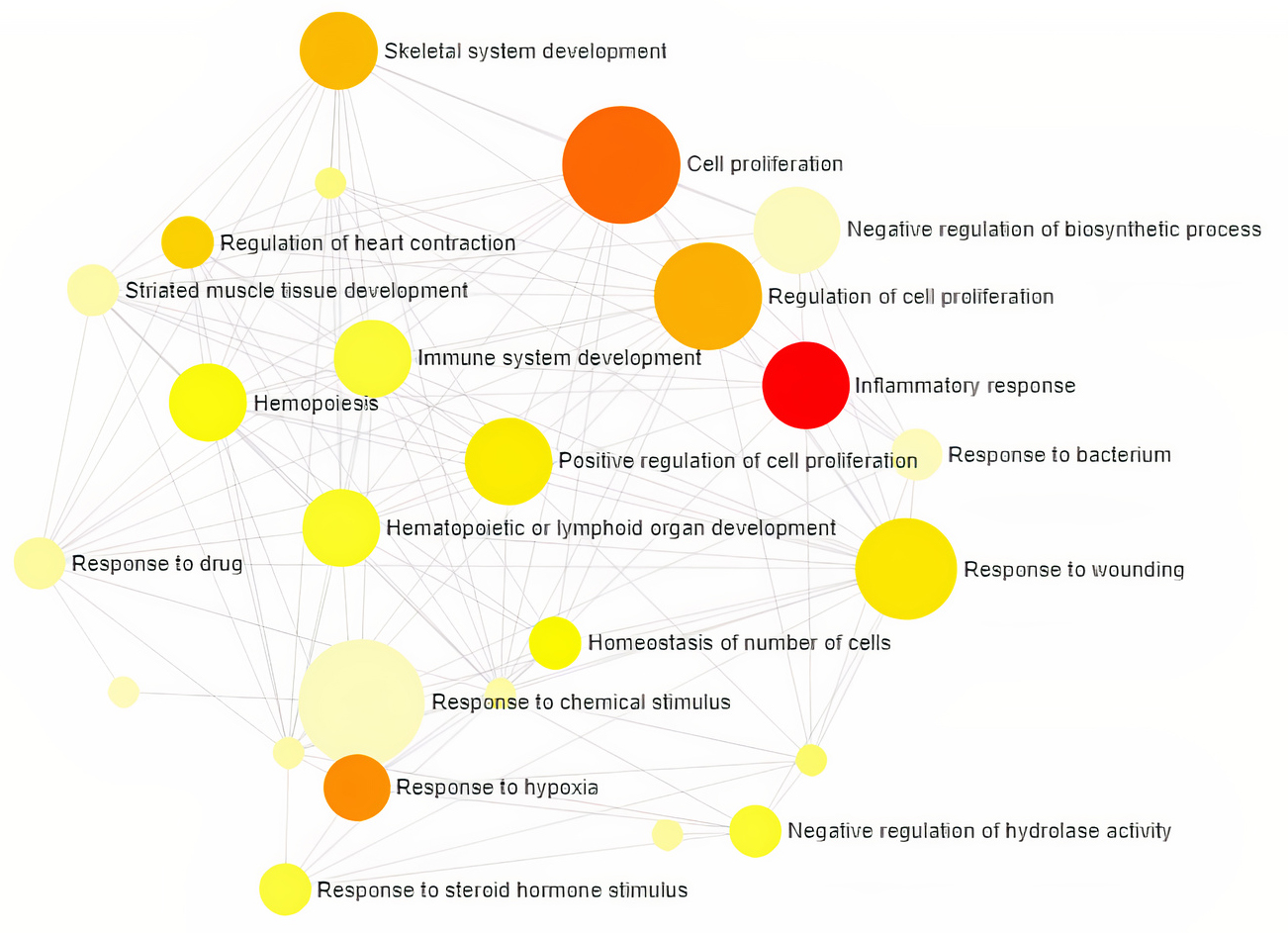 Fig. 1.
Fig. 1.
Target genes involved in Biological Processes and the category
are directly proportional to the node size. The nodes are color shaded
according to the significance level (adjusted p-value
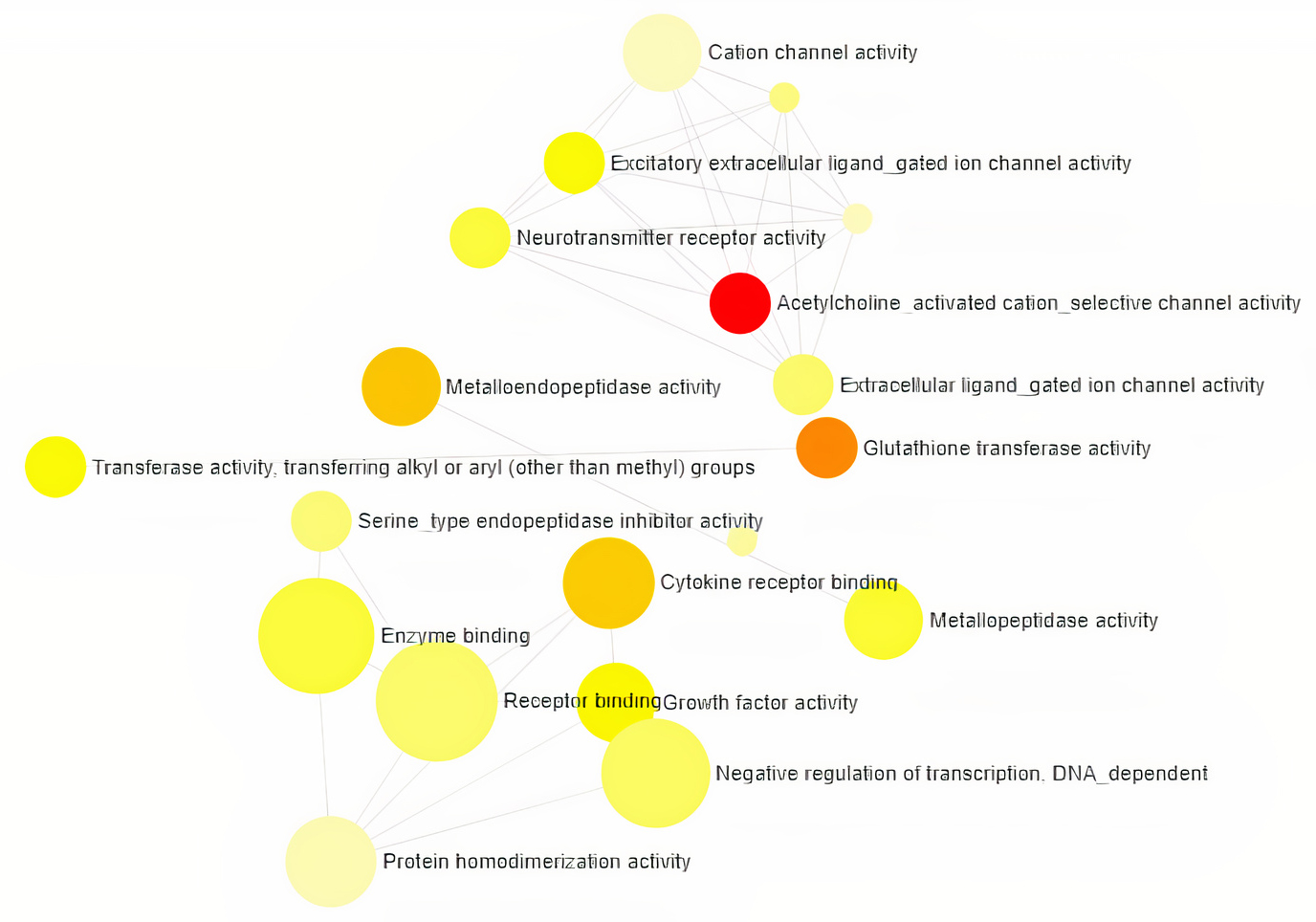 Fig. 2.
Fig. 2.Classification of Human Targets with encoding Molecular
Functions and the category is directly proportional to the node size. The
nodes are color shaded according to the significance level (adjusted
p-value
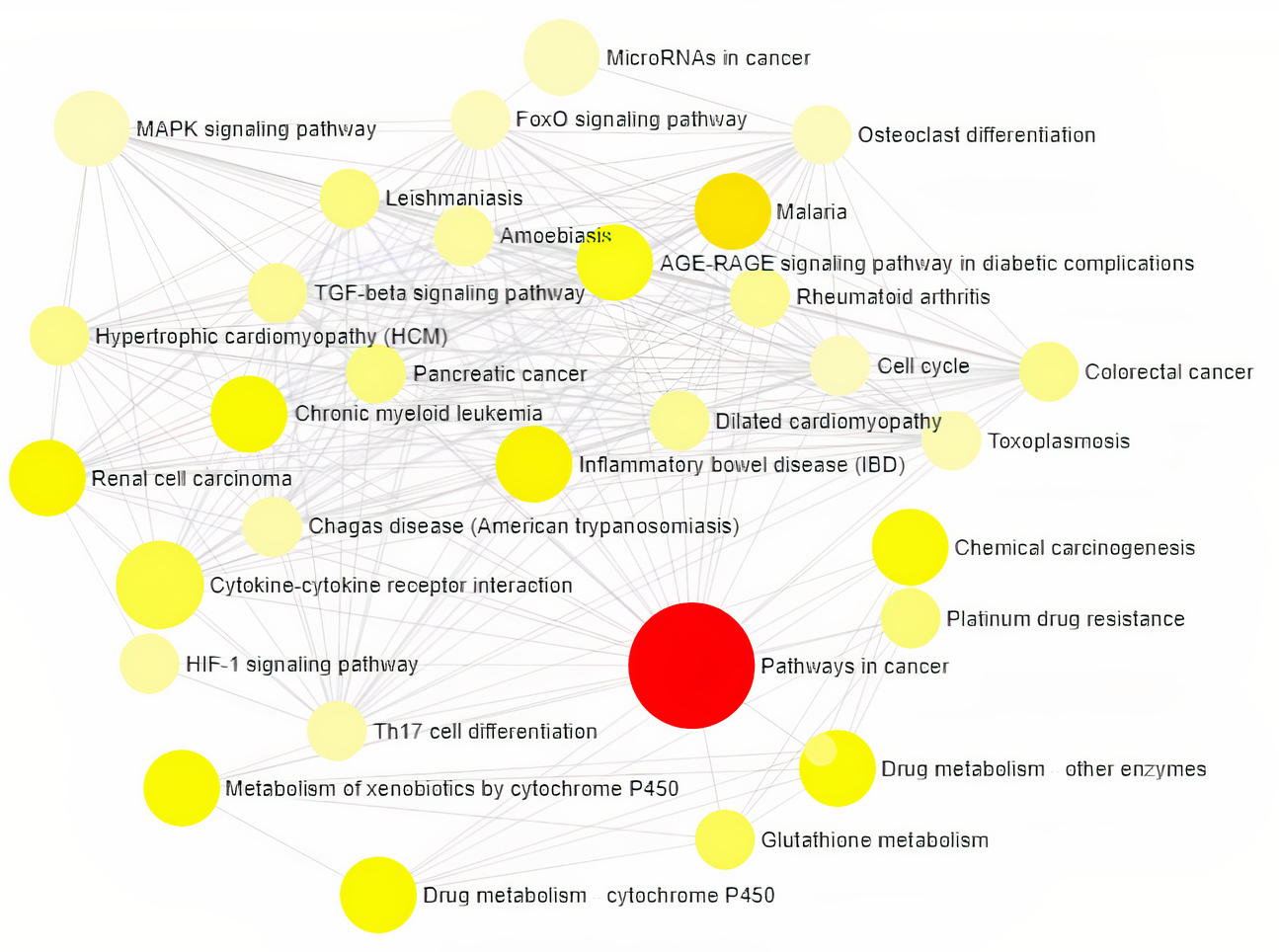 Fig. 3.
Fig. 3.Visualization of Network-based Pathway enrichment Analysis is
directly proportional to the size of the node. The nodes are color shaded
according to the significance level (adjusted p-value
The C-T-N was also constructed using the Cytoscape v3.8.2, which displays the interaction of 86 compounds with 1300 target genes (Fig. 4). These interactions revealed the multi-target properties of compounds, thus increasing the creditability of these compounds as a potent therapeutical drug for treating COPD.
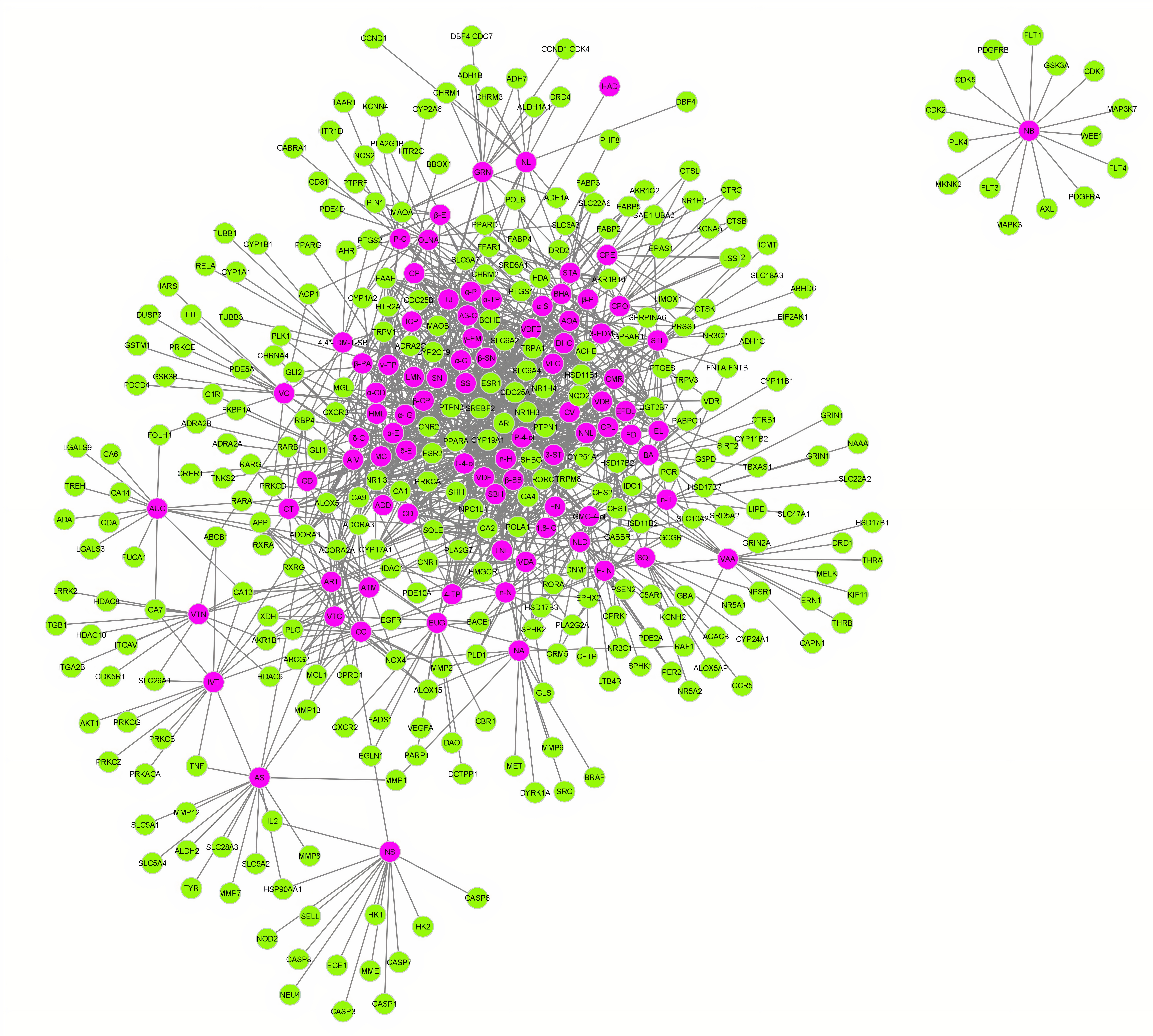 Fig. 4.
Fig. 4.Compound-Target-Network (C-T-N). The violet color indicates compounds and the green color represents human COPD targets.
Interactions between the gene identifiers revealed the multiple interactions between the human targets and were visualized using Gene Mania (Fig. 5).
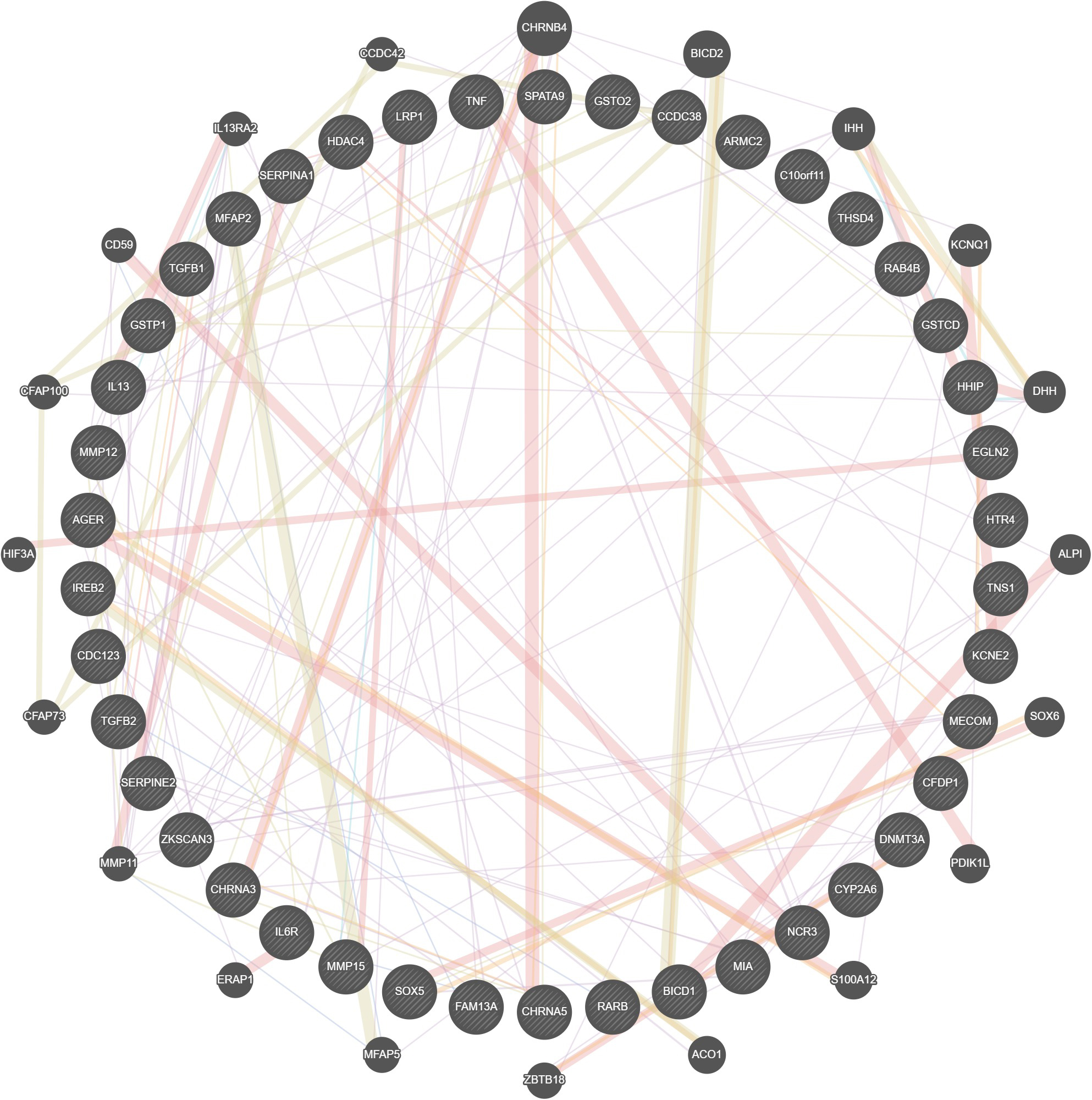 Fig. 5.
Fig. 5.Visualization of Gene Cross-talks.
Unique genes involved in COPD prognosis namely AAT, CHRNA3,
CHRNA5, IREB2, MMP12, SOX5, and
TGFB1 were identified by combinations of network pharmacology and
cheminformatics. The interaction network of these genes has 27 nodes and 133
edges. These interactions have an average of 9.85 nodal degrees within the
neighbor proteins. Protein-protein interaction of COPD unique genes enrichment
p-value was
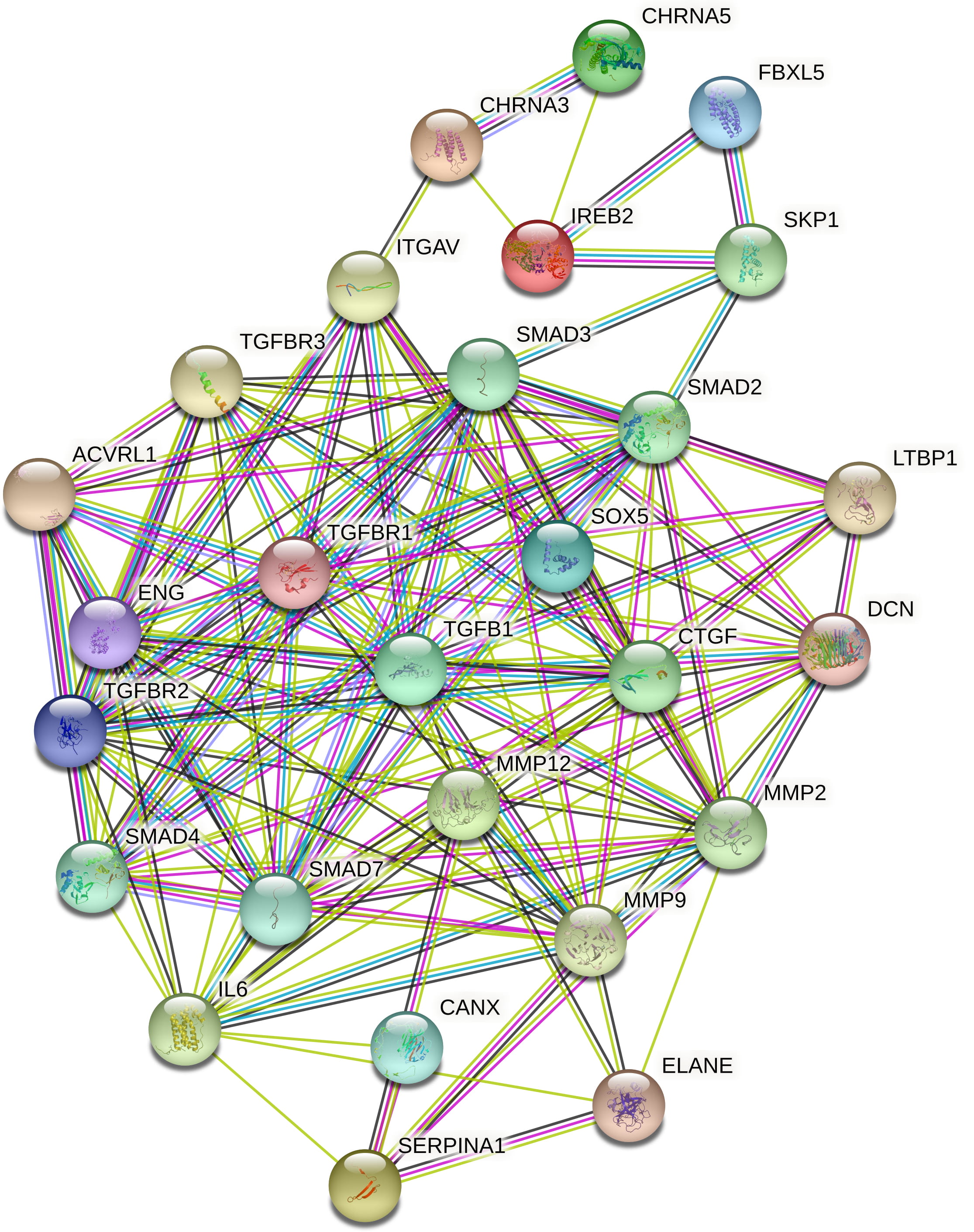 Fig. 6.
Fig. 6.COPD corresponding protein-protein interactions.
The properties of active compounds like GPCR, Ki, Pi, Ei, Ncr, and nVio were fetched from molinspiration tool. Compounds with zero nVio and with enzyme inhibitory scores above 0.5 were considered to be highly significant (Table 3). Thus, Caryophyllene Epoxide, Caryophyllene Oxide, Elemol, and Cedrol were found to be significant phytocompounds.
| Compound | GPCR lg | Ki | Ncr | Pi | Ei | nVio |
| Caryophyllene Epoxide | –0.08 | –0.86 | 0.62 | 0.00 | 0.57 | 0 |
| Caryophyllene Oxide | –0.08 | –0.86 | 0.62 | 0.00 | 0.57 | 0 |
| Elemol | –0.10 | –0.84 | 0.80 | –0.01 | 0.52 | 0 |
| Cedrol | –0.15 | –0.94 | 0.03 | –0.52 | 0.50 | 0 |
| –0.02 | –0.62 | 0.60 | –0.10 | 0.48 | 0 | |
| Vitedoamine A | 0.28 | 0.65 | 0.08 | 0.04 | 0.46 | 0 |
| Germacren-4-Ol | 0.05 | –0.55 | 0.58 | –0.28 | 0.41 | 0 |
| Vitedoin B | –0.04 | –0.33 | 0.55 | –0.08 | 0.40 | 0 |
| –0.29 | –0.81 | 0.53 | –0.32 | 0.40 | 0 | |
| –0.20 | –0.88 | 0.10 | –0.52 | 0.36 | 0 | |
| Negundin A | 0.10 | 0.12 | 0.20 | –0.14 | 0.34 | 0 |
| Carveol | –0.55 | –1.40 | 0.25 | –0.89 | 0.23 | 0 |
| Vitamin-C | –0.53 | –1.09 | –1.01 | –0.81 | 0.20 | 0 |
| –0.51 | –1.45 | –0.02 | –0.78 | 0.14 | 0 | |
| Vitedoin A | 0.04 | –0.19 | 0.07 | –0.04 | 0.14 | 0 |
| Casticin | –0.14 | 0.13 | 0.01 | –0.34 | 0.12 | 0 |
| Vitexicarpin | –0.14 | 0.13 | 0.01 | –0.34 | 0.12 | 0 |
| Artemetin | –0.15 | 0.12 | 0.00 | –0.32 | 0.11 | 0 |
| Linalool | –0.73 | –1.26 | –0.06 | –0.94 | 0.07 | 0 |
| Negundin B | 0.05 | –0.14 | 0.02 | –0.13 | 0.07 | 0 |
| Terpinen-4-Ol | –0.56 | –1.68 | –0.20 | –0.92 | 0.06 | 0 |
| Spathulenol | –0.42 | –0.68 | 0.28 | –0.36 | 0.06 | 0 |
| 4-Terpineol | –0.56 | –1.68 | –0.20 | –0.92 | 0.06 | 0 |
| Terpinen-4-Ol | –0.56 | –1.68 | –0.20 | –0.92 | 0.06 | 0 |
| Caryophyllenol | –0.06 | –0.81 | –0.25 | –0.28 | 0.04 | 0 |
| Neral | –0.86 | –1.29 | –0.42 | –0.57 | 0.02 | 0 |
| Geranial | –0.86 | –1.29 | –0.42 | –0.57 | 0.02 | 0 |
| Myrcene | –1.11 | –1.51 | –0.45 | –1.31 | –0.07 | 0 |
| –0.90 | –1.37 | –0.33 | –1.55 | –0.07 | 0 | |
| Dihydromyrcenol | –0.66 | –1.18 | –0.12 | –0.71 | –0.07 | 0 |
| 4,4ʹʹ-Dimethoxy-TransStilbene | –0.25 | –0.23 | –0.18 | –0.40 | –0.09 | 0 |
| Bornyl Acetate | –0.32 | –1.33 | –0.59 | –0.44 | –0.12 | 0 |
| Viridiflorol | –0.50 | –0.82 | –0.22 | –0.48 | –0.13 | 0 |
| –0.49 | –1.27 | –0.01 | –0.57 | –0.14 | 0 | |
| 1,8-Cineol | –0.93 | –1.60 | –1.07 | –0.90 | –0.15 | 0 |
| Limonene | –0.91 | –2.01 | –0.34 | –1.38 | –0.21 | 0 |
| Sabinene Hydrate | –0.59 | –1.22 | –0.31 | –0.43 | –0.25 | 0 |
| Viridiflorene | –0.96 | –1.08 | –0.33 | –0.61 | –0.26 | 0 |
| –0.99 | –1.55 | –0.28 | –1.31 | –0.27 | 0 | |
| Amyl Isovalerate | –0.67 | –1.19 | –0.79 | –0.58 | –0.28 | 0 |
| Aromadendrene | –0.67 | –0.98 | –0.21 | –0.67 | –0.30 | 0 |
| –0.48 | –1.50 | –0.62 | –0.85 | –0.34 | 0 | |
| –0.53 | –1.45 | –0.50 | –0.80 | –0.34 | 0 | |
| Nonanol | –0.89 | –1.13 | –0.94 | –0.98 | –0.37 | 0 |
| Eugenol | –0.86 | –1.14 | –0.78 | –1.29 | –0.41 | 0 |
| Camphor | –0.79 | –2.12 | –1.21 | –0.95 | –0.52 | 0 |
| ∆ |
–1.29 | –1.51 | –1.28 | –1.28 | –0.53 | 0 |
| Thujene | –0.96 | –1.79 | –1.13 | –1.02 | –0.58 | 0 |
| Sabinene | –1.15 | –1.79 | –0.69 | –0.78 | –0.60 | 0 |
| p-Cymene | –1.18 | –1.40 | –1.21 | –1.42 | –0.78 | 0 |
| Camphene | –1.02 | –1.85 | –1.15 | –1.40 | –0.82 | 0 |
| –0.55 | –0.86 | 0.49 | –0.64 | 0.26 | 1 | |
| Δ-Elemene | –0.36 | –0.69 | 0.63 | –0.60 | 0.46 | 1 |
| Friedelin | 0.02 | –0.39 | 0.39 | 0.02 | 0.21 | 1 |
| Farnesol | –0.13 | –0.60 | 0.20 | –0.43 | 0.42 | 1 |
| Stearic Acid | 0.11 | –0.20 | 0.17 | 0.06 | 0.20 | 1 |
| Behenic Acid | 0.17 | –0.10 | 0.23 | 0.17 | 0.17 | 1 |
| -Caryophyllene | –0.34 | –0.78 | 0.13 | –0.60 | 0.19 | 1 |
| Isocaryophyllene | –0.34 | –0.78 | 0.13 | –0.60 | 0.19 | 1 |
| Humulene | –0.14 | –0.93 | 0.34 | –0.67 | 0.31 | 1 |
| Δ-Cadinene | –0.58 | –0.75 | 0.00 | –0.68 | 0.19 | 1 |
| –0.33 | –0.79 | 0.02 | –0.49 | 0.10 | 1 | |
| Nerolidol | –0.17 | –0.64 | 0.42 | –0.43 | 0.39 | 1 |
| –0.36 | –1.02 | 0.43 | –0.38 | 0.30 | 1 | |
| n-Tritriacontane | 0.03 | –0.03 | 0.03 | 0.03 | 0.02 | 1 |
| n-Hentriacontanol | 0.06 | 0.01 | 0.10 | 0.08 | 0.08 | 1 |
| Epifriedelinol | 0.17 | –0.18 | 0.41 | 0.17 | 0.31 | 1 |
| Oleanolic Acid | 0.28 | –0.40 | 0.77 | 0.15 | 0.65 | 1 |
| n-Nonacosane | 0.04 | –0.04 | 0.04 | 0.03 | 0.02 | 1 |
| 0.14 | –0.51 | 0.73 | 0.07 | 0.51 | 1 | |
| –0.24 | –0.97 | 0.34 | –0.51 | 0.28 | 1 | |
| Acetyl Oleanolic Acid | 0.18 | –0.46 | 0.67 | 0.12 | 0.58 | 1 |
| Sitosterol | 0.14 | –0.51 | 0.73 | 0.07 | 0.51 | 1 |
| (E)-Nerolidol | –0.17 | –0.64 | 0.42 | –0.43 | 0.39 | 1 |
| –0.26 | –0.94 | 0.35 | –0.48 | 0.29 | 1 | |
| –0.24 | –1.07 | 0.01 | –0.70 | 0.41 | 1 | |
| Germacrene D | –0.30 | –0.81 | 0.32 | –0.67 | 0.26 | 1 |
| Hexadecanoic Acid | 0.02 | –0.33 | 0.08 | –0.04 | 0.18 | 1 |
| Valencene | –0.31 | –1.28 | 0.41 | –0.79 | 0.26 | 1 |
| Squalene | 0.04 | –0.10 | 0.19 | –0.03 | 0.16 | 1 |
| Vitexin | 0.13 | 0.19 | 0.23 | 0.03 | 0.46 | 1 |
| Aucubin | 0.23 | 0.03 | 0.10 | 0.35 | 0.55 | 1 |
| Isovitexin | 0.12 | 0.15 | 0.23 | 0.04 | 0.47 | 1 |
| Negundoside | 0.24 | –0.22 | 0.35 | 0.14 | 0.43 | 2 |
| Agnuside | 0.07 | –0.10 | 0.15 | 0.19 | 0.33 | 2 |
| Carotene | 0.02 | –0.14 | 0.49 | –0.12 | 0.27 | 2 |
| The bold format represents essential bioactive compounds to combat COPD. | ||||||
Primary studies have identified four phytocompounds of V. negundo L. as pharmacologically significant. Molecular docking was performed using these compounds against the COPD responsible human targets: AAT, CHRNA3, CHRNA5, IREB2, MMP12, SOX5, and TGFB1 (Supplementary Fig. 1). The binding score of docking were provided in Table 4.
| S.No. | Target | Compound | Binding energy (kcal/mol) |
| 1 | AAT | CPE | –6.3 |
| (Alpha–1 antitrypsin) | CPO | –6.6 | |
| CD | –6.8 | ||
| EL | –5.8 | ||
| 2 | CHRNA3 | CPE | –5.7 |
| (Cholinegic Receptor Nicotinic Alpha 3 Subunit) | CPO | –5.2 | |
| CD | –6 | ||
| EL | –5 | ||
| 3 | CHRNA5 | CPE | –6 |
| (Cholinegic Receptor Nicotinic Alpha 5 Subunit) | CPO | –5.7 | |
| CD | –6.7 | ||
| EL | –5.5 | ||
| 4 | IREB2 | CPE | –7.1 |
| (Iron Responsive Element Binding Protein 2) | CPO | –6.9 | |
| CD | –7.5 | ||
| EL | –5.9 | ||
| 5 | MMP12 | CPE | –5.9 |
| (Matrix Metallopeptidase 12) | CPO | –6.1 | |
| CD | –6.3 | ||
| EL | –5.5 | ||
| 6 | SOX5 | CPE | –5.9 |
| (SRY–Box Transcription Factor 5) | CPO | –5.7 | |
| CD | –5.8 | ||
| EL | –5.5 | ||
| 7 | TGFB1 | CPE | –5.6 |
| (Transforming Growth Factor Beta 1) | CPO | –5.5 | |
| CD | –5.8 | ||
| EL | –5.1 |
The FDA-approved drugs used in the treatment of COPD were collated and the pharmacological properties of the active ingredient of these drugs were computed using the Molinspiration tool. Based on these properties, phytocompounds and commercially available drugs were compared, which leads to the identification of highly potent compounds which combat COPD. The details of potential bioactives are listed in Table 5.
| Drug for COPD | GPCR lg | Ki | Ncr | Pi | Ei | nVio |
| Tudorza Pressair | 0.46 | –0.44 | –0.21 | –0.21 | 0.09 | 0 |
| Seebri Neohaler | 0.58 | –0.35 | –0.21 | –0.06 | 0.41 | 0 |
| Atrovent | 0.56 | –0.29 | –0.37 | –0.07 | 0.17 | 0 |
| Spiriva | 0.63 | –0.39 | –0.15 | 0 | 0.15 | 0 |
| Incruse Ellipta | 0.42 | –0.29 | –0.19 | 0.04 | 0.27 | 0 |
| Brovana | 0.42 | –0.16 | –0.02 | 0.18 | 0.22 | 0 |
| Foradil | 0.42 | –0.16 | –0.02 | 0.18 | 0.22 | 0 |
| Arcapta Neohaler | 0.38 | 0.08 | 0.15 | 0.11 | 0.22 | 0 |
| Serevent | 0.38 | 0.08 | 0.15 | 0.29 | 0.27 | 0 |
| Striverdi Respimat | 0.24 | –0.25 | –0.31 | 0.02 | –0.04 | 0 |
| Pulmicort | 0.21 | –0.64 | 1.27 | 0.27 | 0.67 | 0 |
| Aerobid | 0.08 | –0.48 | 1.49 | 0.45 | 0.7 | 0 |
| Qvar | –0.11 | –0.83 | 0.93 | 0.3 | 0.42 | 1 |
| Flovent | 0.15 | –0.69 | 1.83 | 0.95 | 0.81 | 1 |
| Asmanex | –0.30 | –0.82 | 0.87 | 0.05 | 0.25 | 1 |
| Alvesco | –0.03 | –0.74 | 0.78 | 0.11 | 0.33 | 2 |
| Bioactive Compounds | GPCR lg | Ki | Ncr | Pi | Ei | nVio |
| Caryophyllene Epoxide | –0.08 | –0.86 | 0.62 | 0 | 0.57 | 0 |
| Caryophyllene Oxide | –0.08 | –0.86 | 0.62 | 0 | 0.57 | 0 |
| Elemol | –0.10 | –0.84 | 0.8 | –0.01 | 0.52 | 0 |
| Cedrol | –0.15 | –0.94 | 0.03 | –0.52 | 0.5 | 0 |
| The bold format represents essential bioactive compounds to combat COPD. | ||||||
COPD is a systemic disorder [23] that involves structural alterations in both lung parenchyma and respiratory airways [24], which requires intensive research for the development of proper treatment methods. Since the allopathic formulations are not effective at present, exploration of alternative solutions has become necessary. More than 50% of the world’s population has started the usage of alternative medicines, where the Indian ancient medicinal knowledge become highly significant [25]. India had a well-established ayurvedic system since ancient times [26], with more than 1500 well-studied plants are officially incorporated in ayurvedic formulations [27]. But the exact mode and mechanism of these medicinal plants remain unknown [26]. Hence, in this study, we have integrated the pharmacology, cheminformatics, and molecular docking approaches to identify the highly potent compounds present in V. negundo L. and to reveal its mode of action in treating COPD at the molecular level.
In this investigation, 86 bio-active molecules were identified with the help of cheminformatics. All these 86 molecules have strong interactions with more than 1300 human targets directly. These compounds have been hypothesized to be involved in various molecular and biological processes against COPD. Followingly, Cytoscape 3.8.2 was used to obtain the molecular interactions and C-T-N. These results predict the mechanism of drug compounds involved in various biological processes to achieve therapeutic potential.
As highlighted by previous reports, the binding of compounds to only a single direct target does not yield relevant results [28]. From a network point of view, the receptors or genes that are needed to be targeted have shifted from unique proteins to a whole molecular network of genes involved in causing disease [29]. These multiple interactions of drugs with various targets increase the efficiency of the drugs and the probability of effective treatment [26]. Results from Swiss target prediction and C-T-N have shown that each potential compound can act on multiple targets, that are involved in the development of COPD. Since various gene regulations are intricated in causing COPD, multi-target effects of these compounds can be used to achieve superior results.
Subsequently, molecular docking analysis was performed to verify the potentials of these compounds. Results of docking display the significant binding energies between the plant compounds and the respective human targets. The genes AAT and IREB2 play a significant role in causing lung inflammation [30, 31] genes CHRNA3 and CHRNA 5 play a crucial role in lung cancer development [32], and genes MMP12, SOX5, and TGFB1 involve in the prognosis of emphysema [33, 34, 35]. These medical conditions namely lung inflammation, lung cancer, and emphysema are the hallmarks of COPD and these seven genes can be regarded as candidate genes for COPD. Targeting these genes will pave way for the treatment of COPD. Hence, molecular docking was performed against these candidate gene products. Among the phytocompounds, CD interacts with the highest binding affinities of –7.5 and –6.8 kcal/mol with IREB2 and AAT respectively. In general, CD exhibits the highest affinity with almost all the seven targets followed by CPE, CPO, and EL. These results revealed that V. negundo L. compounds possess significant regulatory actions on COPD target receptors through dynamic interactions.
Finally, highly potent molecules were identified by comparing the properties of phytocompounds with commercially available drugs, which shows the curative potential of V. negundo L. and also the proof for the Indian medical system as a highly potent approach for treating diseases, when modern treatment methods cannot yield desired outcomes.
In brief, among the 1300 genes targeted by phytocompounds present in V. negundo L., seven targets were predicted to be responsible for causing COPD using the combined analyses. Among 86 phytocompounds, four compounds were predicted to be highly potent phytocompounds, which can be employed for treating COPD after laboratory trials and validation.
The present study revealed that the V. negundo L. showed diverse immune–stimulants to treat the airflow obstruction and enhance the breathing capabilities of COPD cases. Biopharmaceutical research on V. negundo L. is still a bottleneck, interestingly our results on potentially specialized molecules and their pharmacological features, active human targets, enrichment analyses, and signaling networks have enabled the floodgates of research with the integration of Ayurveda to the era of modern medicine. This is the first investigation that identified the pivotal aspects of host immune responses to COPD. This study also conjectures that V. negundo L. phytomolecules and their combination with other biomolecules as is recommended by the Ayurvedic and modern medicine system—may result in combined effects and further studies are required.
Hence, we hypothesize that the identified potential compounds from this herb can be used to regulate the gene-targeted pathways, which can ultimately result in the cure of COPD. Yet, in vivo evaluation is required to further validate this hypothesis. Furthermore, it is suggested that obtained interaction can be useful to design competitive human immune targets antagonist which will pave the way to combat COPD.
COPD, Chronic Obstructive Pulmonary Disease; ER, Endoplasmic Reticulum; GO, Gene Ontology; C-T-N, Compound-Target-Network; nVio, Number of Violations; GPCR, GPCR Ligand Activity; Ncr, Enzymes, and Nuclear Receptors; Ki, Kinase Activity; Ei, Enzyme Inhibition Activity; Pi, Protease Inhibitory Activity.
Conceptualization—SA, PM and MR; Data curation—RJ, and MAL; Investigation—SA, PM, RJ, MAL and SKP; Supervision—J-TC, MR; Validation—RS, HS, and J-TC; Writing - original draft—SA and PM; Writing - review & editing—RS, SKP, HS, J-TC, and MR. All authors have read and agreed to the published version of the manuscript.
Not applicable.
Not applicable.
This research received no external funding.
The authors declare no conflict of interest.
