†These authors contributed equally.
Academic Editor: Graham Pawelec
Background: Dynamic contrast-enhanced (DCE) MRI is widely used to
assess vascular perfusion and permeability in cancer. In small animal
applications, conventional modeling of pharmacokinetic (PK) parameters from DCE
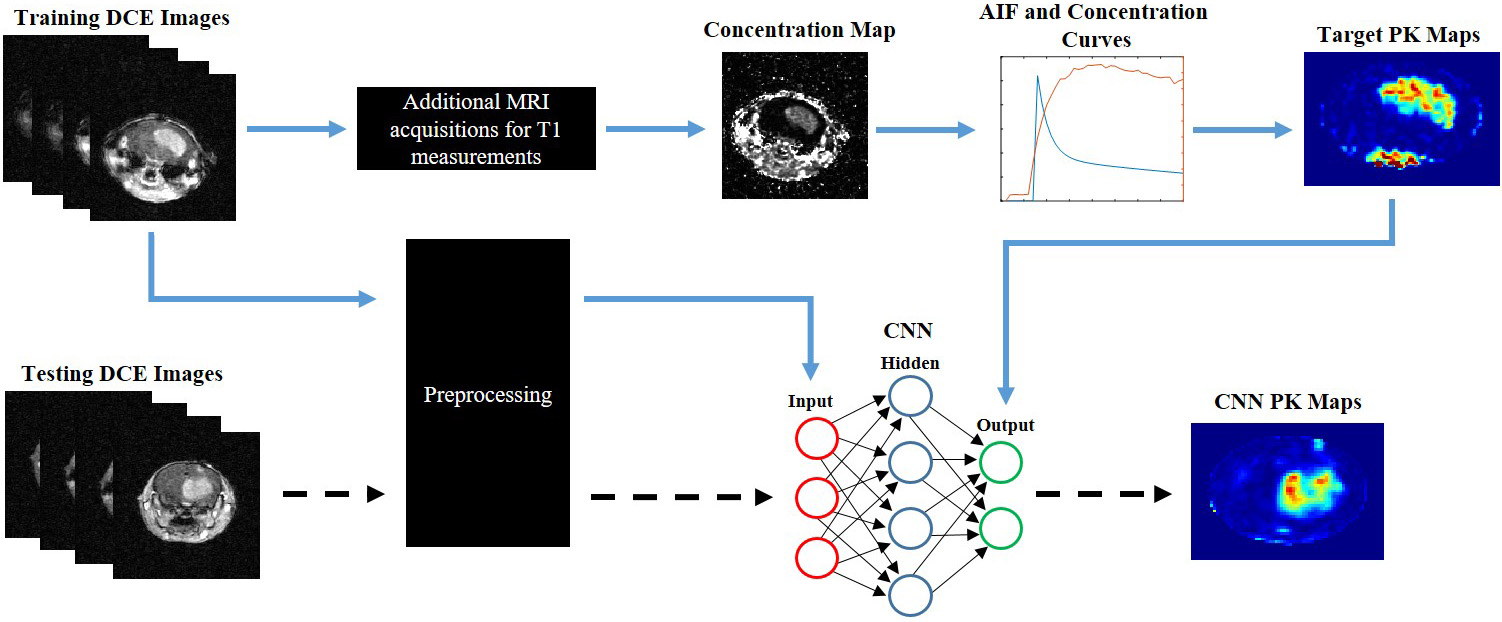
MRI images is complex and time consuming. This study is aimed at developing a
deep learning approach to fully automate the generation of kinetic parameter
maps, Ktrans (volume transfer coefficient) and Vp (blood plasma volume ratio), as
a potential surrogate to conventional PK modeling in mouse brain tumor models
based on DCE MRI. Methods: Using a 7T MRI, DCE MRI was conducted in U87
glioma xenografts growing orthotopically in nude mice. Vascular permeability
Ktrans and Vp maps were generated using the classical Tofts model as well as the
extended-Tofts model. These vascular permeability maps were then processed as
target images to a twenty-four layer convolutional neural network (CNN). The CNN
was trained on T