Here we use immunohistochemistry to examine the expression of Piezo2 in neurons of the mouse dorsal root ganglia and brain. Whereas Piezo2 is expressed in the large majority (
Epiphenomena seldom remain epiphenomena for long; they are the raw material for evolutionary advances [1].
Piezo2 is a pressure-activated channel expressed in peripheral neurons and known to play key roles in peripheral mechanosensation [2, 3, 4] including the regulation of arterial blood pressure [5] and breathing [6]. In comparison, the expression and possible function of Piezo2 in the brain remains relatively unexplored. One group using RT-PCR [7] and Western blotting [8] has reported Piezo2 expression in rat neocortical and hippocampal tissues and shown that this expression increases in response to repetitive mechanical (i.e., blast) injury. These findings raise the intriguing possibility that a sudden mechanical over activation of Piezo2 channels caused by a concussion or mild traumatic brain injury (TBI) might contribute to the rapid and often reversible disruptions in brain functions, including loss of consciousness and memory. However, Piezo2 gene/protein expression measurement in brain tissue alone does not discriminate between expression in specific neurons and/or non-neuronal cell types (e.g., glial and vascular cells). We were further motivated to make this distinction based on a patch-clamp study of mouse brain slices in which pressure-activated single-channel currents were recorded in neocortical and hippocampal pyramidal neurons [9]. In the same study, pressure activation of only a single channel current was shown to trigger action potentials in the whole neuron and with stronger channel activation, high-frequency spiking [9]. Although the channel protein was not identified at the time, it was noted that the currents displayed “Piezo-like” single-channel conductance and gating characteristics [9]. Therefore, we have used a custom generated anti-PIEZO2 antibody (Ab) and immunohistochemistry (IHC) to investigate the expression of Piezo2 in mouse brain slices. We were particularly interested in determining whether Piezo2 shows nonspecific, low-level expression, which could be functionally inconsequential, or instead is expressed in highly select cell types, similar to functionally critical voltage- and transmitter-gated channel subunits expressed in the brain. Recently, another group, using two different anti-PIEZO2 Abs, reported Piezo2 expression in rat peripheral and central neurons [10].
All experimental protocols were approved by the Animal Care and Use Committee at
the University Texas Medical Branch (UTMB) and are in accordance with the NIH
Guide for the Care and Use of Laboratory Animals. Young adult (20–25 g,
~10 weeks old) male C57BL/6J (n = 3) or GAD67-GFP (n = 3) mice
(Jackson Laboratory, Bar Harbor, Maine, USA). No differences were noted in the Ab
staining patterns in the brains and dorsal root ganglia (DRG) from the two
strains of mice. Mice were deeply anesthetized with 5% isoflurane in oxygen.
After exposing the pleural cavity, perfused through the left ventricle first with
the cold heparinized saline aorta and then with 4% paraformaldehyde in 0.1 M
phosphate-buffered saline (PBS). Their DRG and brains were rapidly removed and
stored in the same fixative overnight. Subsequently, the brains and DRG were
dehydrated through an ethanol series/xylene embedded in paraffin, and 10
The anti-PIEZO2 Ab was custom generated by Proteintech (Chicago, Il, USA) against a peptide fragment overexpressed from a cloned fragment of the human PIEZO2 (pF1KE0043) that we purchased from Kazusa DNA. Research Institute (Chiba, Japan). This vector is suitable for expressing tag-free proteins in E. coli cells and in vitro protein-expression systems driven by T7 RNA polymerase. The antigen (protein epitope signature tag (PrEST)) was a 188 amino acid peptide corresponding to amino acids 2440-2628 located at the C-terminus of the human PIEZO2 isoform (NP_071351.2) corresponding to amino acids 2512-2700 of the mouse Piezo2 isoform (NP_001034574.4). The plasmid insert was sequenced to ensure that the correct PrEST sequence was cloned. The size of the resulting recombinant protein (including the specific PrEST) was checked to ensure that the correct antigen was produced and purified. The rabbit Ab was affinity purified using an antigen-bound column. To control cross-reactivity, the affinity-purified Ab was tested for sensitivity and specificity on protein arrays consisting of glass slides with spotted PrEST fragments, including PIEZO1 and PIEZO2 fragments.
A primary aim in selecting the specific PIEZO2 PrEST was to generate an anti-PIEZO2 Ab with maximum binding affinity for the native mouse Piezo2 protein still embedded in the membrane (i.e., with its tertiary structure retained). We chose the C-terminal 188 amino acid peptide that encompasses part of the extracellular domain that contributes to Piezo2 pressure sensitivity [11, 12]. Another reason for this choice was that it should generate an Ab with high specificity for the mouse Piezo2 protein rather than, or in addition to, its paralog, Piezo1. The human PIEZO2 PrEST did display relatively high amino acid identity (i.e., 84%, with 158 out of 188 residues identical) for the corresponding region (2512--2700) of the mouse Piezo2 isoform (NP_001034574.4) but relatively low identity (i.e., 34.6% with 65 out of 188 residue identity) with the corresponding region (3226--3414) of the mouse Piezo1 isoform (NP_001032375.1). Finally, the PrEST we chose, coincidently as it turned out, also encompasses the shorter C-terminal peptide (122 amino acids) of the corresponding amino acids 2446-2568 of PIEZO2 used to generate the commercial anti-PIEZO2 Ab (Atlas antibodies, HPA015986)1 (1https://www.atlasantibodies.com/api/print_datasheet/HPA015986.pdf).
The Piezo2 immunoreactivity was detected using the anti-PIEZO2 Ab at a 1:600
dilution for 60 minutes. The brain and DRG tissues were processed using the Dako
Autostainer. The secondary Ab used was biotinylated goat anti-rabbit IgG (Vector
Labs, Burlington, CA, #BA-1000) at 1:200 for 30 minutes, followed by Dako LSAB2
streptavidin-HRP (#SA-5704) for 30 minutes. Slides were developed with Dako DAB
chromagen (#K3468) for 5 minutes and counterstained with Harris Hematoxylin for
one minute. Imaging was carried out under bright field Olympus Bx51 microscope
with Olympus DP imaging software. Because DAB staining does not fulfill the
Beer-Lambert law, it is not stoichiometric [13], and staining intensity will not
be related to the number of antigen molecules. Therefore, quantitative
measurement of expression level was not possible. DRG and brain slices were
viewed, and Ab stained cells were counted in high magnified (typically
40
Results were based on studies of six mice in which typically 3–4 slices of DRG and brains were examined. Only slices from mice that showed neurons with well-preserved morphology (i.e., when observed under high magnification) were selected for analysis. Our analysis focused on Piezo2 expression in brain regions where neuron types were unequivocally identifiable based on their characteristic morphology and layered arrangement. As it turned out, pyramidal neurons within the neocortex and hippocampus, Purkinje cells (PC) in the cerebellar cortex and mitral cells (MC) of the olfactory bulb (OB) expressed Piezo2. In contrast, the more abundant granule cells in the dentate gyrus, cerebellar cortex, and OB were absent. We considered these yes/no patterns of Piezo2 expression reproducible if the same pattern was replicated in brain slices from at least 2 different mice. Because relative counts of stained/unstained cell types/morphologies measured in brain slices from different mice appeared very similar, counts were combined.
The critical requirement for our Ab generated against a human PIEZO2 peptide was
high sensitivity and high selectivity in detecting the endogenous expression of
the mouse ortholog Piezo2 but not its mouse paralog Piezo1. The latter was
important because Piezo1 is expressed in both mouse peripheral DRG [14, 15] and central neurons [16, 17, 18]. We used mouse DRG sensory neurons to confirm Ab
sensitivity, where Piezo2 was first identified [2]. As indicated in Fig. 1A,B,
the Ab stained the large majority of mouse DRG neurons (~90%),
including the smallest (~10
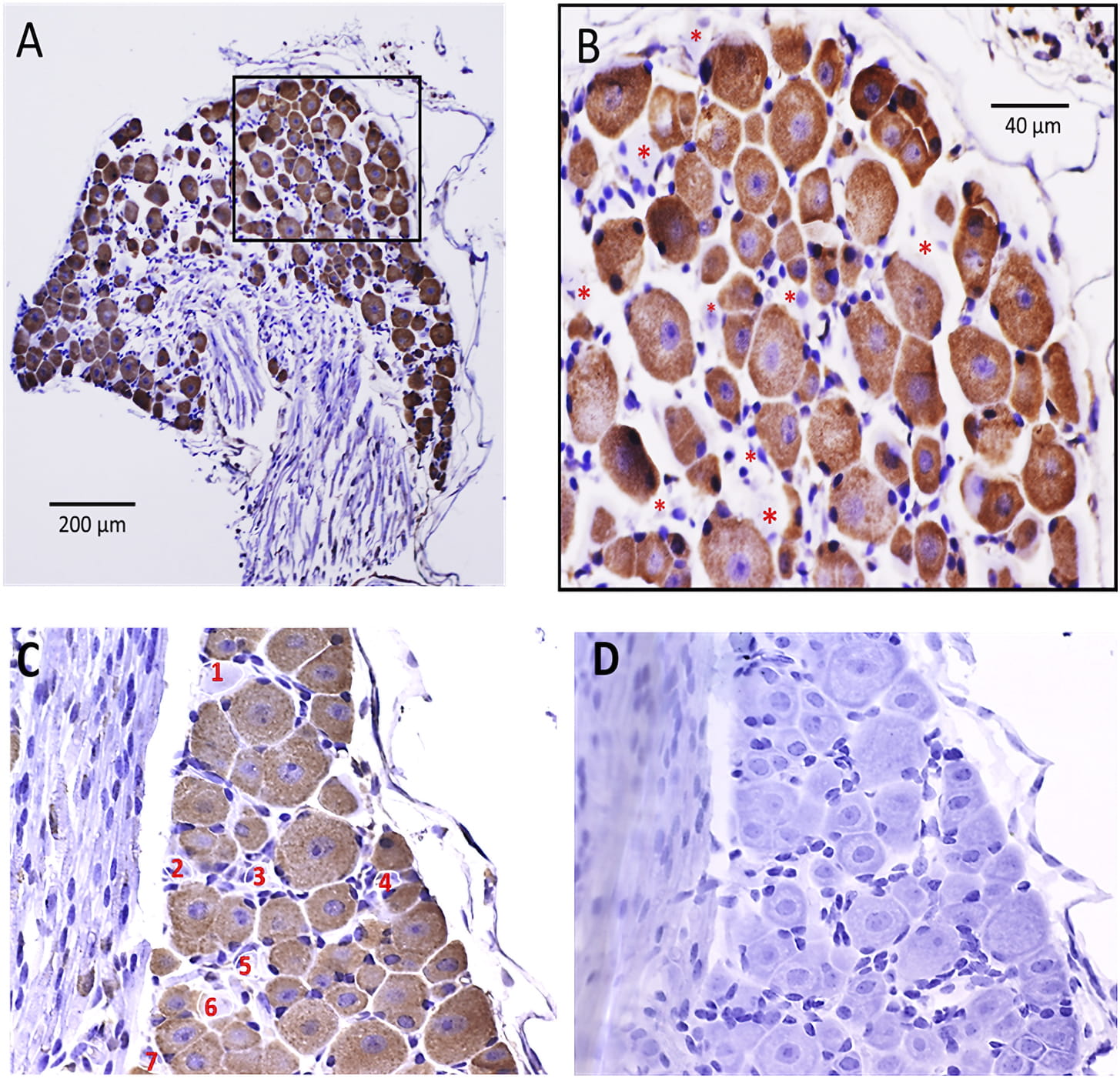 Fig. 1.
Fig. 1.Immunohistochemical localization of Piezo2 in mouse
DRG. (A) The anti-PIEZO2 Ab stained most DRG neurons independent of soma size
(range 10–50
Using brain slices from the same mice that the DRG were isolated and following identical IHC protocols, we examined the expression of Piezo2 in different brain regions. In particular, the Ab stained many cells in the grey matter of the neocortex, most notably, cells spanning across layers V and VI (Fig. 2A). Examination at high magnification indicated that the stained cells were mostly pyramidal-like according to their triangular-shaped cell body and prominent unitary (apical) dendrite oriented perpendicular to the cortical layers and directed towards the pia (Fig. 2B). In contrast, we saw a much lower abundance of stained multipolar cells or cells with no processes, which may have represented stellate interneurons, astrocytes or even pyramidal neurons upon 3-D reconstruction (not carried out here). Counting these morphological types across different neocortical regions within sagittal slices in 2 different mice indicated that for a total of 1912 Ab stained cells, 1277 (67%) showed a pyramidal-like morphology, 193 (10%) displayed multiple processes, and 442 (23%) were round with no observable processes.
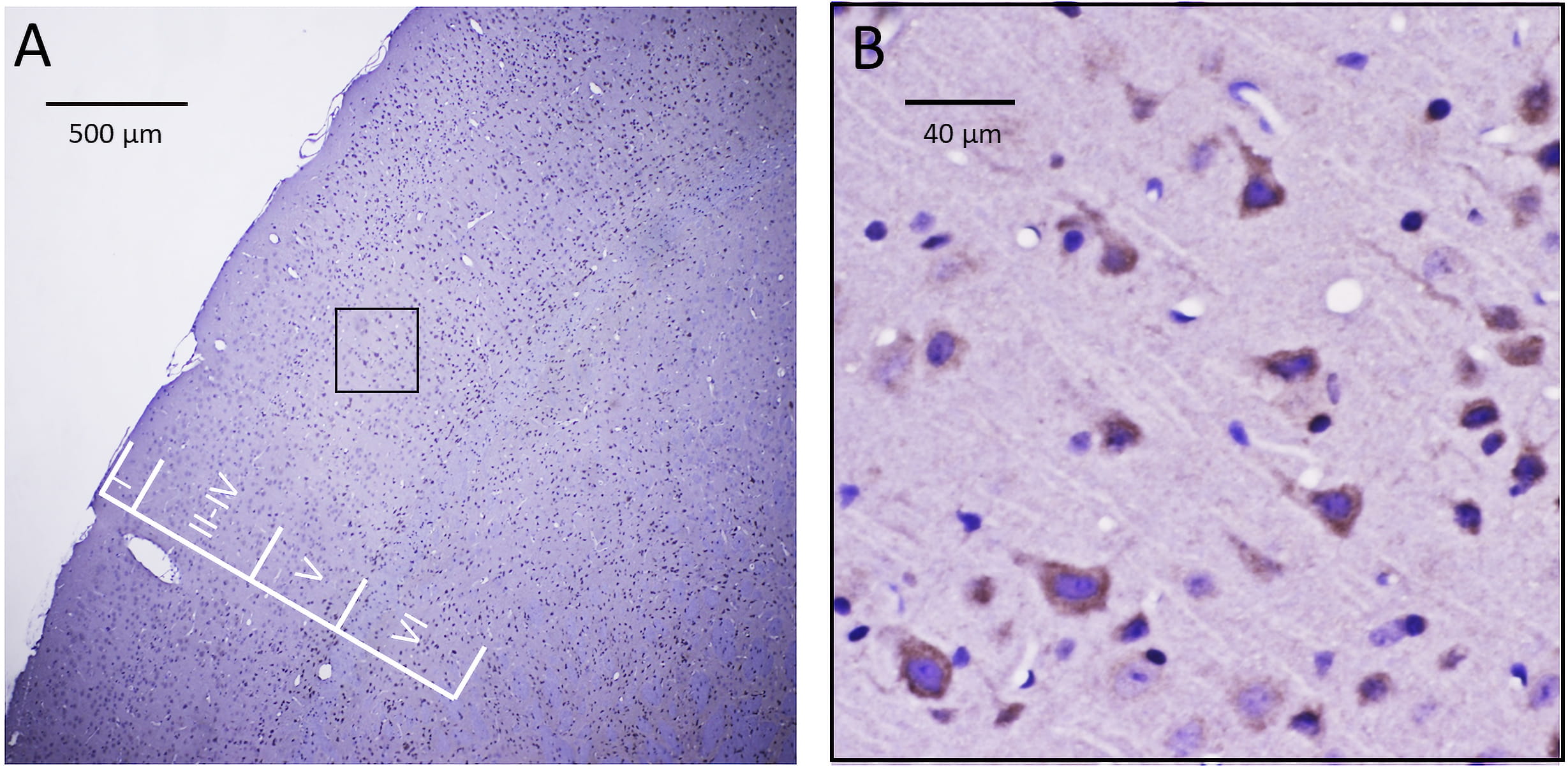 Fig. 2.
Fig. 2.Immunohistochemical localization of Piezo2 in a sagittal slice of mouse neocortex. (A) The anti-PIEZO2 Ab stained many cells throughout layers II–VI of the neocortex with minimal staining of cells in layer I. (B) The selected region in A (i.e., within the black box) representing a region within layer V is shown at higher magnification and indicates the most commonly stained cells displayed a pyramidal-shaped cell body with a thick process directed towards the pia (i.e., consistent with the apical dendrite of a pyramidal neuron). In contrast, there was less clear evidence of the staining of multipolar cells indicative of either stellate neurons or astrocytes.
In the case of the hippocampus, the Ab also showed selective Ab staining of neurons in specific regions of the hippocampus. Fig. 3A shows at low magnification the different regions (i.e., dentate gyrus (DG), CA1-4), and Fig. 3B shows a more magnified CA3 region in which several neurons show clear brown staining, some of which appeared pyramidal shaped with a prominent process. Fig. 4 shows individual regions of the hippocampus from the same slice indicating a higher abundance of Ab stained cells in the CA3-2 regions compared with other regions (i.e., DG, hilus/CA4 and CA1 regions). What appeared most obvious for the DG (see Fig. 4A–C) was that the most abundant cells in this region, the granule cells, showed no Ab staining (i.e., the round blue cells). However, near the crest of the DG, there was a small cluster of stained cells (red arrow Fig. 4A) and spindle-shaped cells located in the subgranular layer (red arrows Fig. 4B, C). These cells may reflect possibly pyramidal basket cells or even neural precursor cells. The CA3-2 regions (Fig. 4E–I) had a higher density of stained cells. Some cells had a classical pyramidal neuron appearance with a triangular-shaped cell body and a single prominent dendritic process. In transitioning from the CA2 to the CA1, there was a marked decrease in the number of stained cells. Similar staining patterns were seen in other slices from a different mouse with a lack of staining of dentate granule cells and the most abundant staining of pyramidal-like neurons in the CA3– CA2 regions. Cell counts made in these hippocampal regions from brains slices from two different mice indicated stained cells in DG represented only ~2% of the cells counted (i.e., 50/2350) with no evident staining of DG granule cells. Similarly, in the hilus/C4 regions, only ~4% of neurons (i.e., 13/370) showed staining. In comparison, in the CA3 and CA2 regions, there were ~33% (i.e., 105/315) and 24% (i.e., 39/160), respectively, but only ~5% (15/305) in the CA1 region.
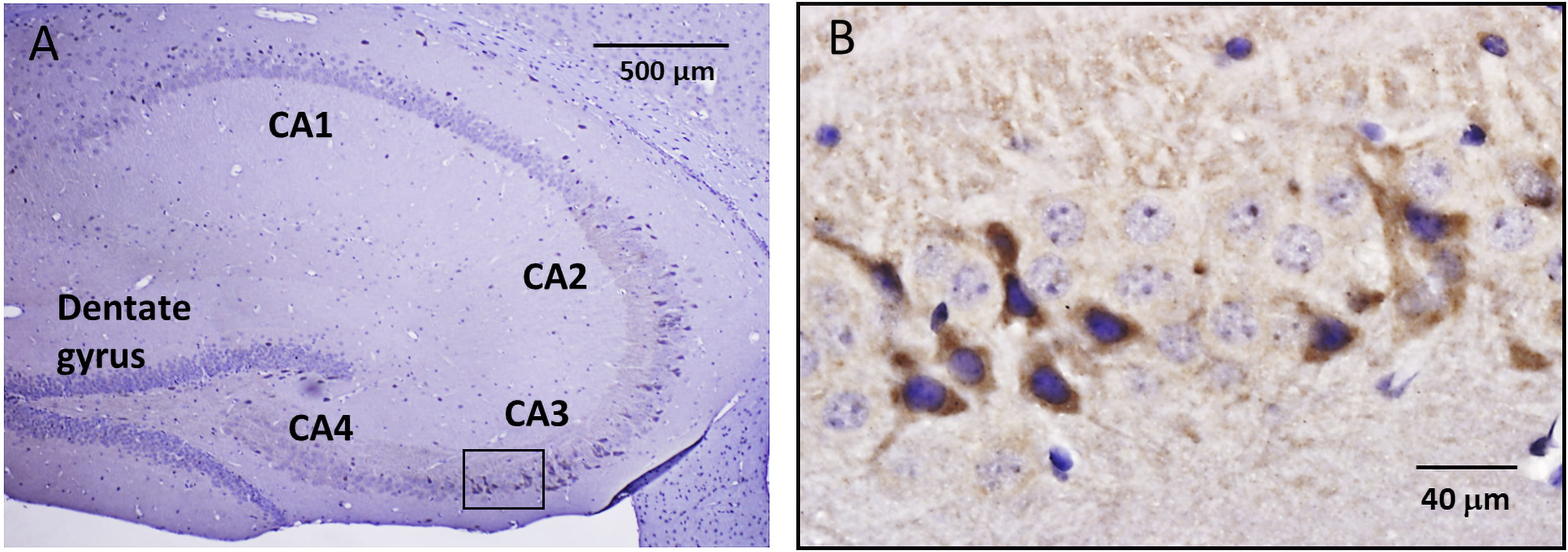 Fig. 3.
Fig. 3.Immunohistochemical localization of Piezo2 in mouse hippocampus. (A) Low magnification image of a hippocampal sagittal slice showing distinct divisions, including the dentate gyrus (DG) and CA4–CA1 regions. (B) The selected region in A (i.e., within the black box) that involves part of the CA3 region is shown at higher magnification and indicates at least 11 heavily stained cells, some of which displayed a distinct pyramidal-shaped cell body and a single prominent process. Within the same region, ~25 cells were stained blue by hematoxylin but showed no Ab staining. This all-or-none staining is at least consistent with the idea of a stochastic on-off switch controlling neuron type-specific Piezo2 expression.
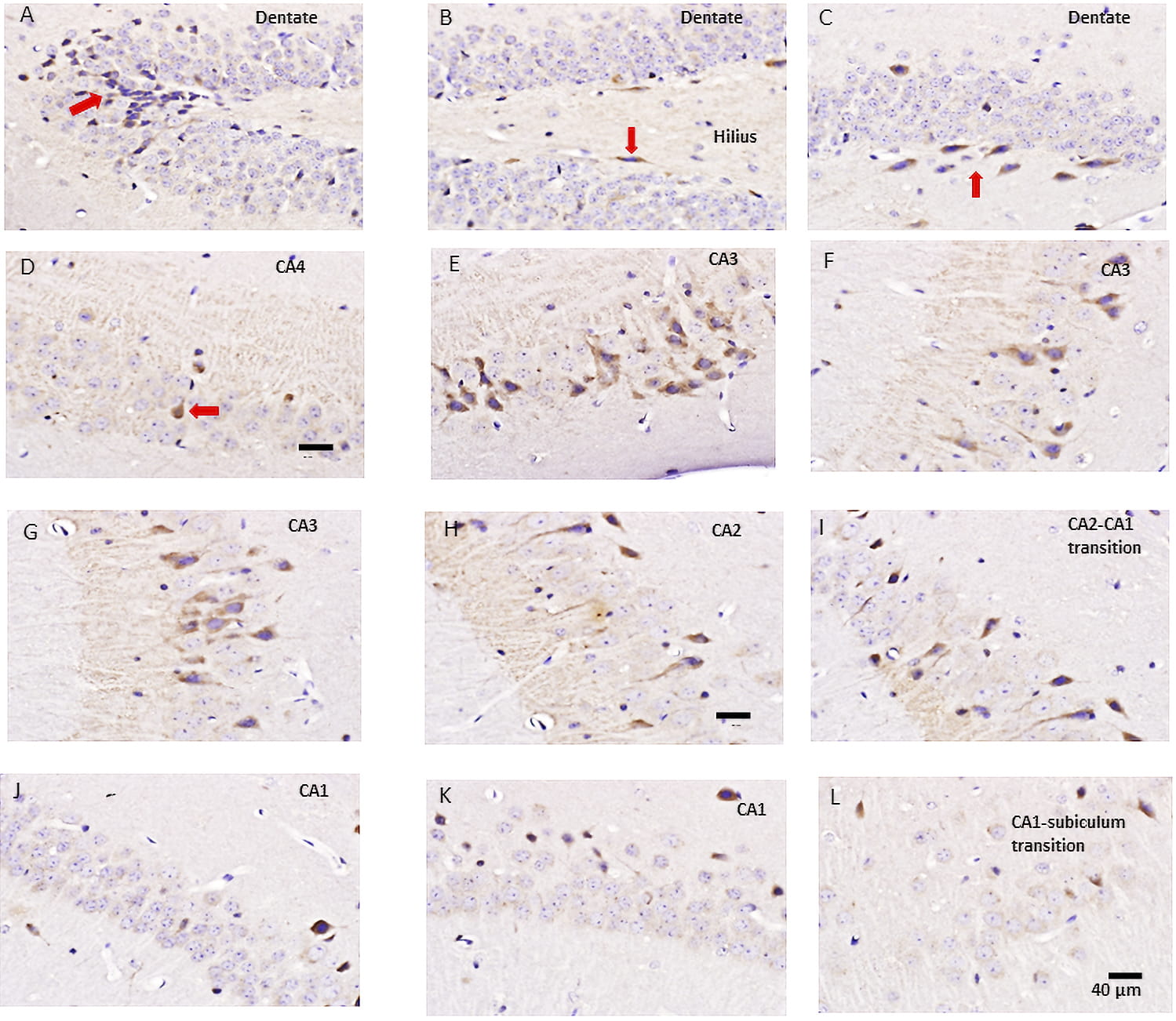 Fig. 4.
Fig. 4.Ab staining of cells in different regions of the hippocampus. (A) Shows the beginning of the DG in which most of the small round cells (i.e., granule cells) are unstained by the anti-PIEZO2 Ab. However, there was a cluster of stained cells at the crest/tip of the gyrus (red arrow). (B) Shows the adjacent region of the DG in which only 2–3 spindle-shaped cells (one indicated by red arrow) located in the subgranular zone show Ab staining. (C) A continuing region further along the DG showing 7 spindle-shaped cells stained by the anti-Piezo2 Ab. At least in this slice, the DG region only showed 16 out of 1149 cells with clear brown staining (i.e., 1.4%). (D) Shows the hilus/C4 region with at least one darky-stained cell (red arrow) but only 2 out of the 72 neurons Ab stained. (E–G) In this slice, the CA3 region showed the highest density of Ab staining, with 50 out of 140 cells stained (36%) and displayed unitary thick processes (i.e., apical dendrites) consistent with pyramidal neurons. (H) The CA2 region shows a somewhat lower density of stained neurons, with 15 out of 71 cells stained (22%). (I) A region near the CA2-CA1 transition displays a further reduced density of stained neurons (23/305, i.e., ~8%). (J–K) CA1 regions showing even fewer stained neurons. (L) CA1 region transitioning into the subiculum that displayed no Ab-stained cells.
Our results for Piezo2 expression in mouse neocortex (NC) and hippocampus (HC) can be compared with PIEZO2 expression reported by the Human Protein Atlas (HPA) using a different Ab generated against a shorter PIEZO2 peptide (see Table 1, Ref. [8, 10, 22]). This Ab also stained neurons but not glial cells in the human NC and HC (see Supplementary Fig. 2A). In this case, the Ab results were orthogonally validated by PIEZO2 gene expression in human NC and HC tissues [24, 25], see Supplementary Fig. 2B2 (2https://www.proteinatlas.org/ENSG00000154864-PIEZO2/tissue). On this basis, HPA has assigned this Ab result an “Approved” reliability score in the 2021 revision of their website (Dr. Borbala Katona, HPA Tissue Atlas Team, Uppsala University, personal communication). In another recent study of Piezo2 expression in rat peripheral and central neurons using two different anti-PIEZO2 Abs (Table 1), the Ab stained NC and HC neurons but not astrocytes [10]. Finally, a scRNA-seq study [26] has confirmed Piezo2 gene expression in mouse HC, with relatively higher gene expression in CA3 compared with CA2 and CA1 regions but with the DG cells showing a high transcript copy number ([26], and personal communication, Dr. Andres Vieira). It remains to be determined whether this reflects a real difference between gene transcription and translation of functional protein in hippocampal neurons.
| Ab | Antigenic peptide | DRG | CNS | Ref/Antibody |
| 1 | C-termini 2440-2628 | 90% | Yes | Current, custom |
| 2 | N-termini 351-485 | N/T | [22], custom | |
| 3 | Not reported | N/T | Yes | [8], ProSci, 26–438 |
| 4 | C-termini 2446-2568 | N/T | Yes | HPA, HPA015986 |
| 5 | N-termini 845-864 | Yes | [10], Prosci/8613 | |
| 6 | Internal 1092-1104 | Yes | [10], Alomone/APC090 |
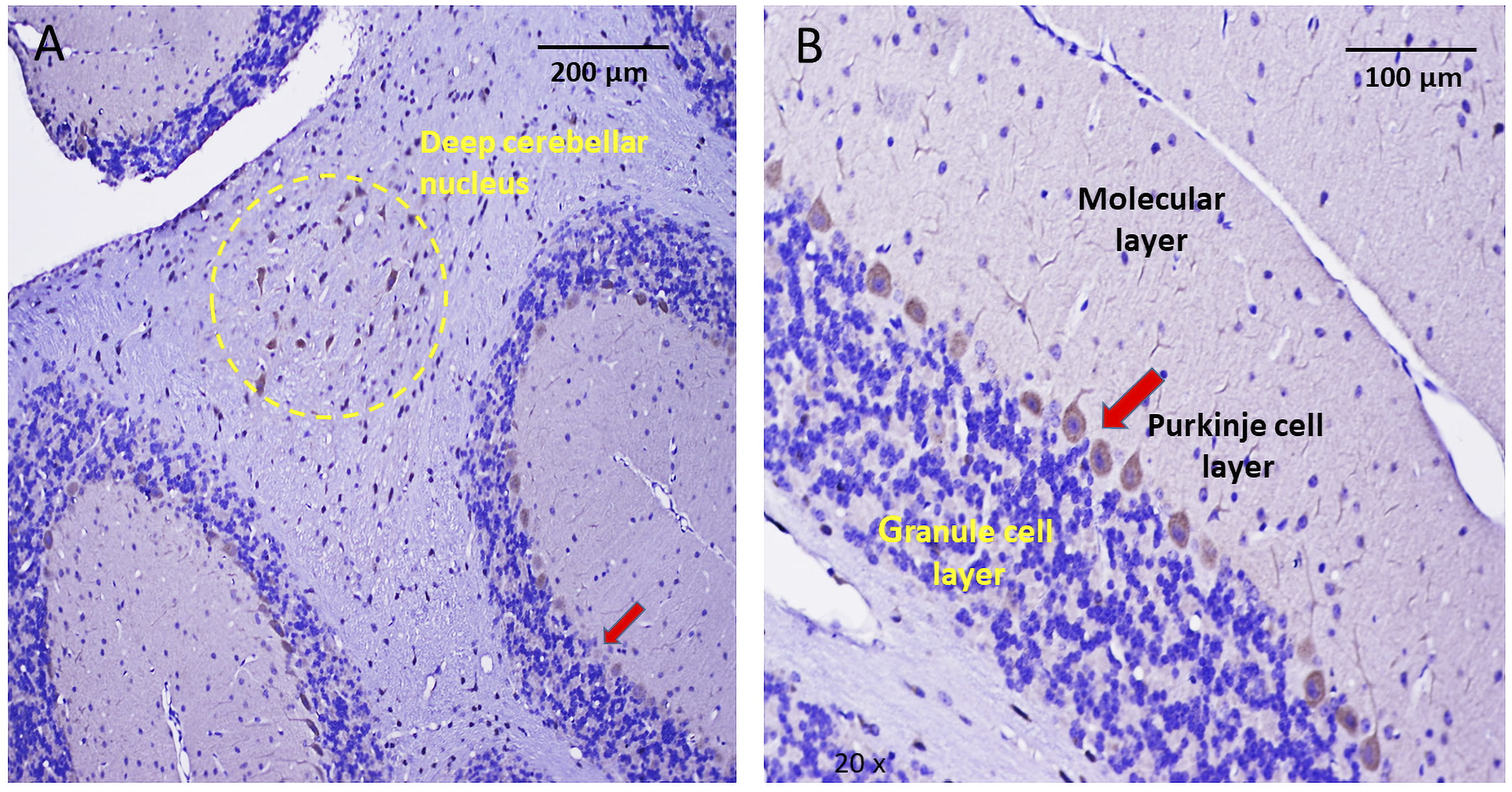 Fig. 5.
Fig. 5.Immunohistochemical localization of Piezo2 in the cerebellum. (A) Low magnification of a cerebellar slice indicating the stained Purkinje cell layer (red arrow) that separates the granule and molecular cell layers. The yellow dashed circle within cerebellar white matter (arbor vitae) encloses a group of stained neurons, presumably part of a deep cerebellar nucleus. (B) Higher magnification of the cerebellar cortex showing the characteristic single layer of Purkinje cells, which appear uniformly stained by the anti-PIEZO2 Ab. In contrast to the stained Purkinje cells, the more numerous granular cells within the granular cell layer were not stained. Similarly, apart from Purkinje cell dendrites, there was no pronounced Ab staining of cells within the molecular layer.
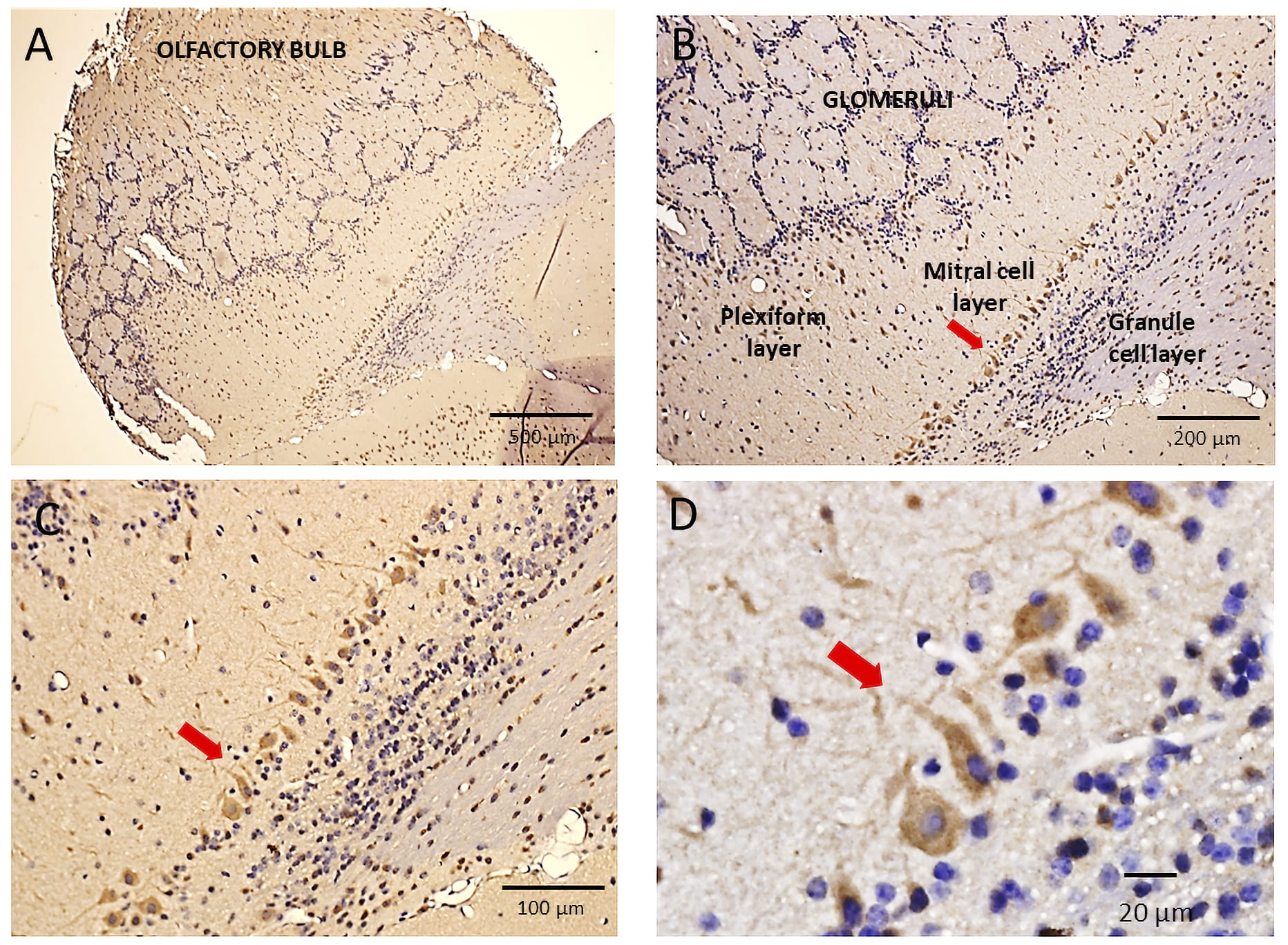 Fig. 6.
Fig. 6.Immunohistochemical localization of Piezo2 in the mouse
olfactory bulb. (A) Shows a low magnification image of the mouse olfactory bulb
(OB) with its characteristic circular/spherical glomeruli structures spanning the
OB. These glomeruli include the synaptic connections formed between primary
olfactory nerve (PON) axons and mitral cell dendrites. (B–C) Higher
magnification images (10
As indicated in Fig. 5, the Ab selectively stained Purkinje cells (PC) in the cerebellar cortex, including their relatively large cell bodies and the branching dendrites that extend into the molecular cell layer (Fig. 5B). In marked contrast, the granule cells (GC) that form the granular layer showed no evidence of staining. This all-or-none selectivity in staining PC vs. GC again highlights the selectivity in Piezo2 expression given a PC:GC ratio of 1:3000 in the cerebellar cortex [27]. Interestingly, as indicated in Fig. 5A (i.e., yellow circle), some neurons embedded within the cerebellar white matter (arbor vitae) and possibly representing projection neurons within a deep cerebellar nucleus also showed Ab staining. Similar selective staining of PC in the human cerebellar cortex has been previously reported by the HPA using another anti-PIEZO Ab [24] (see Supplementary Fig. 2C and Table 1)3 (3https://www.proteinatlas.org/ENSG00000154864-PIEZO2/tissue/cerebellum).
Fig. 6A shows a low magnified image of the mouse OB and indicates Ab staining of the OB neuropil with the circular glomeruli delineated by the hematoxylin stain. Fig. 6B–D show progressively magnified images of the same OB indicating the strongest brown staining was of the mitral cells (MC) that form a distinct boundary cell layer separating the plexiform from the granular cell layers. The neuropil staining in the external plexiform layer presumably represents stained mitral cell dendrites that project to the glomeruli, where they receive synaptic inputs from primary olfactory neurons (PONs). Tufted cells located in the external plexiform layer, which also receive synapses from PONs, did not appear stained. In addition, as indicated in Fig. 6D, the most abundant cells of the OB, the granule cells, showed no evidence of positive (brown) staining.
This expression of Piezo2 in the MCs of the OB was unanticipated, especially given their well-recognized role in processing primary olfactory sensory inputs with no prior suspected intrinsic mechanosensitive function. However, a recent study using single nucleus RNA sequencing indicates that Piezo2 is a genetic marker of specific MCs within the mouse OB [28].
The findings here that Piezo2 is expressed in select neurons of mouse brain augments earlier studies showing Piezo2 expression in isolated rodent brain tissues [2, 7, 8]. Furthermore, as described in each specific results section, the Piezo2 expression we observed in select mouse brain neurons has been confirmed by other independent proteomic and/or transcriptomic studies of rodent and/or human brain [10, 24, 25, 26, 28] (and see Supplementary Fig. 2). The all-or-none, cell-type-specific Piezo2 expression in mouse brain neurons is also interesting in relation to the stochastic on-off transcriptional switch that seems to regulate Piezo1 expression in mouse and human brain [16]. In this case, pathological conditions, including Alzheimer’s disease (AD) and infection, can switch Piezo1 expression between neurons and astrocytes [16, 17, 18].
In contrast to the increasing proteomic and transcriptomic evidence for Piezo2
brain expression in brain neurons, there is no direct evidence for functional
expression. However, a recent patch-clamp study of mouse brain slices has
reported single “Piezo-like” pressure-activated single-channel currents in
neocortical and hippocampal pyramidal neurons [9] and another study, using
Ca
Interestingly, TBI is only one of several neuropathological conditions,
including hydrocephalus, cerebral hemorrhage and brain tumors, known to elevate
the normally low (
Although expression of Piezo2 in brain neurons may confer no survival advantage,
the apparent Piezo2 related response to concussion/TBI (i.e., with ICP
Direct experimental evidence for the actual idea that breathing can induce entrainment of neural networks goes back 80 years, with the discovery by Edgar Adrian that nasal breathing in rodents causes the rhythmic firing of mitral neurons within the olfactory bulb and neurons within neurons the piriform cortex [42]. Many subsequent studies have extended his discovery by showing that nasal breathing not only entrains OB oscillations at the rodent’s breathing frequency (0.5–5 Hz) but also modulates the amplitude of higher frequency oscillations (80–120 Hz) in the OB as well as other downstream brain regions including the hippocampus and neocortex [43, 44, 45]. Significantly, these neural network oscillations have also been associated with specific changes in rodent behaviors, including whisking, memory formation and emotional (fear) responses [46, 47, 48, 49, 50, 51, 52, 53, 54, 55]. Moreover, the many observations related to respiration-locked oscillations seen in rodents have now been confirmed in humans, using either intracranial EEG recording from epileptic patients or high-density EEG recording from healthy subjects [40, 41, 56, 57].
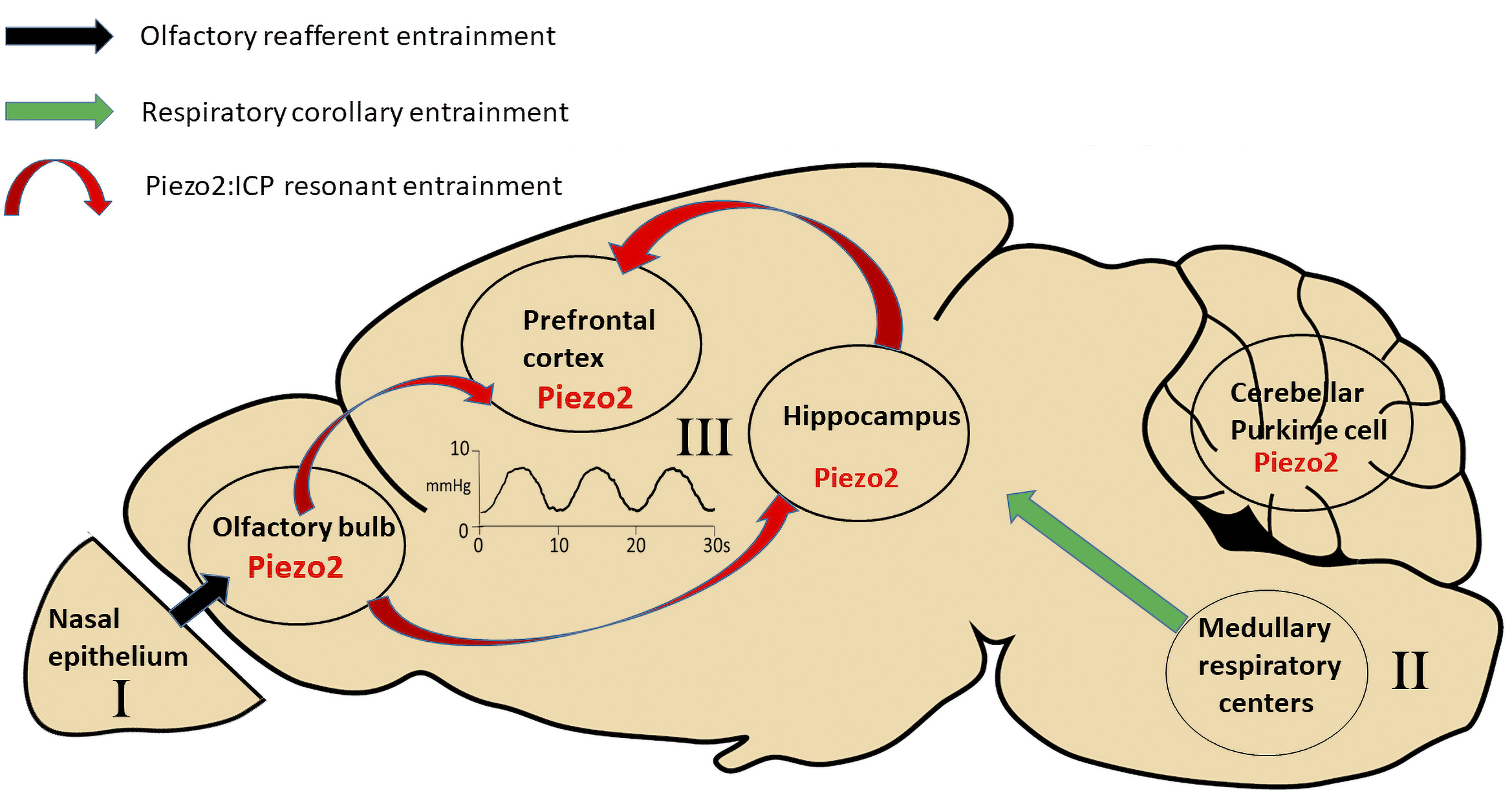 Fig. 7.
Fig. 7.Different mechanisms for respiration-induced
entrainment of neural networks in widely spaced regions of the brain. The first
mechanism involves respiratory olfactory re-afferent discharge (ORD). It
originates in the nasal epithelium, where primary olfactory neurons (PONs), which
express olfactory G protein-coupled receptors (GPCRs) that are also
mechanosensitive, can detect the nasal airflow pressure changes associated with
nasal breathing. The PONs, through their direct synaptic connections (black
arrow) with mitral cells and tufted cells in the OB, cause respiratory
entrainment of the neural activity within the OB. In turn, the OB entrains neural
activity in the hippocampus and prefrontal cortex through their monosynaptic
and/or polysynaptic connections (not shown) with these brain regions. The second
mechanism depends upon respiratory corollary discharge (RCD) arising from the
neurons in the medullary respiratory nuclei that control the diaphragm muscles
and send efferent copy discharges to high brain regions (e.g., hippocampus and
neocortex) via yet to be identified synaptic pathways. The third mechanism,
intrinsic resonance discharge (IRD), depends upon the pressure-sensitive channel,
Piezo2, expressed in mitral cells of the OB and pyramidal cells of the
hippocampus and neocortex. Piezo2, based on its demonstrated high sensitivity to
low-pressure pulses (
At least three different, non-mutually exclusive mechanisms may explain respiration-induced neural network oscillations [43, 44, 45, 46]. The first, called olfactory re-afferent discharge (ORD), proposes that nasal airflow during nasal breathing activates mechanosensitive PON that, in turn, activate via their direct synaptic connections, MCs within the OB [58] (see Fig. 7). In this mechanism, the OB acts as a “global clock” for other brain regions, synchronizing via its synaptic connections, neural networks across widespread brain regions, including the hippocampus and neocortex [44, 45]. Evidence supporting this mechanism is that tracheotomy or ablation of the nasal epithelium (or removal of the OB) in rodents reduces the synchronized activity in the OB and/or downstream brain regions [42, 43, 59, 60]. Moreover, in both humans and rodents, air pressure pulses delivered to the nostrils can restore and/or alter the oscillation frequency [51, 57].
The second mechanism referred to as “respiratory corollary discharge” (RCD)
depends upon efferent copy discharges from neurons in the brain stem nuclei that
regulate breathing [61, 62, 63] (see also Fig. 7). Support for this mechanism includes
the finding that although tracheotomy or ablation of the nasal epithelium reduces
the neural oscillations in rodents, they do not completely block them [46, 63].
Moreover, in humans, the oscillations persist during mouth breathing and,
therefore, in the absence of any nasal airflow [56, 63]. Further evidence for the
RCD mechanism is that neurons in the brain stem respiratory nuclei form
connections with neurons in the locus coeruleus that via their projections may
alter activity in widely spaced neural networks throughout the brain [62]. An
added twist to both the RCD and ORD mechanisms is the recent report [64] that low
frequency (i.e.,
The third mechanism and the least investigated, we refer to as intrinsic resonance discharge (IRD). In this case, the intrinsic properties of the neural networks involving either intrinsic neuronal membrane properties or synaptic micro-circuitry dictate that they fire or resonate at a frequency close to the respiratory frequency [41]. The evidence for this mechanism is again that respiration-entrained oscillations are retained during mouth breathing or in the absence of nasal airflow or functional PONs [46, 56, 63]. Moreover, IRD may result in part account for a somewhat puzzling feature of the ORD mechanism, which relates to the mechanosensitivity of the PON and their odor-sensitive G-protein coupled receptors (GPCRs) [58]. This GPCR mechanosensitivity depends upon a second messenger pathway and consequently displays long latencies (~300 ms) in response to pressure pulses [58].
Furthermore, because of GPCR adaptation, their maximum frequency response to
repetitive pressure pulse stimuli is only ~0.5 Hz [58]. This is
significantly lower than the rodent’s range of breathing frequency (1–10 Hz but
as high as 15 Hz during sniffing) as well as the entrainment frequency induced in
the MCs of the OB (1–10 Hz) [65]. Although the convergence of many PON inputs to
the MCs may still generate rhythmic activity that follows respiration, even when
only a fraction of the PONs responding with each respiratory cycle [58], an
alternative or reinforcing mechanism could be that the MCs themselves possess an
intrinsic mechanosensitive resonance to the breathing cycle. Specifically, we
hypothesize that Piezo2 in MCs and neurons in the neocortex and hippocampus
confer an intrinsic resonance that reinforces their entrainment by the breathing
rhythm via their transduction pulsatile ICP changes (Fig. 7). Because
stretch-activated channels, including Piezo channels, are characterized by their
fast gating [66] and ability to accurately track relative high frequency (e.g.,
An intrinsic Piezo2/ICP resonance mechanism may also have acquired a more predominant role for human breathing-induced entrainment because of additional selective pressures. In particular, in humans, nasal breathing is not obligatory as in rodents. In addition, olfaction has become a relatively unimportant sensory function, adding little advantage for survival, compared with rodents where olfaction, sniffing and whisking are tightly linked and critical for their normal behavior and survival. Moreover, humans appear to be the only species capable of volitionally switching breathing patterns (e.g., forced inspiration/expiration cycles) to promote mood, cognition, and emotional changes. Humans may also be unique in that they can choose to modulate volitionally through their breathing, both their heartbeat and emotions [69, 70]. Interestingly, a study of single-unit recording from epileptic patients indicated a higher proportion of hippocampal and amygdala neurons show entrainment with the cardiac cycle compared with the respiratory cycle [71].
Regardless of the exact mechanism, a key issue for all mechanisms is their reliability and degree of synchronization that they can induce within widely separated neural networks. This synchronization has special implications for the unity of experience related to perception, motor control, and memory retrieval, especially in the larger human brain. In the case of the first two mechanisms, the overall synchronization will depend upon signaling delays within multi-synaptic, axonal transmission pathways. In comparison, a Piezo2/ICP pulse dependent IRD mechanism will only be rate-limited by the speed of ICP pulse transmission throughout the brain, powered by the perennial actions of the respiratory and cardiac pumps. In the ideal case, when Pascal’s law applies, namely a rigid, fluid-filled and closed container, a pressure pulse will spread throughout the container almost instantaneously at the speed of sound (i.e., ~1500 m/s) [31]. However, this may represent an upper estimate since the brain contained within the skull is a semi-closed system that involves cerebral venous outflow and arterial inflow [30, 31, 32]. Moreover, CSF and blood circulate between the brain and spinal cord, particularly during inspiration and expiration [36, 37, 38]. In addition, the brain parenchyma, ventricles, and vasculature are themselves compliant rather than rigid, which will also tend to dissipate and slow the spread of ICP changes.
In one notable study [74], aimed at measuring ICP transmission in the human
brain, two ICP sensors, placed 5 cm apart in the lateral ventricle and brain
parenchyma measured a phase shift of 10 ms [74]. This would predict a pulse
velocity of only 5 m/s. (i.e., which compares with a predicted 50
The current immunohistochemical evidence for Piezo2 expression in specific mouse brain neurons is now supported by independent rodent and human brain studies using different anti-PIEZO2 Abs generated against different regions of the PIEZO2 protein (Table 1). In specific cases, the Abs have met additional validation criteria4 (4https://www.proteinatlas.org/about/antibody+validation). including orthogonal (i.e., IHC/ transcriptomic data correspondence) [24, 25, 26, 28] (see Supplementary Figs. 1,2) and genetic criteria [10]. Nevertheless, and ideally, verification of the IHC results with Ab-independent techniques is still required (i.e., using GFP-tagged Piezo2 transgenic mice [76], GENSAT). Furthermore, since Piezo1 shows expression in rodent and human brain neurons [16, 17, 18] and both Piezo1 and Piezo2 function in peripheral baroreception [5], it will be important to establish their individual expression patterns and roles in specific brain neurons. However, the biggest future challenge will be demonstrating that Piezo channel activity participates in the entrainment of neural network oscillations in living, breathing animals. One approach may involve using conditional Piezo knockout mice to compare respiratory entrained neural network oscillations with those reported previously for wild-type mice [43, 44, 45, 46, 47, 48, 49]. It should also be interesting to study human patients that show specific loss of PIEZO2 function [77] and compare how their measured EEG and behavioral responses to specific breathing protocols differ from those seen in normal human subjects [39, 40, 41, 56, 57].
Finally, recalling the fuller and particularly insightful quote from Hutcheon and Yarom [1]:
“A question therefore arises: are resonances used by neurons, or are they simply epiphenomena? A broad answer to this question is that, in nature, epiphenomena seldom remain epiphenomena for long; they are the raw material for evolutionary advances. It would be surprising to find that the brain has not found a use for a set of mechanisms capable of tuning neurons to specific frequencies, particularly in light of the prevalence of robust brain rhythms.”
Here, Hutcheon and Yarom are referring to, at the time, the relatively newly discovered roles of the various EEG recorded electrical brain rhythms [78]. However, it may also turn out the metronomic-like and perennial ICP pulses, transduced by pressure-sensitive channels, prove even more pivotal in tuning and synchronizing the brain to the fundamental rhythms of life [44, 45].
Ab, antibody; AD, Alzheimer’s disease; CSF, cerebrospinal fluid; DG, dentate gyrus; DRG, dorsal root ganglion; EEG, electroencephalography; GFP, green fluorescent protein; GC, granule cell; GPCR, G-protein coupled receptor; HC, hippocampus; HPA, Human Protein Atlas; ICP, intracranial pressure; IHC, immunohistochemistry; IRD, intrinsic resonance discharge. MC, mitral cell; NC, neocortex; NPH, Normal Pressure Hydrocephalus; OB, olfactory bulb; ORD, olfactory re-afferent discharge; PC, Purkinje cell; PON. primary olfactory neuron; PrEST, protein epitope signature tag; RCD, respiratory corollary discharge; snRNA-seq, single nucleus RNA sequencing; scRNA-seq, single cell RNA sequencing; TBI, traumatic brain injury.
JW surgically isolated and prepared the mouse brains and DRG for slicing. OH conceived the study, analyzed and interpreted the IHC experiments and wrote the manuscript. All authors read and approved the final manuscript.
Mice were housed in the animal facility at UTMB, Galveston and all experimental protocols were approved by the Animal Care and Use Committee at the UTMB (The ethical approval number for our animal protocol is 0907049) and are in accordance with the NIH Guide for the Care and Use of Laboratory Animals.
We thank Jin Mo Chung for providing his laboratory resources and support for Jigong Wang. We also thank Thomas Green for critical reading the manuscript, and Daniel Jupiter, and Andrzej Kudlicki for useful discussions. We also acknowledge the technical assistance of Kerry Graves of the UTMB Pathology core facility for preparing the tissues for IHC and Kenneth Escobar in providing the microscopy facilities.
The project was supported by a UTMB bridging grant to OPH and an NIH Grant NS03168 to Jin Mo Chung.
The authors declare no conflict of interest.
