Merlin ((Moesin-ezrin-radixin-like protein, also known as schwannomin) is a tumor suppressor protein which is encoded by the neurofibromatosis type 2 gene, NF2. Loss of function mutations or deletions in NF2 which normally restrains tumor growth, leads to the formation of multiple tumors including schwannoma, meningioma and ependymoma. We tested whether NF2/Merlin is expressed and exerts similar control on proliferation of colorectal cancer cells and modulates the rate of their apoptosis. Expression of NF2/Merlin was reduced in colorectal cancer cells as compared with adjacent non-cancerous cells. Overexpression of NF2 inhibited colony formation by tumor cells and inhibited proliferation of cancer cells both in vitro and in vivo. The rate of apoptosis was also increased in colorectal cancer cells by overexpression of NF2. These findings show that NF2/Merlin is also reduced in tumors that do not arise in the context of neurofibromatosis and that induction of its expression might be used to control tumor growth.
Colorectal Cancer (CRC) is one of the most frequently diagnosed neoplasms across the world (1-2). The prognosis of patients with colorectal cancer is closely related to the stage at which the tumor is diagnosed with late detection leading to a poor prognosis (3-4). Despite great advances in cancer treatment, the prognosis of colorectal cancer is still quite poor (5). Thus, there is a need to further understand the complex progression of colorectal cancer to identify better therapeutic targets (6).
Merlin ((Moesin-ezrin-radixin-like protein, also known as schwannomin), also known as neuro-fibromin 2, is a tumor suppressor protein with 595 amin acid residues which is encoded by the neurofibromatosis type 2 gene, NF2 (7-8). Loss of function mutations or deletions in NF2 which normally restrains tumor growth, leads to the formation of multiple tumors including schwannoma, meningioma and ependymoma (9-10). Merlin restrains tumor growth and reverts malignant phenotype by inhibiting p21 Ras-activated kinase, disrupting Rac-dependent signaling, constraining activation of mTOR complex1 and mediating contact-dependent inhibition of EGFR mobility and internalization (11). To extend the evidence regarding the role of Merlin in tumors developed in the context of neurofibromatosis, here, we tested whether NF2/Merlin is expressed and exerts similar control on proliferation of colorectal cancer cells and modulates the rate of their apoptosis. We show that NF2/Merlin is also reduced in colorectal tumor cells that do not arise in the context of neurofibromatosis and that induction of Merline expression might be used to control tumor growth.
CRC (Colorectal Cancer) cell lines, HCT116 (ATCC® CCL-247) and SW620 (ATCC® CCL-227) were purchased from ATCC (VA, USA). CRC cells were grown in Dulbecco’s modified Eagle’s medium (DMEM, Invitrogen, Shanghai, China) supplemented with 10% fetal bovine serum (Gibco-ThermoFisher, EU-Approved) and100 U/mL penicillin, and 100 mg/mL streptomycin (Invitrogen, Shanghai, China) at 37°Cin a humidified incubator with 5% CO2 atmosphere. Specimens of colorectal cancers were obtained from Shanghai Municipal Hospital of Traditional Chinese Medicine, Shanghai, after the informed consents from patients or their guardians were obtained.
The microarray data from 104 laser microdissected tumor tissue samples from colorectal cancer patients (GSE21510) were downloaded from GEO DataSets. These data were used to separate those that show low versus a high NF2 expression level by finding the cut-off value of median expression of this gene. Then, we carried out the Gene Set Enrichment Analysis (GSEA) and GSEA enrichment analysis (http://www.broadinstitute.org/gsea/index.jsp) with KEGG_PATHWAY_IN_ CANCER, KEGG_COLORECTAL_CANCER and KEGG_APOPTOSIS gene set.
Western blotting was performed as previously described (12). The primary antibodies used in this study were anti-Merlin (#12888, Cell Signaling Technologies, Danvers, MA, Dilution:1:1000), anti-GAPDH (#5174, Cell Signaling Technologies, Dilution:1:1000), and anti-Cleaved-Caspase3 (#9661, Cell Signaling Technologies, Dilution: 1:1000).
Total RNA was extracted from cultured cells by using Total RNA extracting Kit (Fastagen,Shanghai, China) following the manufacturer’s instruction. For qRT-PCR, mRNA was reverse transcribed to cDNA by using the Reverse Transcription Kit (Takara, Dalian, China). Real-time PCR analyses were conducted with SYBR Premix Ex Taq (Takara, Dalian China). Results were normalized to the expression of GAPDH. The sequence of the primers used in this study are listed in Table 1.
| Gene | Forward primer (5' to 3') | Reverse primer (5' to 3') |
|---|---|---|
| NF2 | AGTCGGAAACTGGCAGATAGC | GGTAGTGTTGTACTGGGCCAAT |
| GAPDH | TGGACTCTGTTCGCTCAGGT | TGCCTCCTTCCGTACCACAT |
NF2-overexpression (OE-NF2) lentivirus vectors and corresponding non-targeting control vector were purchased from (HANBIO, Shanghai, China). Both oligos were cloned into the lentivirus expression plasmid pFH-L which carried a green fluorescence protein (GFP) reporter gene (HANBIO, Shanghai, China). Then, HCT116 and SW620 cells were transfected with OE-NF2 lentivirus vectors and corresponding non-targeting control vector, respectively. After 48 h, the transfection efficiency was confirmed by detecting the GFP-positive cells under a fluorescence microscope.
HCT116 and SW620 cells were seeded into 96-wells plates at a density of 3000 cells/well in six replicate wells. Cell proliferation was measured by cell counting kit-8 reagent (CCK-8, DOJINDO, Japan) according to the manufacturer’s protocol and recorded by fluoroanalyzer (Floskan Ascent) at OD of 450 nm after incubating cells at 37°C for 2 h.
HCT116 and SW620 cells were seeded into 6-wells plates at a density of 500 cells/well in three replicate wells and cultured for 10 days. Colonies were then fixed in 4% paraformaldehyde for 15 min and stained for 15 min with 0.5% crystal violet. Then, the numbers of colonies were photographed, by an Olympus microscope (Olympus Corporation, Tokyo, Japan). Photographs were analyzed using ImageJ program (National Institutes of Health, Maryland).
Immunohistochemical (IHC) was conducted by using an EnVision™ Kit (DAKO, Denmark) according to the manufacturer’s protocol. The primary antibodies used in this experiment included anti-NF2/Merlin (ab88957, Abcam, Cambridge, United Kingdom, Dilution:1:400), anti-Ki67 (#9449, Cell Signaling Technologies, Dilution:1:400). Stained cells photographed and photographs were analyzed using ImageJ program.
Apoptotic cells were identified by using the Alexa Fluor® 647 Annexin V (640911, Biolegend, USA) and Propidium Iodide (PI)/RNase Staining Solution (#4087, Cell Signaling Technologies) according to the manufacturer’s instruction. Apoptotic cells were then detected by using FACS Aria II cell sorter (BD, USA).
The animal studies were performed in accordance with humane treatment of animals and consistent with a protocol approved by the Shanghai Municipal Hospital of Traditional Chinese Medicine, Shanghai. HCT116 cells were washed and re-suspended Matrigel (BD Biosciences) diluted in PBS at a ratio of 1:1. Cell suspensions (100 microliters) containing 5×105 cells were introduced subcutaneously in the left and right flanks of 5 week-old female NOD/SCID mice. Tumor growth was monitored by their bioluminescence using In vivo Imaging System (IVIS), Spectrum (Perkin-Elmer, Waltham, MA) and Living Image Software for IVIS.
Each experiment was carried out in triplicates and all experiments were performed at least three times. Statistical analysis was performed by using SPSS 16.0 software (SPSS, USA). The significant differences between two groups were analyzed by using Student’s t -test; One-way ANOVA was performed to analyze more than two groups. Statistical significance was expressed as, ***: p < 0.001, **: p < 0.01, *: p < 0.05. The survival curves were deciphered by Kaplan-Meier analysis, and P values were calculated by Log-rank (Mantel-Cox) test.
We used TCGA_COLORECTAL CANCER database to decipher whether the expression level of NF2 correlates with prognosis of the disease. The analysis revealed that the prognosis of patients with colorectal cancer is better only when NF2 is highly expressed in the tumors (Figure 1A). However, in the majority of patients with colorectal cancer, immunostainting and Western blotting showed that, as compared with normal colonic tissues, Merlin was significantly reduced in colorectal cancer (Figure 1B-C).
 Figure 1
Figure 1NF2/Merlin is reduced in colorectal cancer tissue compared with normal colonic tissue and its expression correlates with good prognosis. (A) Kaplan–Meier analyses of the correlation between the expression level of NF2 and the prognosis of colorectal cancer (derived from TCGA_COLORECTAL CANCER database). (B) Representative images of Merlin staining in colorectal cancer and normal colonic tissue (scale bar = 50 micrometers). (C) Westerb blotting of Merlin in 5 colorectal cancer and corresponding normal colonic tissues.
To assess the function of NF2, we first constructed a GFP labeled NF2-OE lentivirus. Forty eight hours after transfection of NF2-OE lentivirus into HCT116 and SW620 cells, the transfection efficiency was assessed. qRT-PCR, immunofluorescence and Western blot assays confirmed the overexpression of NF2/Merlin in transfected colorectal cancer cells. The results demonstrated that NF2/Merlin significantly increased in infected HCT116 and SW620 cells compared with control cells (Figure 2A-C). Overexpression of NF2 markedly reduced the viability and the efficiency of colony formation by HCT116 and SW620 cells (Figure 3A-B). Moreover, the HCT116 cells overexpressing NF2 introduced subcutaneously to NOD/SCID mice, grew much slower and weighed significantly less than the control cells (Figure 4A-B). Consistent with the effect of NF2 on proliferation rate of cancer cells, sections of xenografted tumors immunostainted for Ki67 showed a substantial decrease in Ki67 positivity in NF2 overexpressing cells (Figure 4C).
 Figure 2
Figure 2Lentivirus-mediated NF2-overexpression increases the expression of Merlin in HCT116 and SW620 cells. (A) Representative images of GFP-positive HCT116 and SW620 cells and corresponding control cells. (B) qRT-PCR of NF2 in HCT116 and SW620 cells. (C) Western blotting of Merlin was detected in HCT116 and SW620 cells. Data are means ± SD.*, P < 0.05; **, P < 0.01; ***, P < 0.001.
 Figure 3
Figure 3Overexpression of NF2 inhibits colony formation and proliferation of colorectal cancer cells. (A) Representative images of colonies formed in HCT116 and SW620 cells overexpressing NF2 and by control cells. Histogram showing the efficiency of colony formation. (B) CCK-8 assay showing the proliferative ability of in HCT116 and SW620 cells overexpressing NF2 and control cells. Data are means ± SD.*, P < 0.05; **, P < 0.01; ***, P < 0.001.
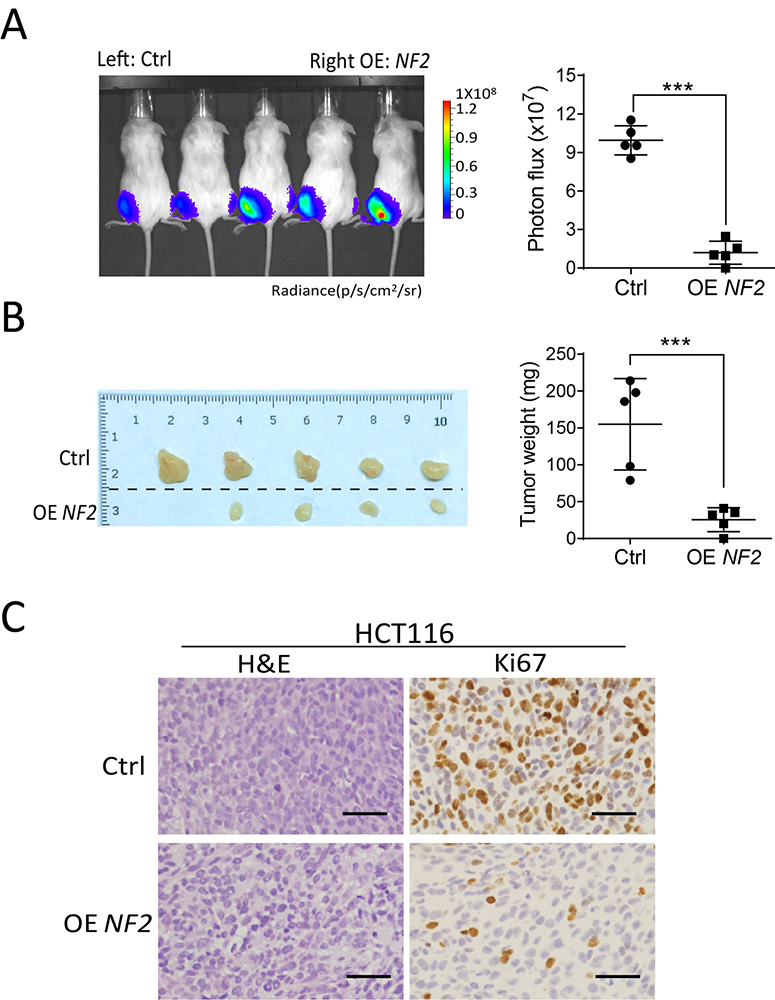 Figure 4
Figure 4Overexpression of NF2 inhibits proliferation of colorectal cancer cells in vivo. (A) In vivo bioluminescent images and the quantification of xenografts formed by in HCT116 and SW620 cells overexpressing NF2 and control. (B) Subcutaneous xenografts formed by in HCT116 and SW620 cells overexpressing NF2 and contros. (C) H&E staining and of Ki67 immunohistochemically detected in xenograft tumors formed by in HCT116 and SW620 cells overexpressing NF2 and controls. Scale bar=50 um. Data are means ± SD. ***, P < 0.001.
The GSE21510 from GEO DataSets of microarrays of 104 laser microdissected tumor tissue samples of colorectal cancer patients showed a direct correlation between the expression of NF2 and apoptosis of colorectal cancer in the context of KEGG_PATHWAY_IN_ CANCER, KEGG_COLORECTAL_CANCER and KEGG_APOPTOSIS gene sets (Figure 5). To validate such findings, the rate of apoptosis in cancer cells overexpressing NF2 was compared with control cells. Annexin V and PI staining followed by flow cytometry showed that cells overexpressing NF2 led to an early apoptosis in HCT116 and SW620 cells compared with control cells (Figure 6A). Cleaved-Caspase3 was measured to confirm the apoptotic mode of cell death. We then conducted Western blot assays to test the expression of Cleaved-Caspase3 and we found Cleaved-Caspase3 significantly increased in NF2-overexpression HCT116 and SW620 cells which was consistent with the flow cytometric results. (Figure 6B) Taken together, these results above illustrated that overexpression of NF2 induced apoptosis in colorectal cancer cells.
 Figure 5
Figure 5GSEA enrichment analysis of gene expression profiles in colorectal cancer. (A) Heat map of expression values for differentially expressed genes from 104 colorectal cancer obtained from the GEO DataSets. (GSE21510) (B) GSE21510 gene set was analyzed with GSEA enrichment plots of KEGG_PATHWAY_IN_CANCER, KEGG_COLORECTAL_CANCER and KEGG_APOPTOSIS gene set .NES:normalized enrichment score.
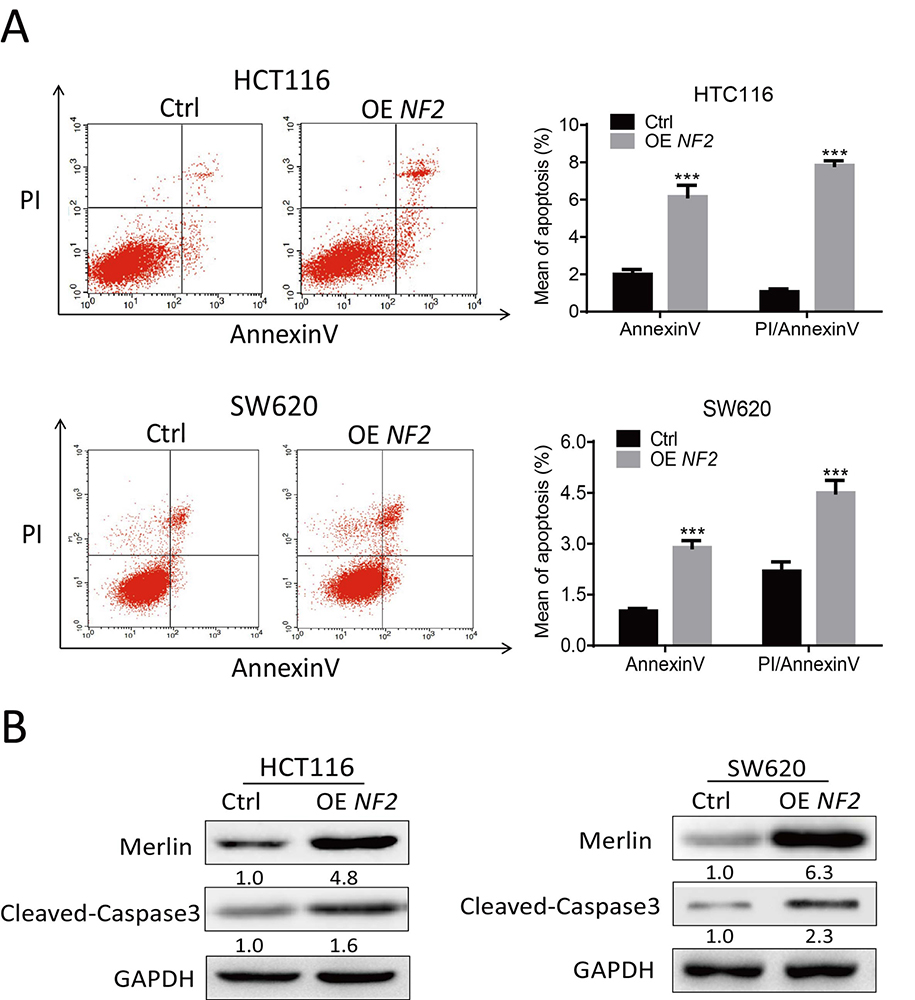 Figure 6
Figure 6Overexpression of NF2 induces apoptosis in colorectal cancer cells. (A) Flow cytometric analysis of apoptotic cells in in by in HCT116 and SW620 cells overexpressing NF2 and control. Histogram shows the percentage of early/late apoptotic cells. (B) Western blotting of Merlin and Cleaved-Caspase3 by in HCT116 and SW620 cells overexpressing NF2 and control. Data are means ± SD.*, P < 0.05; **, P < 0.01; ***, P < 0.001.
Colorectal cancer (CRC) is the third most common cancer diagnosed around the world (13). Though the incidence and the mortality rate of colorectal cancer has decreased due to effective cancer screening measures, it has been projected that there will be 140,250 new cases of colorectal cancer in 2019, with an estimated 50,630 people dying from this disease (2). Thus, there is a need to further understand the complex progression of colorectal cancer in order to identify therapeutic targets that can better control these tumors and improve its prognosis, particularly, when the tumor is diagnosed at last stages of the disease (6).
NF2 is generally regarded as a tumor suppressor gene and is frequently inactivated in some benign tumors of the central nervous system (CNS), including schwannomas and meningiomas (14-15). Merlin, encoded by NF2, has been reported to restrict tumor growth and cause reverion of the malignant phenotype in some solid tumors (16-17). Mutations of the NF2 have also been reported in renal cell carcinoma, thyroid cancer, mesothelioma and melanoma, suggesting of existence of a more general tumor suppressive role for NF2 (18). Here, we showed that, similar to tumors that are developed in the context of inactivation of NF2, NF2/Merlin is reduced in colorectal cancer cells. Furthermore, overexpression of NF2 inhibited colony formation and proliferation of colorectal cancer cells both in vitro and in vivo. Our analysis of data derived from 104 colorectal cancer samples by GSEA enrichment analysis supported of existence of a direct correlation of NF2 expression with apoptosis. Consistent with such data, the overexpression of NF2 induced apoptosis in colorectal cancer cells. These findings are consistent with the roles of NF2 in proliferation and apoptosis which has been reported to act through regulating beta1-AR/PKA/cAMP pathway or as a gatekeeper of the Hippo-signaling pathway which controls both proliferation and apoptosis (19-22). It is possible that NF2/Merlin may have similar functions under normal conditions. For example, in pancreatic beta-cells, loss of NF2/Merlin inhibited apoptosis which was induced by inactivating LATS2 (23). It is also possible that NF2 modulates cell cycle progression and/or apoptosis by acting as a bridge between cytoskeleton and membranous proteins (18). Together, these data show that NF2/Merlin acts to regulate proliferation and apoptosis of colorectal cancers and for this reason, might be used as a therapeutic target in the control of colorectal cancers.
Xingli Wu and Fajiang Mao contributed to this paper equally. This Study Was Supported by Grants From the National Natural Science Foundation of China (NO.81472551/NO.81873279). Medical Guidance Project of Shanghai Science and Technology Commission (NO.19401935200). Clinical Speciality of Shanghai Health Commission(NO.201840325).
CRC
Colorectal cancer
Phosphatidyl inositol 3-kinase/Akt/ mammalian target of rapamycin
Cell Counting Kit 8
moesin-ezrin-radixinlink protein
Immunohistochemical
Neurofibromatosis type 2
green fluorescence protein
Gene Set Enrichment Analysis
The Cancer Genome Atlas
Kyoto Encyclopedia of Genes and Genomes
glyceraldehyde 3-phosphate dehydrogenase
