- Academic Editors
†These authors contributed equally.
Schizophrenia (SCZ) is a complex and heterogeneous neuropsychiatric disorder that lacks objective diagnostic indicators and the pathogenesis remain unclear. Genetic factors may exert a significant impact on the development of the condition. While obtaining brain tissue for biopsy in the course of adjuvant diagnosis of SCZ patients may not be possible, the collection of peripheral blood is more accessible and easier to implement. In recent years, the development and application of RNA sequencing technology has made seeking biomarkers of SCZ becomes more feasible. There is emerging evidence suggesting that certain non-coding RNAs (ncRNA) are distinctly different in the peripheral blood of SCZ patients and healthy controls. Although the mechanisms remain unclear, these aberrantly expressed ncRNAs may be intimately associated with the onset and development of SCZ and may be of great significance for the diagnosis and treatment of SCZ. Therefore, we reviewed the expression of distinct types of ncRNAs that have been found in the peripheral blood of SCZ patients and explored their potential application as diagnostic biomarkers of SCZ. Differentially expressed ncRNAs in the peripheral blood of SCZ patients could not only serve as potential diagnostic biomarkers and therapeutic targets for SCZ but may also have implications for advancing understanding of the molecular mechanisms underlying the development of SCZ and elucidating the complex etiology of SCZ. Early diagnostic biomarkers obtained directly from peripheral blood are of great significance for the timely diagnosis and treatment of SCZ. Our review will enhance the comprehension of molecular mechanisms of SCZ and contribute to the identification of promising ncRNAs in peripheral blood for both diagnosis and therapy of SCZ.
Schizophrenia (SCZ) [1] is a heterogeneous and chronic neuropsychiatric disorder with sophisticated and diverse clinical manifestations. Patients with SCZ suffer from alterations of emotions, cognition, and behaviors. SCZ impacts approximately 1% of the world’s population [2]. The diagnosis of SCZ is dependent on the clinical manifestations and symptoms of the patient, and takes six months or more to diagnose according to DSM-5 diagnostic criteria [3]. Due to the absence of objective diagnostic criteria for SCZ at an early stage, many patients are already in a severe stage by the time they are diagnosed. While medications are available for alleviating and limiting the progression of symptoms, the prognosis for many patients remains unsatisfactory [4]. Hence, it is important to identify objective diagnostic indicators that can be found during the early stages of SCZ.
The pathogenesis of SCZ remains unclear, with genetic, environmental, and social factors all influencing its development to some extent. Studies investigating genes associated with SCZ have made certain developments and with the continuous application of high-throughput technologies, several genetic variants relating to SCZ have been identified [5]. With unbiased properties and high throughput, RNA sequencing technology has been acknowledged as a powerful method for the recognition of biomarkers of SCZ. Since RNA molecules play essential functions in the development and progression of numerous conditions, to date, studies [6, 7] have reported the aberrant expression of the transcriptome in SCZ patients and it has been suggested that differentially expressed RNA molecules can be considered as diagnostic or therapeutic biomarkers for SCZ. RNA in organisms are categorized into two major groups: coding RNAs and non-coding RNAs (ncRNA) [8]. The former refers to mRNA, while the latter includes microRNA (miRNA), long non-coding RNA (lncRNA), and circular RNA (circRNA). The primary function of mRNA is to allow the expression of the genetic information in a transcribed protein. ncRNAs, despite not being capable of coding proteins, can affect the expression of genes through a variety of mechanisms [9]. Statistically [10], the proportion of mRNAs in the transcriptome is typically less than 2%, while the proportion of ncRNAs accounts for more extensive expression in mammalian cells than mRNAs. Recent studies have revealed a number of unique ncRNAs that play vital roles in the maintenance of normal physiological functions and the regulation of various diseases.
An increasing number of studies have demonstrated that ncRNA expression is relatively specific, is abundant in the brain and peripheral nervous system, and can dynamically modulate a wide range of signaling pathways in the context of neurodegenerative lesions through a variety of mechanisms [11]. Consequently, further exploration of the mechanisms through which ncRNAs regulate gene expression is of great significance for the early diagnosis and treatment of neurodegenerative diseases.
Recently, it has been suggested that numerous ncRNAs are also promising diagnostic biomarkers of SCZ [12, 13]. Evidence from various studies indicates that alterations of ncRNAs in SCZ may provide novel insights into the mechanisms underlying its pathobiology. Further exploration of the alterations of ncRNAs in SCZ could therefore be instrumental for gaining further insight into the mechanisms underlying the development and progression of SCZ, as well as for better therapeutic options and early diagnosis of SCZ.
Accurate recognition of differentially expressed genes among particular conditions is necessary for understanding phenotypic variation [14]. RNA sequencing technology has gradually emerged as a necessary tool for analyzing differentially expressed genes at the whole transcriptome level. It has also been used to investigate the complexity of mRNA splicing and the mechanism of ncRNA-regulated gene expression, which has contributed to our understanding of the molecular mechanisms of SCZ. RNA sequencing is a promising tool for investigating disease-related gene expression alterations at the RNA level with high-resolution and low-cost. It has also been used to enhance comprehension of the roles of multiple genes in the causation of certain psychiatric disorders, including SCZ. Quantitative reverse transcription real-time polymerase chain reaction (RT-qPCR) is a convenient and effective method for mRNA detection, with high sensitivity and specificity. It is currently being widely used in the study of SCZ pathogenesis [15].
The utilization of peripheral blood to identify biomarkers of SCZ is more feasible compared to brain tissue samples as it is easily accessible and less invasive. Additionally, there are multiple confounding factors that may affect the expression of genes in post-mortem brain tissue, ranging from cause of death, substance-use history, gender, and age [16]. There are studies that have detected high concordance between the expression of genes in peripheral blood and brain tissue. Liew et al. [17] demonstrated that genes expressed in human peripheral blood share approximately 80% homology with those expressed in brain tissue with the use of microarray hybridization as well as expressed sequence tags. Further, by employing a contrastive gene expression trail extrapolation algorithm, Iturria-Medina et al. [18] identified that approximately 85-90% of the most predictable regulatory pathways identified in brain were also top predictors in the peripheral blood. Therefore, the use of peripheral blood samples to detect biomarkers of SCZ is an optimal alternative method for brain tissue samples.
In the present review, the various types of ncRNAs aberrantly expressed in the peripheral blood of SCZ patients are discussed and the potential value of these ncRNAs are assessed as diagnostic biomarkers and potential therapeutic targets for SCZ. This review will provide further insight and contribute to the translation of ncRNAs as biomarkers in the peripheral blood of SCZ in clinical practice. Fig. 1 illustrates an overview of the material covered in this review.
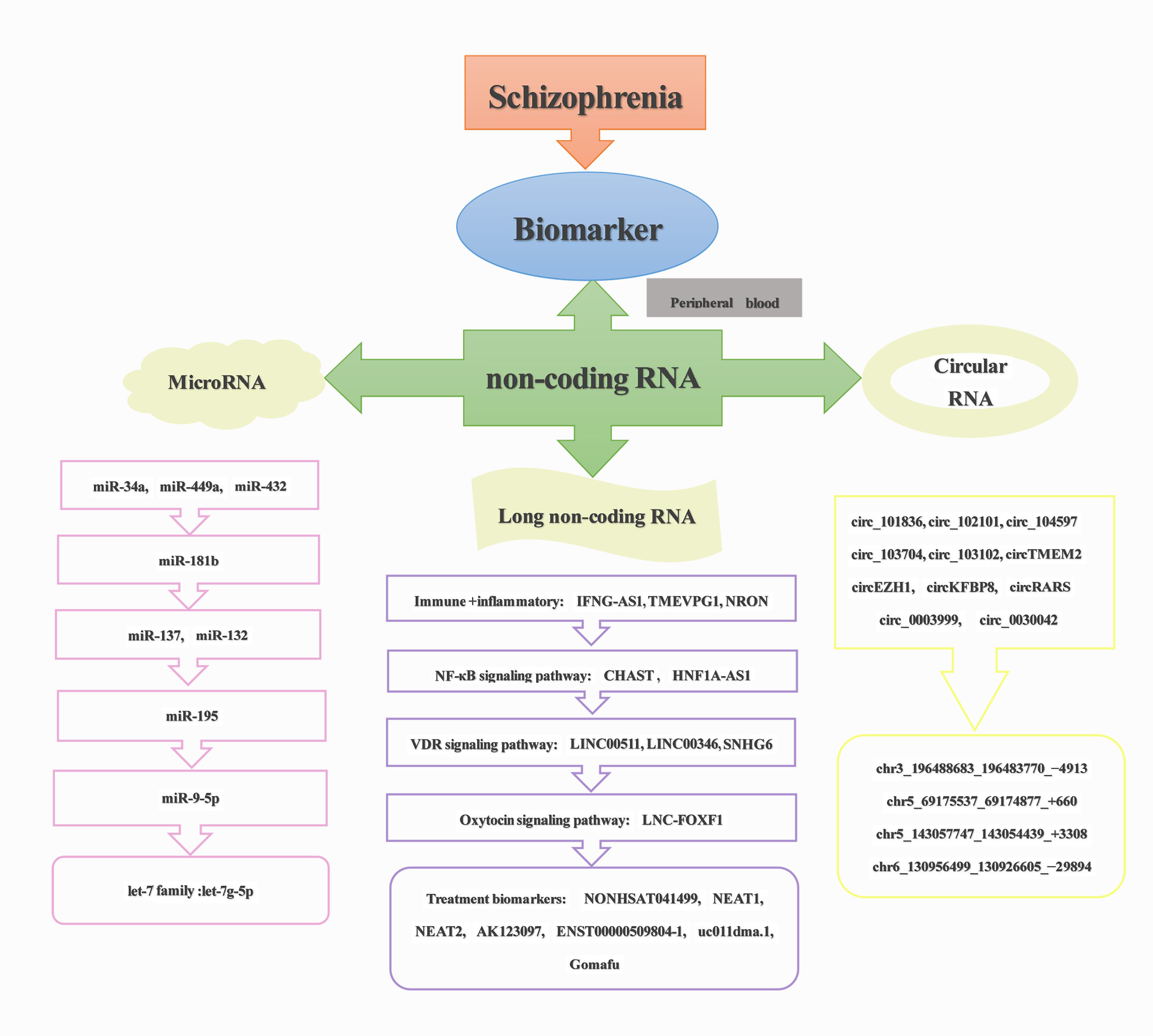 Fig. 1.
Fig. 1.Overview of content reviewed.
miRNAs are an endogenous, minor non-coding RNA that primarily engage in the modulation of gene expression in post-transcriptional processes by disturbing transcription or translation. They play an essential role in the regulatory mechanisms of a variety of biological processes, including time of development, cell proliferation and differentiation, and apoptosis [19]. Recently, an increasing number of studies have identified that dysregulation of the expression of miRNAs is intimately associated with multiple diseases. Additionally, there are hundreds of miRNAs that have been shown to be aberrantly expressed in diseases based on the analysis of global gene expression profiles [20].
It has been demonstrated that miRNAs are abundantly expressed in the nervous system, where they can induce abnormalities on a range of gene expression and functioning pathways. These abnormalities are of significance for numerous neuropsychiatric disorders, including SCZ, as they cause the dysfunction of multiple pathways [21]. Studies [22, 23] have explored the role of miRNAs on both brain function and interneuron development. miRNAs coordinate the regulation of translation, stability, splicing, and localization of related mRNAs, which could contribute to further understanding of the pathogenesis of SCZ. Recently, as studies regarding the role of miRNAs in SCZ are becoming widespread, miRNAs have been identified as critical regulators of gene expression and are promising candidates for biomarkers of SCZ. Research has revealed that expression of miRNAs in peripheral blood alters in response to changes of the body’s physiological or pathological conditions [24]. Accordingly, aberrantly expressed miRNAs in peripheral blood of SCZ patients may be of great significance for the diagnosis of SCZ. Table 1 (Ref. [15, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47]) presents miRNAs that have the potential to serve as biomarkers in the peripheral blood of SCZ.
| miRNA | Method | Sample | Direction | Sample size | Year | Study |
| miR-181b | RT-qPCR | serum | up | 115:40 | 2011 | Shi et al. [31] |
| miR-219-2-3p | up | |||||
| miR-1308 | up | |||||
| let-7g | up | |||||
| miR-195 | down | |||||
| miR-34a | RT-qPCR | PBMC | up | 30:30 | 2011 | Lai et al. [25] |
| miR-449a | - | |||||
| miR-564 | - | |||||
| miR-432 | - | |||||
| miR-548d | - | |||||
| miR-572 | - | |||||
| miR-652 | - | |||||
| miR-134 | RT-qPCR | PBMC | down | 112:76 | 2012 | Gardiner et al. [32] |
| miR-128 | - | |||||
| miR-181b | - | |||||
| miR-31 | down | |||||
| miR-431 | down | |||||
| miR-433 | down | |||||
| miR-107 | down | |||||
| miR-99b | down | |||||
| miR-487b | down | |||||
| miR-130b | RNA-seq | plasma | up | 164:187 | 2015 | Wei et al. [41] |
| miR-193a-3p | RT-qPCR | up | 400:312 | |||
| miR-132 | RNA-seq | PBMC | down | 105:130 | 2015 | Yu et al. [35] |
| miR-30e | RT-qPCR | plasma | up | 61:62 | 2015 | Sun et al. [29] |
| miR-181b | up | |||||
| miR-34a | up | |||||
| miR-346 | up | |||||
| miR-7 | up | |||||
| let-7g-5p | RNA-seq | blood | down | 6:10:8 | 2015 | Rizos et al. [39] |
| miR-98-5p | down | |||||
| miR-183-5p | down | |||||
| miR-132 | RT-qPCR | plasma PBMC | up | 25:13 | 2015 | Sun et al. [28] |
| miR-195 | up | |||||
| miR-7 | up | |||||
| miR-212 | up | |||||
| miR-34a | up | |||||
| miR-30e | up | |||||
| miR-137 | RT-qPCR | peripheral blood | up | 44:44 | 2016 | Wu et al. [34] |
| miR9-5p | RT-qPCR | peripheral blood | up | 16:16 | 2016 | Camkurt et al. [37] |
| miR29a-3p | up | |||||
| miR106b-5p | up | |||||
| miR125a-3p | up | |||||
| miR125b-3p | up | |||||
| miR-34a | RT-qPCR | peripheral blood | up | 25:27 | 2016 | Lai et al. [26] |
| miR-449a | up | |||||
| miR-564 | up | |||||
| miR-432 | up | |||||
| miR-548d | up | |||||
| miR-572 | up | |||||
| miR-652 | up | |||||
| miR-22-3p | RNA-seq | peripheral blood | up | 10:10 | 2018 | Ma et al. [15] |
| miR-92a-3p | RT-qPCR | up | 44:44 | |||
| miR-137 | up | |||||
| miR-34a-5p | RT-qPCR | serum | up | 40:40 | 2019 | He et al. [30] |
| miR-432-5p | down | |||||
| miR-449a | up | |||||
| miR-223 | RNA-seq | plasma | up | 17:17 | 2019 | Zhao et al. [42] |
| RT-qPCR | 21:21 | |||||
| miR-1271-5p | RNA-seq | PBMC | down | 36:15 | 2019 | Geaghan et al. [40] |
| miR-221-5p | RT-qPCR | down | 17 | |||
| let-7 | down | |||||
| miR-320a-3p | RNA-seq | serum | down | 50:60 | 2019 | Wang et al. [43] |
| miR-320b | ||||||
| miR-19b | RT-qPCR | peripheral blood | up | 22:19 | 2020 | Horai et al. [27] |
| miR-373-5p | GSE54578 | peripheral blood | - | 15:15 | 2020 | Pala et al. [44] |
| miR-199a-3p | - | |||||
| miR-148b-3p | RT-qPCR | peripheral blood | up | 44:44 | 2020 | Wu et al. [33] |
| miR-218-5p | RNA-seq | PBMC | up | 34:31 | 2020 | You et al. [45] |
| miR-1262 | RT-qPCR | up | 6:6 | |||
| miR-195 | RNA-seq | peripheral blood | up | 118:47 | 2021 | Pan et al. [36] |
| miR-9-5p | RNA-seq | peripheral blood | down | 15:15 | 2022 | Jin et al. [38] |
| miR-4467 | RT-qPCR | up | 35:60 | |||
| miR-574-5P | GSE54914 | peripheral blood | up | 18:12 | 2022 | Davarinejad et al. [46] |
| miR-1827 | up | |||||
| miR-4429 | up | |||||
| miR-185-5p | qPCR | peripheral blood | down | 50:50 | 2022 | Sabaie et al. [47] |
RT-qPCR, Quantitative reverse transcription real-time polymerase chain reaction; PBMC, peripheral blood mononuclear cells.
In a 2011 study by Lai et al. [25], a comparative analysis of miRNA expression in peripheral blood mononuclear leukocytes was performed with Taqman low-density arrays. Seven miRNAs (miR-34a, miR-449a, miR-564, miR-432, miR-548d, miR-572, and miR-652) were differentially expressed between SCZ patients and healthy controls (CTL), with the most significant difference being in expression of miR-34a. A support vector machine was used to assess the predictive accuracy of the 7-miRNA signature in differentiating SCZ from CTL. The area under the receiver operating characteristic (ROC) curve of its diagnostic prediction model was 93% and the area under the curve (AUC) in the test set was 85%, with good diagnostic performance. Diagnostic prediction models for SCZ serve as an essential approach to distinguish SCZ cases from CTL and to predict whether SCZ will occur. It has become widely accepted to employ machine learning methods to construct diagnostic prediction models, where combining different variables for SCZ prediction may improve the accuracy of prediction. With the help of diagnostic prediction models, clinicians and SCZ patients can make better joint decisions, researchers can screen suitable SCZ study subjects more accurately, and governments can allocate medical resources accordingly.
Their subsequent study in 2016 [26] revealed that hospitalization did not influence expression of these seven miRNAs above that in peripheral blood, leading them to suggest that miRNAs may as trait-dependent markers. Moreover, their study revealed a corresponding correlation between the expression levels of miR-34a in the blood and in cortical Brodmann area 46. In a study conducted by Horai et al. [27] in 2020, miR-19b, which is highly expressed in neural progenitor cells in the hippocampus of SCZ patients, also had increased expression in peripheral blood. The high expression of miR-19b likely increases the vulnerability of SCZ by attenuating the proliferation of neural progenitor cells in the hippocampus. It is possible that these studies may provide further support for the use of miRNAs in peripheral blood as diagnostic biomarkers of SCZ. Sun et al. [28] detected that the expression of miR-132, miR-195, miR-30e, and miR-7 were significantly upregulated in blood plasma and miR-212, miR-34a, and miR-30e were upregulated in peripheral blood mononuclear cells (PBMC). Differences in the tissue microenvironment in which miRNAs function may explain the differences in their expression levels in different tissues. Interestingly, the expression of miR-30e in both plasma and PBMC was significantly different in patients with SCZ compared to CTL. Further logistic regression analysis demonstrated that miR-30e in plasma has greater diagnostic value for SCZ, which further suggests that miR-30e may be considered as a plasma biomarker for the diagnoses of SCZ. Meanwhile, in their other study [29], the combination of miR-30e, miR-181b, miR-34a, miR-346, and miR-7 in plasma was found to be a potential biomarker for SCZ diagnosis. He et al. [30] detected that miR-34a-5p, miR-432-5p, and miR-449a were aberrantly expressed in the serum of SCZ patients. Accordingly, it could be suggested that miR-34a, miR-449a, and miR-432 are performing relatively important roles in the pathogenesis of SCZ and that it is feasible to look for miRNAs in peripheral blood that can reflect aberrant alterations in brain tissue.
Shi et al. [31] detected that miR-181b, miR-219-2-3p, miR-195, miR-1308, and let-7g could act as potential diagnostic biomarkers of SCZ. In an attempt to explore miRNAs relevant to SCZ in non-neural tissues, Gardiner et al. [32] performed an analysis of miRNA expression profiles and discovered that certain miRNAs that are differentially expressed in the brain are also differentially expressed in peripheral blood, such as miR-134, miR-128, and miR-181b. Additionally, from their study, seven miRNAs (miR-31, miR-431, miR-433, miR-107, miR-134, miR-99b, miR-487b) were identified as being differentially expressed in the peripheral blood of patients with SCZ. Aberrant expression of miR-181b in the plasma of SCZ patients was also identified in a study by Sun et al. [29]. Hence, miR-181b may also be potentially valuable for the diagnosis of SCZ.
A study by Wu et al. [33] revealed upregulation of the expression of miR-148b-3p in the peripheral blood of patients with SCZ during their first-episode and predicted that ZNF804A may be the target gene where miR-148b-3p exerts its effect in the pathological mechanism of SCZ. In another study conducted by Wu et al. [34] 2016, the expression of miR-137 was upregulated in the peripheral blood of SCZ patients compared to CTL. The diagnostic ROC curve for distinguishing SCZ from CTL with the utilization of miR-137 showed an area under the curve(AUC) value of 0.795. Furthermore, this study also revealed that miR-137 may target genetic variants impacting the RNA binding site of the EFNB2 gene, causing its down-regulation. Accordingly, they suggested that miR-137 may be a meaningful biomarker for SCZ. Two years later, Ma et al. [15]explored miRNAs in peripheral blood that are potential diagnostic biomarkers for SCZ with second-generation sequencing in combination with RT-qPCR and detected that the combination of three miRNAs, miR-137, miR-22-3p, and miR-92a-3p, may be meaningful diagnostic biomarkers for SCZ. Additionally, a study by Yu et al. [35] identified miR-132 as a promising biomarker in peripheral blood for differentiating SCZ from CTL. Sun et al. [28] have also demonstrated the value of miR-132 in the diagnosis of SCZ. As such, miR-137 and miR-132, may serve as potential diagnostic biomarkers of SCZ that are intimately associated with regulating the expression of SCZ-related mRNAs.
As previously described, Shi et al. [31] identified nine miRNAs, including miR-195, as candidate biomarkers for the diagnosis of SCZ back in 2011. Another study by Sun et al. [28] subsequently revealed significant upregulation of miR-195 expression in the plasma of SCZ. In 2021, Pan et al. [36] also identified significantly elevated levels of miR-195 in peripheral blood of SCZ patients. Additionally, their study demonstrated that in SCZ patients, high expression of miR-195 was associated with a decrease in levels of brain-derived neurotrophic factor (BDNF), where low levels of BDNF protein is associated with cognitive dysfunction. Consequently, the upregulation of miR-195 in peripheral blood likely influences cognitive function in SCZ by modulating the expression of BDNF.
In 2016, Camkurt et al. [37] detected five miRNAs, miR9-5p, miR29a-3p, miR106b-5p, miR125a-3p, and miR125b-3p, significantly upregulated in SCZ. Interestingly, in 2022, Jin et al. [38] revealed that the expression of miR-4467 was significantly upregulated in SCZ, while miR-9-5p expression was significantly down-regulated. The predicted AUC value was 0.709 by combining miR-4467 and miR-9-5p for the diagnosis of SCZ. Notably, miR-9-5p expression appeared to be in opposite directions in different studies although they were all aberrantly expressed; therefore, further exploration is necessary to clarify the diagnostic value of miR-9-5p in peripheral blood for SCZ.
As mentioned previously, a study by Shi et al. [31] identified let-7g as a potential diagnostic biomarker in the serum of SCZ patients. It was also detected by Rizos et al. [39], that the expression of let-7g-5p, miR-98-5p, and miR-183-5p were significantly down-regulated in the blood of patients with cancer and SCZ. Additionally, Geaghan et al. [40], revealed that miRNAs of the let-7 family, miR-1271-5p, and miR-221-5p performed essential functions in regulating the expression of genes in immune cells in the peripheral blood of SCZ patients. It has been determined that the let-7 family of miRNAs are tumor suppressors that modulate the response of macrophages as well as the production of B-cell antibodies, both of which play essential roles in regulating the immune system [48]. Hence, further exploration of the aberrantly expressed let-7 family miRNAs in peripheral blood may be of significance for understanding the pathogenesis of SCZ.
In addition, several studies have detected other miRNAs in peripheral blood that might serve as diagnostic markers for SCZ. In 2015, Wei et al. [41] conducted validation of eight miRNAs (miR-130a, miR-130b, miR-122, miR-193a-3p, miR-193b, miR-502-3p, miR-652, and miR-886-5p) differentially expressed between SCZ patients and CTL by utilizing RT-qPCR. They determined that two of these miRNAs (miR-130b and miR-193a-3p) may have significance for the diagnosis of SCZ. Zhao et al. [42] identified that the expression of miR-223 in plasma of SCZ patients was upregulated both during the first episode and its later stages compared to CTL. This abnormal expression of miR-223 may affect the expression levels of its targeted genes involved in cell migration. An investigation by Wang et al. [43] revealed that the expression of miR-320a-3p and miR-320b was significantly downregulated in the serum of SCZ patients. Pala et al. [44] identified miR-373-5p and miR-199a-3p as potential biomarkers for SCZ diagnosis by analyzing the microRNA expression profile GSE54578. You et al. [45] discovered that the expression of miR-218-5p and miR-1262 were notably upregulated in PBMC of treatment-resistant SCZ patients. The target genes of these two miRNAs, CBX5, NF165, and CACUL1, are intimately associated with brain function and the nervous system. As a result, they proposed that miR-218-5p and miR-1262 might be biomarkers for early diagnosis of treatment-resistant SCZ. Davarinejad et al. [46] identified miR-574-5P, miR-4429, and miR-1827 as potential blood diagnostic biomarkers for SCZ. Sabaie et al. [47] identified down-regulation of miR-185-5p in the peripheral blood in SCZ patients, which could be relatively well differentiated from that of CTLs (AUC = 0.722), but larger samples for validation are still necessary.
It is evident that there have been multiple studies which have detected miRNAs in peripheral blood that can serve as diagnostic biomarkers for SCZ. Interestingly, some of the aberrantly expressed miRNAs detected in peripheral blood are also present at abnormal levels in brain tissue of SCZ patients. Therefore, the use of peripheral blood is promising as a means for the detection of diagnostic miRNAs in SCZ. Additionally, it is noteworthy that several studies have found the same miRNAs in peripheral blood to be of diagnostic value for SCZ, such as miR-34a, miR-181b, miR-137, miR-132, miR-195, miR9-5p, miR-432, miR-7, miR-30e, miR-548d, miR-432, miR-449a, and let-7. Thus, these miRNAs are promising as reliable diagnostic biomarkers for SCZ. However, the accuracy of diagnostic prediction of SCZ using a single miRNA may be low and an attempt should be made to combine multiple miRNAs that are abnormally expressed in peripheral blood for the prediction of SCZ, thus further improving the accuracy of diagnostic results. Consequently, miRNAs differentially expressed in the peripheral blood of SCZ patients may be novel biomarkers that provide non-invasive and accurate diagnosis of SCZ.
lncRNAs are RNA transcripts that encode proteins less than 200 nucleotides in length. They play vital roles in an array of biological functions and cellular processes, such as metabolism, cell differentiation, cell cycle, and have been implicated in multiple diseases [49]. Qian et al. [50] investigated the mechanism and functional role of lncRNAs in regulating RNA metabolism and expression of genes by using high-throughput sequencing, bioinformatics, and automated capillary approaches. They identified that lncRNAs are essential modulators of the function and expression of almost all genes.
Previous studies have revealed the expression of numerous lncRNAs in the brain that are predominantly engaged in the development and function of the nervous system [51]. It is widely accepted that SCZ is a caused by multiple factors with sophisticated genetic constituents. Rusconi et al. [52] demonstrated that the characteristic mutations of numerous psychiatric disorders, including SCZ, occurred in non-coding parts of genes. There has been an accumulation of studies demonstrating the relevance of lncRNAs to the pathogenesis of SCZ [53, 54]. It has been suggested that most genes expressed in the brain and peripheral blood share common regulation pathways. For instance, Rao et al. [55] demonstrated that LINC00461, which is downregulated in the hippocampus of SCZ patients, was also downregulated in peripheral blood. Hence, lncRNAs aberrantly expressed in the brain of SCZ patients may also have abnormal expression in the peripheral blood and it is therefore feasible to detect lncRNAs aberrantly expressed in peripheral blood of SCZ patients [54]. Table 2 (Ref. [8, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72]) presents potential lncRNAs biomarkers in peripheral blood of SCZ patients.
| lncRNA | Method | Sample | Direction | Sample size | Year | Study |
| AC079587.1 | RNA-seq | peripheral blood | down | 19:18 | 2015 | Ren et al. [56] |
| CTD-2194F4.2 | down | |||||
| RP11-146N23.1 | down | |||||
| RP11-383G10.3 | down | |||||
| RP11-698L23.1 | down | |||||
| RP11-167J8.1 | down | |||||
| RP4-803A2.2 | up | |||||
| GAPDHP37 | up | |||||
| RP11-93K22.14 | up | |||||
| CR602933 | up | |||||
| AC093716.1 | up | |||||
| COX6B1P1 | up | |||||
| AC060764.1 | up | |||||
| RP4-559A3.5 | up | |||||
| AC104389.32 | up | |||||
| AC009852.1 | up | |||||
| LOC644246 | up | |||||
| TTC39C | up | |||||
| ATP5G2P1 | up | |||||
| RP11-379B18.3 | up | |||||
| RP11-144C15.1 | up | |||||
| POLR2LP | up | |||||
| RP1-197O17.2 | up | |||||
| PPIHP1 | up | |||||
| AP004242.2 | up | |||||
| DA376252 | GWAS | plasma | - | - | 2016 | Chen et al. [71] |
| BX089737 | ||||||
| LOC101927273 | ||||||
| LINC01029 | ||||||
| LOC101928622 | ||||||
| HY157071 | ||||||
| DA902558 | ||||||
| NONHSAT089447 | RNA-seq | PBMC | down | 3:3 | 2016 | Chen et al. [60] |
| NONHSAT041499 | qRT-PCR | down | 106:48 | |||
| NONHSAT021545 | ||||||
| IFNG-AS1 | RNA-seq | peripheral blood | down | 27:32 | 2017 | Ghafelehbashi et al. [57] |
| Neat1 | RT-qPCR | peripheral blood | down | 18:9 | 2018 | Li et al. [67] |
| Neat2 | down | |||||
| MEG3 | - | |||||
| MIAT | - | |||||
| TMEVPG1 | RT-qPCR | blood | up | 17:16 | 2018 | Melbourne et al. [58] |
| NRON | up | |||||
| MEG3 | RNA-seq | PBMC | up | 86:44 | 2018 | Sudhalkar et al. [63] |
| PINT | down | |||||
| GAS5 | down | |||||
| Gomafu | RNA-seq | PBMC | up | 35:49 | 2018 | Liu et al. [66] |
| HOXA-AS2 | RNA-seq | peripheral blood | up | 60:60 | 2019 | Fallah et al. [62] |
| Linc-ROR | up | |||||
| MEG3 | up | |||||
| SPRY4-IT1 | up | |||||
| UCA1 | up | |||||
| FAS-AS1 | RT-qPCR | peripheral blood | down | 50:50 | 2019 | Safari et al. [64] |
| PVT1 | down | |||||
| TUG1 | down | |||||
| THRIL | up | |||||
| GAS5 | - | |||||
| NEAT1 | - | |||||
| OIP5-AS1 | - | |||||
| NONHSAT089447 | RNA-seq | PBMC | up | 40:40 | 2019 | Chen et al. [61] |
| PACER | RT-qPCR | peripheral blood | down | 50:50 | 2020 | Safa et al. [68] |
| CHAST | up | |||||
| CEBPA | up | |||||
| H19 | up | |||||
| HNF1A-AS1 | up | |||||
| ASHG19A3A011462 | RNA-seq | peripheral blood | up | 14:18 | 2020 | Ren et al. [72] |
| ASHG19A3A026335 | up | |||||
| ASHG19A3A049471 | down | |||||
| ASHG19A3A049556 | down | |||||
| ASHG19A3A044112 | down | |||||
| BDNF-AS | RNA-seq | peripheral blood | - | 50:50 | 2021 | Badrlou et al. [8] |
| MIR137HG | - | |||||
| MIAT | - | |||||
| PNKY | up | |||||
| Gomafu | RNA-seq | plasma | up | 48:49 | 2021 | Jia et al. [65] |
| AK096174 | down | |||||
| AK123097 | down | |||||
| DB340248 | down | |||||
| uc011dma.1 | up | |||||
| ENST00000509804-1 | down | |||||
| ENST00000509804-2 | down | |||||
| AC006129.1 | RNA-seq | peripheral blood | up | 151:134 | 2021 | Ni et al. [59] |
| SNHG6 | RNA-seq | venous blood | up | 50:50 | 2022 | Ghafouri-Fard et al. [69] |
| LINC00346 | up | |||||
| LINC00511 | up | |||||
| LINC00461 | RT-qPCR | peripheral blood | down | 32:48 | 2022 | Rao et al. [55] |
| LNC-FOXF1 | RNA-seq | peripheral blood | up | 60:60 | 2022 | Eghtedarian et al. [70] |
GWAS, genome-wide association study.
To investigate the potential regulatory effects of lncRNAs on the expression of genes and the pathogenesis in SCZ, Ren et al. [56] performed a Weighted Gene Co-expression Network Analysis (WGCNA) which identified two modules that are relevant to SCZ, the blue and brown modules. It is possible that these two modules are engaged in the pathogenesis of SCZ by causing dysfunction of mitochondria through the regulation of their targeted mRNAs. In 2020, to explore lncRNAs associated with the prodromal stage of SCZ, which is known to be ultra-high risk for psychosis, they conducted an additional WGCNA and detected that the expression of ASHG19A3A011462 and ASHG19A3A026335 was upregulated, while the expression of ASHG19A3A049471, ASHG19A3A044112, and ASHG19A3A049556 was downregulated. Subsequently, by analyzing the function of mRNAs with corresponding expression patterns with these five lncRNAs, the study observed that these mRNAs appeared to be notably abundant in functional pathways associated with immunity and inflammation. Consequently, they suggested that lncRNAs may participate in immune and inflammatory abnormalities in the pathogenesis of ultra-high-risk psychosis, which may have great implications for further exploration of the pathogenesis of SCZ.
To explore the regulatory role of IFNG-AS1 on the gene locus of
IFNG in SCZ, Ghafelehbashi et al. [57] compared the expression
levels of IFNG-AS1, IFNG, and IL-1B in the blood cells
of SCZ patients and CTL. They detected that the expression level of
IFNG-AS1 was significantly downregulated in the blood cells of SCZ
patients when compared to CTLs and there was a positive correlation with
expression levels of IFNG and IL-1B. IFNG and
IL-1B are known to be of significance in the regulation of inflammation,
so it was hypothesized in this study that IFNG-AS1 is intimately
involved in inflammation and immunity, and may be one of the essential
inflammatory regulators in the pathogenesis of SCZ. Additionally, a study by
Melbourne et al. [58] demonstrated a positive correlation between the
expression of lncRNA TMEVPG1, NRON, and the expression of
IL-6 and IFN-
By analyzing lncRNAs microarray data from SCZ patients and CTL, Chen et al. [60] determined that NONHSAT089447, NONHSAT021545, and NONHSAT041499 were significantly upregulated in peripheral blood of SCZ. These three lncRNAs were co-expressed with various mRNAs involved in regulating biological processes such as memory, cognition, neuronal apoptosis, and Ras protein signaling. Additionally, the down-regulated expression of NONHSAT041499 was correlated with the alleviation of positive symptoms in SCZ patients following drug treatment, indicating that NONHSAT041499 may be a potential prognostic factor for the outcome of SCZ treatment. Subsequently, Chen et al. [61] carried out a study to further examine the associations between these lncRNAs and SCZ. They found that the expression of NONHSAT089447 was higher than NONHSAT041499 in SCZ patients and showed either activation or regulation of the dopamine signaling pathway. The expression level of NONHSAT089447 may regulate downstream dopamine signaling, thus affecting the occurrence and development of SCZ. Consequently, NONHSAT089447 may be a potentially valuable diagnostic biomarker of SCZ.
To investigate the relations between NEAT1, NEAT2, MEG3 and MIAT, and SCZ, Li et al. [67] evaluated the levels of these lncRNAs in peripheral blood. What they identified was that the expression levels of NEAT1 and NEAT2 were markedly downregulated but were elevated in SCZ after treatment. However, MIAT and MEG3 were at lower expression levels. Furthermore, they investigated the distribution of these lncRNAs in the body and identified that MIAT was abundantly expressed in the brain, while NEAT1, NEAT2, and MEG3 were abundantly expressed in both the brain and peripheral tissues. Fallah et al. [62] identified differences in the expression levels of HOXA-AS2, Linc-ROR, MEG3, SPRY4-IT1, and UCA1 between female SCZ patients and CTL. However, there were no differences in the expression levels of lncRNAs between male SCZ patients and CTL, suggesting a potential sex difference. Moreover, they suggested that MEG3 may affect SCZ by impacting the glutamatergic, dopaminergic, and GABA ergic pathways. Sudhalkar et al. [63] revealed that the expression levels of MEG3 were upregulated in PBMCs of SCZ patients, while that of PITT and GAS5 were downregulated, and the ROC curve analysis showed strong diagnostic predictive capability of MEG3 for SCZ. Furthermore, this study also detected an association between MEG3 and PITT. This may be attributed to the fact that MEG3 is a lncRNA which could regulate the binding specificity of transcription factor P53. P53 exerts a regulatory and activating effect on the expression of PINT, whereas GAS5 is involved in the mechanism of the development of SCZ by serving as the decoy nucleotide-binding site for the glucocorticoid receptor. Subsequently, in 2019, Safari et al. [64] demonstrated that the expression levels of FAS-AS1, PVT1, and TUG1 were down-regulated and THRIL expression was up-regulated in the peripheral blood of SCZ patients compared to CTL. While the expression of GAS5, NEAT1, and OIP5-AS1 did not differ significantly between SCZ and CTL, there were notable differences in the expression of GAS5, NEAT1, and OIP5-AS1 in female subjects. There was 86.96% specificity and 100% sensitivity of GAS5 for the prediction of the diagnosis of SCZ in females and the level of GAS5 exhibited negative correlation with the other six lncRNAs. Thus, it is speculated that there may be sex differences in certain lncRNAs in peripheral blood of SCZ patients.
Jia et al. [65] attempted to explore prospective diagnostic biomarkers for SCZ and detected that the expression levels of Gomafu and uc011dma.1 were markedly upregulated in plasma of those with SCZ. Meanwhile, AK096174, AK123097, DB340248, ENST00000509804-1, and ENST00000509804-2 were downregulated. The combination of seven lncRNAs for the diagnostic prediction of SCZ showed excellent predictive performance with the area under the ROC curve reaching 0.925. Additionally, the expression of AK123097 and ENST00000509804-1 in plasma were upregulated with the amelioration of patients’ symptoms after drug treatment, while that of uc011dma.1 was greatly reduced. Accordingly, AK123097, uc011dma.1, and ENST00000509804-1 may be promising therapeutic targets for SCZ.
Badrlou et al. [8] evaluated the diagnostic performance of four BDNF-related lncRNAs for SCZ with ROC curves. The diagnostic capabilities of MIAT, MIR137HG, BDNF-AS, and BDNF were 68%, 67%, 72%, and 71%, respectively. A study by Liu et al. [66] explored the expression levels of Gomafu in PBMC of SCZ patients before and after drug treatment. Gomafu was found to be significantly higher in PBMC of untreated SCZ patients compared to CTL. Subsequently, the expression level of Gomafu in PBMC of SCZ patients was markedly increased after 12 weeks of drug treatment. It is well-known that Gomafu, also named MIAT, is located on 22q12.1 and is intimately associated with SCZ. Although MIAT was principally distributed in the brain [67], it is also expressed in peripheral blood and several studies have detected upregulation of its expression level in peripheral blood of SCZ. Hence, Gomafu may be a promising diagnostic biomarker for peripheral blood of SCZ.
The NF-
The oxytocin-related signaling pathway can interact with dopaminergic signaling, which is linked with the pathophysiology of SCZ. There are certain lncRNAs that mediate the activity of the oxytocin system and thus exert influence on the development of SCZ. Eghtedarian et al. [70] assessed the aberrant expression of nine oxytocin-related lncRNAs as well as mRNAs in the venous blood of SCZ patients, where the expression of LNC-FOXF1 was significantly upregulated. LNC-FOXF1 is an oxytocin system related lncRNA. There is also an association between LNC-FOXF1 and the immune response. There are common genetic mechanisms between SCZ and nicotine dependence. Chen et al. [71] identified multiple lncRNAs associated with these genetic mechanisms, including DA376252, BX089737, LOC101927273, LINC01029, LOC101928622, HY157071, and DA902558.
In summary, through comprehensive review and evaluation of the existing studies
on relevant lncRNAs in peripheral blood of SCZ, it can be concluded that these
lncRNAs may exert a certain influence on the pathogenesis of SCZ. They may do so
through a variety of mechanisms, such as regulating the expression of genes
associated with inflammatory cytokines and the function of signaling pathways
that influence glutamatergic and dopaminergic signaling pathways. Additionally,
there are studies that reveal the abnormal expression of lncRNAs in SCZ, such as
NONHSAT089447, NEAT1, MEG3, GAS5, and
Gomafu. Furthermore, a number of lncRNAs have been closely associated
with immune and inflammatory responses in the SCZ, such as IFNG-AS1,
TMEVPG1, and NRON. Moreover, certain signaling pathway-related
lncRNAs have corresponding effects on the pathogenesis of SCZ, such as
NF-
CircRNAs are a type of single-stranded, ncRNA molecule that perform diverse functions in cells. They are generated during retrosplicing of the precursor mRNA and are covalently enclosed with highly specific expression in the cells of numerous organisms [74]. For instance, circRNAs can regulate gene expression and chromatin modification, moderate transcription and splicing, act as molecular sponges by repressing the interaction of miRNA with mRNA or proteins, and serve as templates for translation in several biological and pathophysiological contexts [75]. Studies have also established that there are certain linkages between circRNAs interfering with cellular processes and signaling pathways, modulating immune responses, and the biological mechanisms of multiple conditions, such as tumors [76] and psychiatric disorders [77].
With the continued application of high-throughput sequencing technologies, several studies [78, 79] have recently been conducted to investigate the biologic functions of circRNAs in brain and peripheral nervous system. It has been shown that circRNAs are abundantly expressed in the nervous system, are remarkably hyperactive in synapses of neurons, and that the expression of certain genes in the nervous system are regulated by the expression levels of circRNAs. circRNAs serve critical roles in the maintenance of proper function of the brain and preventing the progression of neurological diseases. Accordingly, dysregulation of circRNA expression may be associated with neurological damage or neurodegenerative diseases [80]. In a study [81] that sequenced the RNA molecules in postmortem brain tissue from SCZ and CTL, it was revealed that the expression of numerous circRNAs was decreased in the brain of SCZ patients compared to CTLs. The stability of these circRNAs was also diminished, suggesting these circRNAs might be playing a vital role in the etiology of SCZ by regulating the expression of miRNAs or the translation of proteins. Table 3 (Ref. [13, 82, 83, 84]) lists circRNAs with potential as biomarkers in peripheral blood of SCZ.
| circRNA | Method | Sample | Direction | Sample size | Year | Study |
| circ_101836 | RNA-seq | PBMC | down | 9:9 | 2019 | Yao et al. [82] |
| circ_102101 | RT-qPCR | down | 102:103 | |||
| circ_104597 | down | |||||
| circ_103704 | up | |||||
| circ_103102 | up | |||||
| circTMEM2 | RNA-seq | PBMC | down | 20:20 | 2021 | Mahmoudi et al. [13] |
| circEZH1 | qRT-PCR | down | 21:21 | |||
| circKFBP8 | down | |||||
| circRARS | down | |||||
| chr3_196488683_196483770_−4913 | RNA-seq | plasma exosomes | up | 5:5 | 2021 | Tan et al. [83] |
| chr5_69175537_69174877_+660 | qRT-PCR | up | 6:6 | |||
| chr5_143057747_143054439_+3308 | up | |||||
| chr6_130956499_130926605_−29894 | up | |||||
| circ_0003999 | RNA-seq | peripheral blood | down | 3:3 | 2022 | Liao et al. [84] |
| circ_0030042 | RT-qPCR | down | 18:20 |
While circRNAs are abundantly expressed in the brain, it is possible that shared pathway alterations or genetic variants that are involved in the etiology of SCZ are also manifested in the periphery, such as in peripheral blood. In an attempt to determine whether circRNAs in peripheral blood could function as diagnostic or therapeutic biomarkers of SCZ, Yao et al. [82] comparatively analyzed the expression of circRNAs in PBMCs of nine SCZ and nine CTL. They found nine differentially expressed circRNAs. RT-qPCR in 102 SCZ patients and 103 CTL further validated that the expression levels of circ_104597, circ_102101, and circ_101836 were remarkably down-regulated and circ_103102 and circ_103704 were notably up-regulated. The presence of a combination of the three downregulated circRNAs predicted a relatively high success rate in the diagnosis of SCZ with a ROC curve of 0.8967. Additionally, it was detected that circ_104597 was down-regulated before treatment but up-regulated following eight weeks of treatment. Thus, it was concluded that circ_104597 might serve as a potential therapeutic biomarker of SCZ.
In 2020, Mahmoudi’s team [13] conducted a study that analyzed circRNA expression in PBMC from 20 patients with SCZ, 19 patients with bipolar disorder (BD), and 20 CTL. It was revealed that circTMEM2, circEZH1, circKFBP8, and circRARS were downregulated and that there were interactions between these circRNAs and miRNAs related to SCZ, such as miR-564 and miR-572. To explore whether circRNAs in plasma could be potential diagnostic and therapeutic biomarkers for SCZ, Tan et al.’s [83] research recognized four up-regulated circRNAs, chr5_69175537_69174877_+660, chr3_196488683_196483770_-4913, chr6_130956499_130926605_-29894, and chr5_143057747_143054439_+3308. With the use of bioinformatic analysis, these circRNAs were found to play potential roles in the stress response, histone ubiquitination, metabolic processes, and other mechanisms associated with SCZ, suggesting they may have the potential to become diagnostic circRNAs for SCZ. circRNAs contain abundant binding sites for miRNAs, which can control gene expression by binding to miRNA, and could thereby be involved in the initiation and progression of SCZ. Liao et al. [84] investigated this potential mechanism of SCZ by constructing a circRNA-miRNA-mRNA network. They determined that circ_0006151/ miR-4685-3p/ZBTB16, circ_0007963/miR-3127-3p/UBE2K, and circ_0000008/miR-1976/ZBTB16 were the top three core competitive endogenous RNA (ceRNA) networks with essential roles in SCZ.
There are currently few studies on circRNAs in peripheral blood in SCZ. However, based on the existing studies, it can be concluded that circRNAs in peripheral blood may have significant implications for SCZ as they provide a basis for the molecular mechanisms involved in the development and pathogenesis of SCZ. As a result, circRNAs could be utilized for early diagnosis and treatment of SCZ.
SCZ is a psychiatric disorder with unknown etiology [85] that affects approximately 1% of the world’s population [86]. Although there are medications [87] which have proven helpful in alleviating the acute symptoms of SCZ and impeding its recurrence, the prognosis for many patients remains unsatisfactory. Consequently, timely detection, diagnosis, and intervention are crucial to manage the progression of SCZ and optimize patient outcomes. Since the current practice of diagnosing SCZ is still highly subjective, it is essential to investigate objective biomarkers for diagnosis. It is evident from various studies [88, 89] that identifying biomarkers of SCZ in peripheral blood has recently emerged as a promising diagnostic tool. At the transcriptional level, there are numerous studies [89] involving mRNA as a biomarker for SCZ. However, given the stability of ncRNA in peripheral blood, and the continuous advancement and application of second-generation sequencing technology, ncRNA in peripheral blood has been considered a potentially innovative biomarker. Identifying ncRNA using this method has high sensitivity, specificity, and feasibility in addition to being non-invasive, making it a promising method for the diagnosis, prognosis, and treatment of SCZ [1, 90]. Accordingly, this review has focused on three types of ncRNAs (miRNA, lncRNA, and circRNA) and their potential application as biomarkers for the diagnosis or treatment of SCZ in peripheral blood. Through the comprehensive review of the literature, we have identified that ncRNAs in peripheral blood exert influence on the mechanisms involved in the ontogenesis and development of SCZ.
Studies have confirmed that miRNAs are essential modulators involved in the regulation of gene expression [12]. Their role in the maintenance of neurological development and the regulation of brain function, aberrantly expressed miRNAs likely participate in the development of numerous neuropsychiatric disorders. Through the review of miRNA-related studies in peripheral blood of SCZ, it has been discovered that there are certain miRNAs that are aberrantly expressed in peripheral blood, including miR-34a, miR-181b, miR-137, miR-132, miR-195, miR9-5p, miR-432, miR-7, miR-30e, miR-548d, miR-432, miR-449a, and let-7, which might be of significance for further understanding into the pathogenesis of SCZ as well as its diagnosis and treatment. Furthermore, several studies [26, 27] have detected corresponding correlations between aberrantly expressed miRNAs in peripheral blood and in the brain.
lncRNAs account for a substantial portion of the total amount of ncRNAs. They can function both independently and in conjunction with other proteins to serve numerous biological roles, including regulation of transcription, mediating the activation of proteins participating in histone modifications, chromatin remodeling, and regulating development of the nervous system [91]. lncRNAs have been shown to be broadly expressed in the central nervous system, where they participate in the regulation of brain development at both the transcriptional and post-transcriptional levels and are closely connected with multiple neuropsychiatric disorders, including SCZ [92]. Through the review of studies associated with lncRNAs in peripheral blood of SCZ, it should be noted that there are specific alterations in the expression of lncRNAs in peripheral blood samples of SCZ that may be of essential diagnostic value for SCZ and may also be predictors of the response to treatment. Close associations between certain lncRNAs, such as ASHG19A3A011462, ASHG19A3A02633, ASHG19A3A049471, ASHG19A3A044112, ASHG19A3A049556, IFNG-AS1, TMEVPG1, and NRON, in peripheral blood of SCZ patients and pathways involved in immunity and inflammation have been detected. Meanwhile, studies have also revealed apparent regulatory relationships between lncRNAs in peripheral blood of SCZ patients and the transduction of signaling pathways, such as NONHSAT089447, MEG3, CHAST, LINC00511, and DA376252. Additionally, it has been determined that the expression levels of NONHSAT041499, NEAT1, NEAT2, AK123097, ENST00000509804-1, uc011dma.1, and Gomafu in peripheral blood of SCZ patients appear to be altered after treatment. These lncRNAs may have potential as either promising therapeutic targets or for prognosis evaluation of patients by their expression level.
circRNAs are also essential constituents of ncRNAs, which primarily exert their regulatory functions at the post-transcriptional level and are intimately associated with numerous cellular and biological functions [93]. It has been shown in the brain that circRNAs are predominantly abundant in synapses and that the aberrant expression of circRNAs could be closely associated with the occurrence and development of neuropsychiatric disorders, such as SCZ [77]. It has been established that circRNAs exhibit high stability, organizational specificity in peripheral blood, and are easily detectable. Through the review of relevant studies on circRNAs in peripheral blood of SCZ, it was noted that certain circRNAs, such as circRNA_104597, circTMEM2, circEZH1, circKFBP8, and circRARS, also had aberrant expression and may have the potential to serve as diagnostic biomarkers or therapeutic targets for SCZ.
Exploration of the ncRNA expression profile in peripheral blood of SCZ patients has profound implications for identifying diagnostic and therapeutic biomarkers as well as further elucidating SCZ pathogenesis. This could have additional implications for early detection and diagnosis of SCZ, subtype differentiation of SCZ, and prediction of drug response. This review further elucidates the mechanisms involved in the regulation of gene expression and the molecular mechanisms of SCZ pathogenesis by further integrating studies related to the analysis of ncRNA expression profiles in peripheral blood of SCZ. Future studies integrating mRNA with different types of ncRNA may be of great significance for the diagnosis and treatment of SCZ, such as the miRNA-mRNA regulatory network, the circRNA-miRNA-mRNA regulatory network, and the ceRNA network. Examination of these pathways would also be of great value for elucidating the pathogenesis of SCZ.
The various types of ncRNAs that are differentially expressed in peripheral blood of SCZ patients could not only serve as potential diagnostic biomarkers and therapeutic targets for SCZ but could also have significant implications for further understanding the molecular mechanisms involved in the development of SCZ. In particular, the early diagnostic biomarkers obtained directly from peripheral blood are of great significance for the timely diagnosis and treatment of SCZ.
Nevertheless, relative to mRNA, there are few studies investigating the function of ncRNAs in various neuropsychiatric disorders and the utilization of ncRNAs as biomarkers of SCZ. There are several studies that have identified ncRNAs that could serve as potential biomarkers in the development of SCZ, but the specific mechanisms of these ncRNAs in the development of SCZ require additional investigation. Furthermore, the accuracy of diagnosing SCZ with a single biomarker is still limited, hence, additional exploration of a combination of multiple potential biomarkers in peripheral blood may further enhance diagnostic accuracy. Finally, clinical trials are required to determine the potential of ncRNAs obtained during RNA sequencing and analysis for the clinical diagnosis and treatment for patients with SCZ.
MTX and QY conceptualized and designed the study. MTX, YCZ, XYL, MDJ and LJY designed the figures and conducted a literature review. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work
Not applicable.
Thanks to all the peer reviewers for their opinions and suggestions.
This research was funded by S&T Promotion Project of Health Commission in Jilin province China, grant number 2022JC082, and S&T Development Plan Project in Jilin Province China, grant number 20220203125SF.
The authors declare no conflict of interest.
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
