- Academic Editor
Background: This study used
bioinformatics combined with statistical methods to identify plasma biomarkers
that can predict intracranial aneurysm (IA) rupture and provide a strong
theoretical basis for the search for new IA rupture prevention methods.
Methods: We downloaded gene expression profiles in the
GSE36791 and GSE122897 datasets from the Gene
Expression Omnibus (GEO) database. Data were normalized using the “sva” R
package and differentially expressed genes (DEGs) were identified using the
“limma” R package. Gene Ontology (GO) and Kyoto Encyclopedia
of Genes and Genomes (KEGG) pathway enrichment analyses were used for DEG
function analysis. Univariate logistic regression analysis, least absolute
shrinkage and selection operator (LASSO) regression modeling, and the support
vector machine recursive feature elimination
(SVM-RFE) algorithm were used to identify key
biomarker genes. Data from GSE122897 and GSE13353 were
extracted to verify our findings. Results: Eight
co-DEG mRNAs were identified in the GSE36791 and GSE122897
datasets. Genes associated with inflammatory responses were clustered in the
co-DEG mRNAs in IAs. CD6 and C-C chemokine receptor 7 (CCR7) were identified as key genes associated with IA. CD6
and CCR7 were upregulated in patients with IA and their expression
levels were positively correlated. There were significant differences in the
infiltration of immune cells between IAs and normal vascular wall tissues
(p
An intracranial aneurysm (IA) is a fusiform, balloon-like bulge in a weak area of a blood vessel in the brain. It is caused by abnormal local changes in the intracranial arterial blood vessel walls, which may be a result of local congenital defects of the blood vessel wall muscle layers or degeneration of the internal elastic layer of the blood vessel wall under the influence of acquired factors that are simultaneously affected by other factors, including hemodynamics [1]. IAs constitute the main cause of spontaneous subarachnoid hemorrhage, and their bleeding probability is second only to that of ischemic stroke and hypertensive cerebral hemorrhage [2]. When an IA ruptures without warning, approximately one-third of patients reportedly die before reaching the hospital. With the advancements in surgical techniques and microscopic instruments, the diagnosis and treatment of IAs have improved in recent years [3, 4, 5]. However, if the ruptured aneurysm hemorrhages again, the mortality rate is 60–80%. Approximately half of the surviving patients endure moderate-to-severe lifetime disability, which can cause a significant economic burden to their families and society [6]. Studying and analyzing the risk factors related to IA rupture can therefore help us to appropriately treat unruptured aneurysms in a timely manner, with obvious benefit to the patient.
Various factors cause IA rupture, including size and location of the IA, and
patient-related risk factors such as chronic hypertension and cigarette smoking
[7, 8]. With recent developments in molecular biology,
genetics, and other disciplines, our understanding of the differences between the
walls of IA blood vessels and those of normal blood vessels, and the role of
IA-related genes in the formation and rupture of IAs and their mechanism of
action, has improved through genome and differential gene expression analyses,
providing a strong basis for the development of new IA prevention and treatment
methods [9, 10]. An increasing number of researchers aim to identify effective
biomarkers to monitor IA formation and rupture by revealing its etiology and
pathogenesis [11]. Inflammatory cytokines such as matrix metalloproteinases
(MMPs), Monocyte Chemoattractant Protein-1
(MCP-1), Tumor Necrosis Factor-
The expression profiling (mRNA) datasets GSE36791 and GSE122897 were downloaded
from the Gene Expression Omnibus (GEO) database
(https://www.ncbi.nlm.nih.gov/geo/).
The GSE36791 dataset (plasma samples) contained 43 patients with
ruptured aneurysm (subarachnoid hemorrhage) and 18 control group patients (people
with unruptured IA). The GSE122897 dataset (arterial tissue
samples) contained 44 patients with ruptured aneurysm (subarachnoid hemorrhage)
and 16 control group patients. The GSE36791 and GSE122897
datasets were normalized using the “sva” package
(http://www.bioconductor.org/) respectively.
Differentially expressed genes (DEGs) were identified using the
“limma” R package
(http://www.bioconductor.org/). Statistical
significance was set at
Gene ontology (GO)
enrichment analysis was used to annotate the cellular components (CCs),
biological processes (BPs), and molecular functions (MFs) of the DEGs. The
biological pathways and functions of the DEGs were analyzed using the Kyoto
Encyclopedia of Genes and Genomes (KEGG) database
(https://www.kegg.jp/).
Statistical significance was set at p
The Student’s t-test was used for normally distributed variables. The
Wilcoxon rank-sum test was used for non-normally distributed variables. The
Two microarray datasets (GSE36791 and GSE122897) were used in this study. The
datasets were normalized and the DEGs were identified based on p
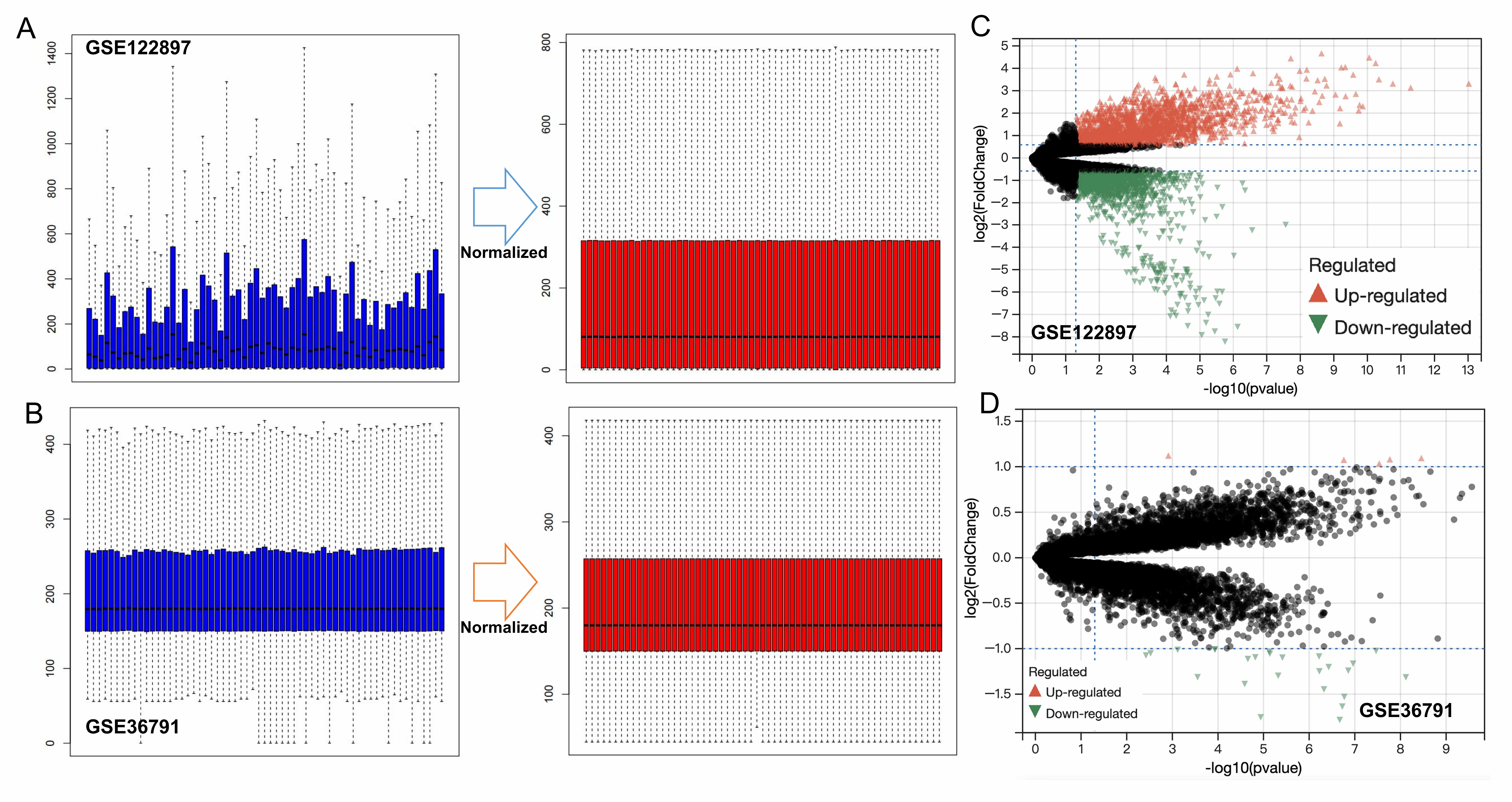 Fig. 1.
Fig. 1.Differentially expressed gene analysis. (A,B) Merged and normalize the chip data (GSE122897 and GSE36791) in the Gene Expression Omnibus (GEO) database. (C,D) Volcano plot of the differentially expressed genes (DEGs). Red and green dots represent genes that are significantly up- or down-regulated (|logFC|
 Fig. 2.
Fig. 2.DEG functional annotation. (A) Venn analysis between the DEGs from GSE36791 and GSE122897. Bubble charts showing the KEGG (B) and GO analysis results (including GO_BP (C), GO_MF (D) and GO_CC (E)) for eight co-DEG mRNAs. The size of the points represent the number of genes in the representative pathway. GO, gene ontology; BP, biological process; CC, cellular component; MF, molecular function; KEGG, Kyoto Encyclopedia of Genes and Genomes; mRNA, messenger RNA.
The “Cluster Profiler” package used to perform the GO and KEGG pathway enrichment analyses to determine the function of eight co-DEG mRNAs. KEGG pathway analyses revealed that the eight co-DEGs were enriched for cytokine-cytokine receptor interaction, inflammatory bowel disease, and arginine biosynthesis pathways (Fig. 2B). The top three terms included immune response, cytokine production, and leukocyte activation in the biological process category. The enriched terms were related to plasma membrane receptor complex, receptor complex, and plasma membrane protein complex in the cellular component category. The enriched terms mainly included cytokine receptor activity, cytokine binding, and lipoteichoic acid binding in the molecular function category (Fig. 2C–E).
Univariate logistic regression analysis was performed on the eight co-DEG mRNAs in the GSE36791 dataset. The results suggested that ARG1, C-C chemokine receptor 7 (CCR7), CD6, CLC, IL18RAP, IRAK3, and IL1R2 could influence IA rupture (Fig. 3A). These seven genes were also identified as possible diagnostic markers using the SVM-RFE algorithm (Fig. 3B). In addition, using the data from these seven genes in 61 patients, two potential predictors based on non-zero coefficients in the LASSO regression model were identified (Fig. 3C,D). ROC AUC analysis indicated that the expression levels of CD6 and CCR7 show potential diagnostic value and that these may be regarded as biomarkers for ruptured IAs (Fig. 3E). Binary transformation of CD6 and CCR7 was performed according to the cut-off value to construct the ROC curve (Fig. 3F,G, Table 1). Binary logistic regression analysis revealed that CD6 and CCR7 were independent predictive factors of IA rupture (Table 2).
 Fig. 3.
Fig. 3.Filter predictors of hub DEGs. (A) Binary logistic regression identification of key DEGs. The binary logistic regression algorithm was used to identify seven DEGs (ARG1, CCR7, CD6, CLC, IL18RAP, IRAK3, and IL1R2) as potential diagnostic markers for intracranial aneurysm rupture. (B) Biomarker selection using the support vector machine recursive feature elimination (SVM-RFE) algorithm. (C) Least absolute shrinkage and selection operator (LASSO) logistic regression identification of hub DEGs. (D) Optimal parameter (lambda) selection in the LASSO model used fivefold cross-validation via minimum criteria. (E) Venn diagram showing genes identified by all methods. (F,G) According to the cut-off value, binary transformation of CD6 and CCR7 was performed to construct the receiver-operating characteristic (ROC) curve. CCR7, C-C chemokine receptor 7; AUC, area under the curve.
| Test result variable | Cut-off value | Area under curve | Standard error | 95% CI | p |
| CD6 | 1.22 | 0.99 | 0.42 | 0.82–0.98 | 0.00 |
| CD6 |
/ | 0.85 | 0.56 | 0.74–0.96 | 0.00 |
| CCR7 | 2.21 | 0.86 | 0.61 | 0.71–0.97 | 0.00 |
| CCR7 |
/ | 0.84 | 0.67 | 0.51–0.84 | 0.00 |
CI, confidence interval.
| Variable | IA | Control | p | B | OR | 95% CI | p |
| CD6 | 0.00 | 2.82 | 11.49 | 1.78–74.03 | 0.01 | ||
| CD6 High | 16 | 8 | |||||
| CD6 Low | 2 | 35 | |||||
| CCR7 | 0.00 | 2.44 | 16.78 | 2.43–115.73 | 0.00 | ||
| CCR7 High | 13 | 2 | |||||
| CCR7 Low | 5 | 41 |
CI, confidence interval; IA, intracranial Aneurysm; OR, odds ratio; B, Regression coefficient.
Nomogram maps were constructed to predict the probability of IA rupture in patients with IA using CD6 and CCR7 (Fig. 4A). The nomogram was validated using a calibration curve. The calibration plots indicate that the nomogram performed well in predicting the rupture probability in patients with IA (Fig. 4B,C). The AUC indicates that CD6 and CCR7 (in the nomogram model) more accurately predict the rupture probability in patients with IA. The ROC curve of the nomogram model was evaluated and the AUC was found to be 0.90 (Fig. 4D). These results suggest that the nomogram constructed using the two-gene signature has high accuracy in predicting the rupture probability in patients with IA. In addition, CD6 and CCR7 were upregulated in patients with IA and their expression levels were positively correlated (Fig. 4E,F).
 Fig. 4.
Fig. 4.Construction of the predictive model. (A) A ruptured intracranial aneurysm (IA) nomogram was constructed for CD6 and CCR7. Each gene is given a score; the total score is calculated by adding each score and projecting this onto a lower total score scale. (B) Calibration curves of the ruptured IA nomogram prediction. The y-axis represents net benefit. The blue line represents the nomogram. The gray line represents all patients. The net benefit is calculated by subtracting the proportion of all false-positive patients from the proportion of true positives. (C) Calibration plot of predicted ruptured IAs, where actual probability is the actual outcome. Each dot mark at the top represents a patient. The x-axis represents the estimated ability to predict IA rupture derived from the nomogram and the y-axis represents the actual ability. (D) Receiver operating characteristic analysis of the ruptured IA nomogram. (E) CD6 and CCR7 mRNA expression in the ruptured IA and control groups in the GSE36791 dataset. *p
These key genes were validated using the GSE122891 and GSE13353 datasets.
CD6 and CCR7 were upregulated in patients with IA and their
expression levels were significantly positively correlated (Fig. 5A1,A2,B1,B2). Hence, CD6 and CCR7 show potential diagnostic
value (p
 Fig. 5.
Fig. 5.Effectiveness test of the predictive model. (A1,B1) CD6 and CCR7 expression levels in the ruptured IA and control groups (GSE122897 and GSE13353 datasets), *p
We analyzed immune cell infiltration and immune cell function in the ruptured IA and control groups (Fig. 6A) Heat map showing differential immune cell infiltration and immune functions between the ruptured IA and control groups (Fig. 6B). Heat map showing correlations between immune cell infiltration and (Fig. 6C) immune functions. Immune cell infiltration analysis showed that CD8+ T, mast, macrophage, Th1, Th2, T helper, Tfh, TIL, and Treg cells were up-regulated in IA patients compared with normal patients. Immune cell function analysis showed that cytolytic activity, antigen presenting cell (APC) co-inhibition, APC co-stimulation, inflammation-promotion, CCR, T cell co-inhibition, human leukocyte antigen (HLA), major histocompatibility complex (MHC) class I, cell cycle checkpoint, para-inflammation, T cell co-stimulation, and Type II interferon (IFN) response functions were more active in IA patients compared with normal patients. Additionally, we performed correlation analysis between the up-regulated cells (Fig. 6B) and between the functions found to be more active in the IA group (Fig. 6C). Differential immune cell infiltration and immune functions between the ruptured IA and control groups show in Fig. 6D.
 Fig. 6.
Fig. 6.Immune cell infiltration and
immune function analysis.
(A) Heat map showing
differential immune cell infiltration and immune functions between the ruptured
IA and control groups. (B,C) Heat map showing correlations between immune cell
infiltration and immune functions. Numbers in the squares represent the strength
of the correlation. The redder the color, the stronger the correlation. (D)
Differential immune cell infiltration and
immune functions between the ruptured IA and control groups. *p
Furthermore, we performed correlation analysis between the differentially infiltrated immune cells identified above and CCR7 and CD6 respectively. CCR7 was positively correlated with activated mast cells, neutrophils, M0 macrophages, and activated dendritic cells, but significantly negatively correlated with M2 macrophages M2, activated natural killer (NK) cells, resting dendritic cells, and M1 macrophages (Fig. 7A). CD6 was positively correlated with neutrophils, plasma cells, activated mast cells, activated dendritic cells, and resting NK cells, but significantly negatively correlated with activated NK cells, M1 macrophages, resting dendritic cells, and M2 macrophages (Fig. 7B).
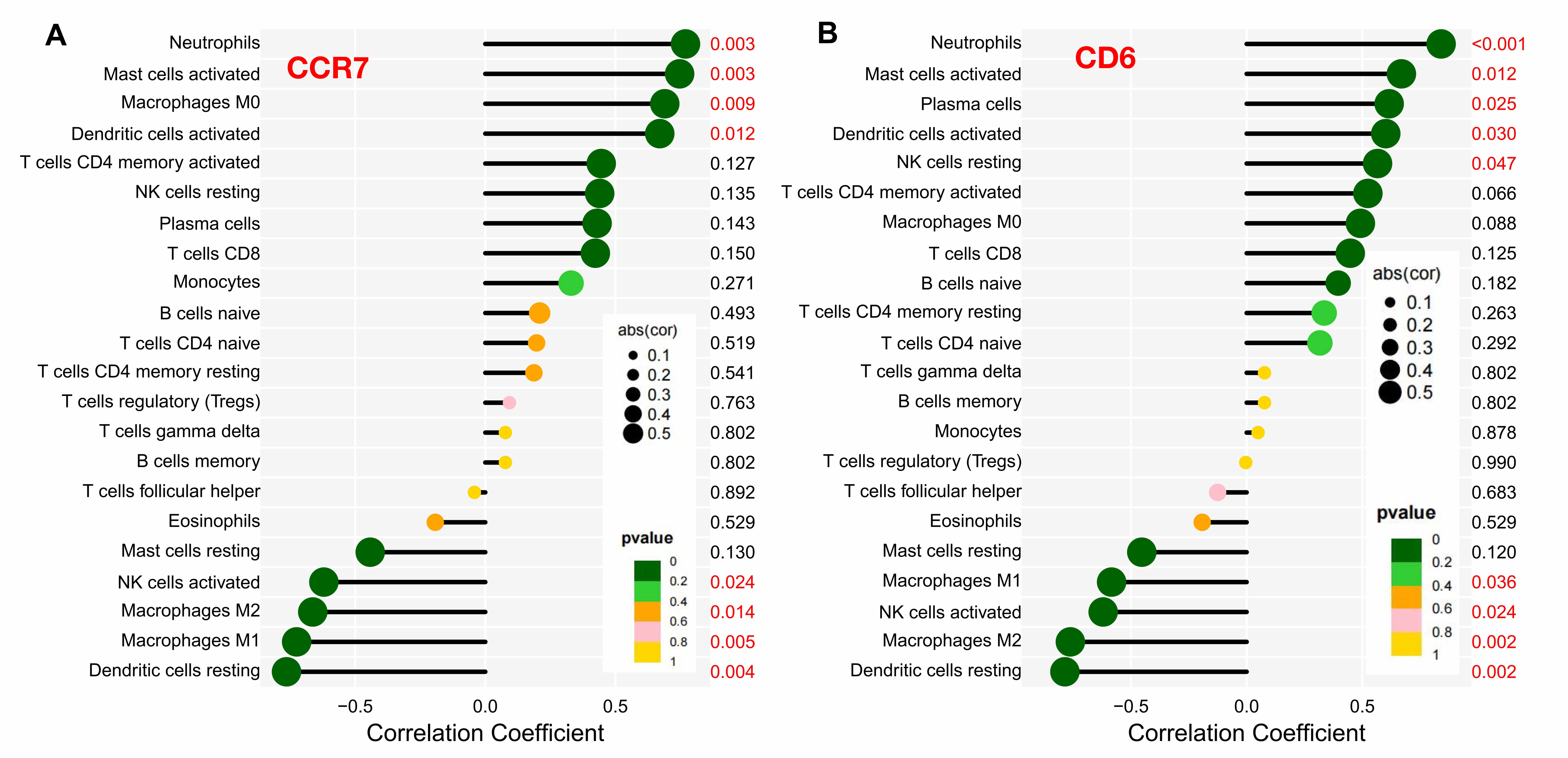 Fig. 7.
Fig. 7.Correlation analysis of infiltrating immune cells and functions. (A) Correlation between CCR7 and infiltrating immune cells and functions in the ruptured IA group. (B) Correlation between CD6 and infiltrating immune cells and functions in the ruptured IA group.
To identify differentially expressed mRNAs between the IA and control groups, high-throughput ChIP-sequencing data were extracted from the aorta tissue and plasma of patients. Based on the co-DEG mRNAs, nomogram maps were constructed to predict the rupture probability in patients with IA using bioinformatics and statistical analysis. The GSE122891 and GSE13353 datasets were used for validation. The transcriptional levels of CD6 and CCR7 were increased in patients with IA compared with those in control individuals, and the expression of CD6 was significantly and positively correlated with that of CCR7. Furthermore, functional annotation of the co-DEG mRNAs, particularly those associated with inflammation-related signaling pathways, was performed.
In recent years, with the advances in IA research, it has been demonstrated that extracellular mechanisms of chronic inflammation and pathological remodeling of the arterial wall may be associated with the occurrence and development of IA [15]. Studies show that while inflammatory cells may infiltrate the normal arterial wall, large quantities of inflammatory cells are found in the arterial walls of IAs, which is related to the massive infiltration of immunoglobulins and macrophages during aneurysm formation [16, 17]. Inflammatory cells can secrete cytokines, which can cause abnormal secretion of pro-inflammatory factors by vascular endothelial and smooth muscle cells, thereby destroying the inner wall of the blood vessels. A variety of inflammatory cells and factor indicators, alone or in combination, have been identified as potential new immune and inflammatory response markers. These include the platelet to lymphocyte ratio, neutrophil to lymphocyte ratio and lymphocyte to mononuclear cell ratio, which have been shown to participate in the development and occurrence of cerebrovascular disease, and to be closely related to its prognosis [18, 19, 20]. Studies have shown that monocyte chemoattractant protein 1 (MCP1) upregulation in patients with aneurysm rupture is similar to that in patients with acute cerebral ischemia. This may be related to the inflammatory reactions in brain ischemia and tumor rupture caused by strong vasoconstriction in these patients [21, 22]. MMPs are proteolytic enzymes that degrade the extracellular matrix, and increased MMP expression has been detected in aneurysms. The expression levels of MMP-9 and MMP-2 were significantly increased in patients with ruptured aneurysms compared with those with unruptured aneurysms [23].
Our study showed that the expression of CCR7 and CD6 were significantly upregulated in IA patients. Analysis of immune cell infiltration and function showed increased immune-inflammatory cell infiltration in IA, and CCR7 and CD6 expression was significantly positively correlated with immune-inflammatory cell functions. T cells express CD6, a co-stimulatory membrane glycoprotein. CD6 and its ligand can activate the leukocyte cell adhesion molecule (ALCAM), and they are involved in the activation and trafficking of T cells [24]. This has been implicated in multiple autoimmune and inflammatory diseases, including multiple sclerosis, rheumatoid arthritis, and inflammatory bowel disease [25, 26]. Chalmers et al. [27] demonstrated that the CD6/ALCAM pathway possibly promotes lupus nephritis through CD6, which is a strong biomarker of the disease. CCR7 plays an important role in the immune response, mediating the delivery of antigens by dendritic cells to secondary lymphoid organs or tissues, and activating T lymphocytes to induce an immune response. CCR7 is expressed in a variety of inflammatory and immune cells and participates in the regulation of inflammatory and immune responses [28, 29]. Changes in CCR7 expression have been reported to be closely associated with the progression of carotid atherosclerosis [30]. In atherosclerosis, CCR7 promotes the adhesion and infiltration of macrophages to the vascular endothelium and the phagocytosis of oxidized low-density lipoproteins leading to the formation of foam cells, which accumulate lipids causing atherosclerotic plaque formation [31, 32]. Llodrá et al. [33] proposed that CCR7 signaling is critical for the outward migration of macrophages during the early stages of atherosclerosis resolution. Several factors can cause IA rupture. However, inflammation can directly cause wall degeneration and lead to IA enlargement, which is the key factor that ultimately increases rupture risk [34]. When the vascular wall is remodeled, it becomes decompensated by inflammation and cannot resist hemodynamic stress, resulting in aneurysm rupture [35].
Our study had some limitations. The results have not been confirmed by in vitro studies or other functional experiments. Although our results are based on the GEO database, we applied different statistical methods to validate the results and showcase our new findings. These results have certain implications for subsequent mechanism research.
Using bioinformatics and statistical analysis, we have shown that CD6 and CCR7 in inflammation-related signaling pathways are closely associated with IA rupture and may play an important role in its pathogenesis. Further studies are needed to investigate the potential value of CD6 and CCR7 as diagnostic biomarkers.
All the data used in this study are available in GEO (https://www.ncbi.nlm.nih.gov/geo) which are public functional genomics data repositories.
DDX and ZSW designed the research study. DDX and XQL drawd the images. DDX and XQL analyzed the data. DDX write the article. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
Not applicable.
This work was supported by the Natural Science Foundation of Fujian Province (grant number: 2023J01122595).
The authors declare no conflict of interest.
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
