Angiotensin-converting enzyme 2 (ACE2), the host cell-binding site for SAR-CoV-2, poses two-fold drug development problems. First, the role of ACE2 itself is still a matter of investigation, and no specific drugs are available targeting ACE2. Second, as a consequence of SARS-CoV-2 interaction with ACE2, there is an impairment of the renin-angiotensin system (RAS) involved in the functioning of vital organs like the heart, kidney, brain, and lungs. In developing antiviral drugs for COVID-19, ACE2, RNA-dependent RNA polymerase (RdRp), and the specific enzymes involved in the viral and cellular gene expression have been the primary targets. SARS-CoV-2 being a new virus with unusually high mortality, there has been a need to get medicines in an emergency, and the drug repurposing has been a primary strategy. Considering extensive mortality and morbidity throughout the world, we have made a maiden attempt to discover the drugs interacting with RAS and identify the lead compounds from herbal plants using molecular docking. Both host ACE2 and viral RNA-dependent RNA polymerase (RdRp) and ORF8 appear to be the primary targets for the treatment of COVID-19. While the drug repurposing of currently approved drugs seems to be one strategy for the treatment of COVID-19, purposing phytochemicals may be another essential strategy for discovering lead compounds. Using in silico molecular docking, we have identified a few phytochemicals that may provide insights into designing herbal and synthetic therapeutics to treat COVID-19.
With the declaration of Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) as the pandemic by WHO, it has emerged as a public health emergency worldwide. The dreaded face of the SARS-CoV-2, commonly known now as Coronavirus disease-19 (COVID-19), can be realized by the fact that there is a consistent increase in the number of cases reaching above 32 million cases and above 1 million deaths worldwide (World Health Organization., 2020). Although the fight against any disease has been an endless task for humanity, the emergence of COVID-19 disease with characteristics of the top five causes of mortality in the world has posed the most significant challenges for drug discovery for the treatment of this disease. There is current guidance for early ambulatory treatment of COVID-19 based on pathophysiologic principles regarding viral replication, immune dysregulation, and thrombosis. This algorithm is designed to welcome new drug candidates for testing in clinical trials and when demonstrated to be effective, incorporation into clinical practice (McCullough et al., 2020).
COVID-19 being the novel virus, the first challenge is to develop an antiviral
drug that can effectively stop its replication after entry into the human body.
The targets in the development of antiviral drugs include the receptor protein to
which the virus binds, the enzymes or nucleosides that facilitate the replication
of viral RNA or DNA, and the proteins involved in uncoating, shedding of viruses
or release of its contents (Paintsil and Cheng, 2009). It is a well-known
fact that the development of the auto-resistance (immunity) is the ultimate
answer for the cure of the viral disease once the virus enters the human system.
The development of specific interferons is a strategy for developing new
antiviral drugs (Samuel, 2001). Although SARS-CoV-2 nonstructural
proteins nsp13, nsp14, nsp15, and orf6 may inhibit interferon signaling (Y. Yuen et al., 2020), IFN
SARS-CoV-2 binds to Angiotensin-Converting Enzyme 2 (ACE2), a site that is also involved in the pathology of leading causes of mortality in the world, including cardiovascular, respiratory, and GI dysfunction. ACE2 is a mystery to the medical scientists as it regulates the renin-angiotensin system (RAS), which in turn, has an influence on the cardiovascular system, including the vital organs, kidney, heart, brain, lungs, and vasculature (Patel et al., 2016). Further, it is still unclear whether ACE2 alone or in concert with angiotensin receptors AT1, AT2, or Mas receptors contribute to the pathology of COVID-19.
Third and yet another critical challenge is to control the harmful effects induced by the interference of ACE2. The immediate problem when a virus enters the human body is to control the patient’s intermittent pathological damage. The antiviral drug could prevent morbidity and mortality if given before the virus has produced sufficient copies or has not infected an adequate number of cells in the body. It is also efficacious if the virus’s time to produce morbidity and mortality is long enough. The present review article focuses on these challenges identifying new drug targets and strategies for developing treatments for ACE2-induced derangement of RAS responsible for morbidity and mortality (Danser et al., 2020; Messerli et al., 2020).
It is now well established that membrane-anchored ACE2 is the site of cell entry for COVID-19. Epithelial cells of the lung, intestine, kidney, heart, and blood vessels express ACE2. ACE2 is also present in the oral and nasal mucosa, skin, lymph nodes, thymus, bone marrow, spleen, liver, and brain (Bilinska et al., 2020; Chen et al., 2020; Hamming et al., 2004; Li et al., 2020a; Zhang et al., 2020), regulating the vasculature, inflammation, oxidative stress, fibrosis, and cell proliferation. Indeed, COVID-19 is associated with both pulmonary pathology and extrapulmonary manifestations in multiple vital systems, including hematologic-immune, cardiovascular, renal, gastrointestinal and hepatobiliary, endocrine-neurologic, and ophthalmic dysfunctions (Gupta et al., 2020; Wiersinga et al., 2020). Furthermore, the interindividual clinical variability associated with COVID-19 vulnerability and severity, are influenced by differences in the expression of ACE2 (Babadaei et al., 2020; Benetti et al., 2020).
ACE2 regulates Ang II action by hydrolyzing Ang II to Ang1-7 (Pai et al., 2017; Verdecchia et al., 2020). Ang1-7 exerts anti-inflammatory, antithrombotic, antihypertensive, antiarrhythmic, and cardioprotective action through G-protein-coupled Mas receptors (MasR) (Bourgonje et al., 2020; Kuba et al., 2013; Santos et al., 2018; Uhal et al., 2011). Ang II is produced by ACE, a zinc metalloproteinase that converts the hormone Ang I to Ang II. Ang II acting via AT1 receptors causes vasoconstriction, inflammation, cytokine secretion, thrombosis, endothelial and myocardial dysfunction. Ang II also acts on AT2 receptors to reduce inflammation and oxidative stress (Mehta and Griendling, 2007). Thus, ACE and ACE2 arms of RAS counterbalance their actions fine-tuning many physiological functions (Fig. 1).
 Fig. 1.
Fig. 1.Overview of SARS-CoV-2 infection and renin-angiotensin system:
During the COVID-19 infection, the SARS-CoV-2 binds to the angiotensin-converting
enzyme 2 (ACE2) receptors present on the cell surface which undergo endocytosis,
and the virus enters inside the cell. Upon entering the cell, it synthesizes RNA
using the host metabolic processes in which enzyme RNA dependent RNA polymerase
(RdRp) is essential. Due to cellular damage caused by the virus, the alveolar
macrophages generate various proinflammatory cytokines like interleukin-6 (IL-6),
macrophage inflammatory protein 1
ACE2 has an extracellular receptor site for the binding of viral spike (S) proteins leading to the virus’s entry into the host cell, especially into lung alveolar epithelial cells and intestinal enterocytes due to high expression of ACE2 in these epithelial cells. Binding of SARS-CoV-2 to ACE2 triggers the internalization of ACE2-virus complex and consequent reduction in ACE2 levels. This leads to an imbalance between ACE2 and ACE arms of RAS. Reduced Ang1-7 action and increased AT1 activation trigger a cascade of inflammatory responses (Tay et al., 2020). Administration of exogenous Ang1-7 and recombinant circulating ACE2 that could serve as a decoy for SARS-CoV-2 binding to membrane ACE2 are under investigation for COVID-19 treatment. Furthermore, angiotensin receptor blockers (ARBs) and ACE inhibitors are also under investigation in the treatment of COVID-19 (Birnbaum, 2020; Inal, 2020; Lopes et al., 2020).
It is very well documented that imbalance in the levels of ACE, and ACE2
function during COVID-19 infection is one of the factors augmenting the cytokine
storm (Crowley and Rudemiller, 2017; Gheblawi et al., 2020). There is
a crosstalk between RAS and complement system since Ang II and AT1 receptor
activation activates complement cascade viz. C5a and C5b-9 (Java et al., 2020; Van Laecke and Van Biesen, 2017). Ang II causes phosphorylation of the p65
subunit of NF-
Moreover, activating the AT1 receptor, Ang II causes regulation of the
Mitogen-Activated Protein Kinases pathway, which ultimately regulates the release
of TNF
Siddiqi and Mehra (2020) proposed three stage-sequence of SARS-CoV-2 infection (Siddiqi and Mehra, 2020). These three stages are also referred to as mild-to-moderate no risk, mild-to-moderate risk, and severe disease groups (Dehelean et al., 2020). During the Stage-I (mild-to-moderate no risk) infection involving innate immunity, treatment with convalescent sera, and non-specific (polyvalent) immunoglobulins may be effective. During Stage-II, there is virus replication, which leads to severe lung inflammation (viral pneumonia) resulting from the release of proinflammatory mediators and consequent multiple organ failure. Moreover, massive release of proinflammatory cytokines is also responsible for neuropsychiatric symptoms associated with COVID-19 that are mediated by proinflammatory cytokines such as depression (Ettman et al., 2020; Petralia et al., 2020).
Antiviral therapy and anti-inflammatory therapy, such as anti-IL-6, may be combined. In Stage-III, there is a marked hyperinflammation. In the RECOVERY trial, low dose dexamethasone reduces deaths in patients hospitalized with COVID-19 who need ventilation. Dexamethasone also reduced mortality by 20% in other hospitalized patients receiving oxygen only. However, there was no benefit among patients who did not require respiratory support. In patients hospitalized with COVID-19, the use of a low dose of dexamethasone resulted in lower 28-day mortality among those receiving either invasive mechanical ventilation or oxygen alone, but not among those receiving no respiratory support (Horby et al., 2020; Mahase, 2020). Dexamethasone benefits for patients with COVID-19 likely arise from its immunosuppressive properties. Moreover, the use of low-dose dexamethasone may reduce the development of glucocorticoid resistance.
Glutathione depletion, as observed in Glucose 6-phosphate dehydrogenase (G6PD) deficient subjects, predisposes them to hemolysis upon SARS-CoV-2 infection. When administered to G6PD-deficient patients with severe COVID-10 disease, N-acetylcysteine (NAC) reverses glutathione depletion, suppressed hemolysis, and elevated liver enzymes, C-reactive protein (CRP), and ferritin, leading to full recovery of the G6PD-deficient patient. NAC also produced clinical improvement in 9/10 respirator-dependent COVID-19-infected patients without G6PD deficiency, possibly by suppressing the cytokine storm (Ibrahim et al., 2020b). In the last two decades, there has been an outburst of two major similar coronavirus severe respiratory pandemics in humans, i.e., Severe Acute Respiratory Syndrome (SARS) in 2003, Middle East Respiratory Syndrome (MERS) in 2012. SARS and MERS are caused by SARS-CoV and MERS-CoV viruses enters into host cells via their interaction with ACE2 and DPP4, respectively (Gorbalenya et al., 2020; Hilgenfeld and Peiris, 2013; Zhong et al., 2003). Various studies also depict the relationship between cardiovascular, metabolic diseases, and SARS and MERS infections (Badawi and Ryoo, 2016; Yang et al., 2006). However, there are no therapeutically proven inhibitors of ACE2 for the treatment of coronavirus infections. In the absence of authenticated medicine available, clinical management of the patient in the ICU is a great challenge.
Various parts of the world have revealed that cardiovascular disturbances, including hypertension and thrombotic events, diabetes, and acute respiratory distress, are emerging comorbidities. Patients with existing cardiovascular diseases face a higher risk of COVID-19 (Kuno et al., 2020; Li et al., 2020b; Richardson et al., 2020b). In patients with cardiovascular diseases, diabetes mellitus, and hypertension treated with ACE inhibitors and ARBs, the expression of ACE2 contributes to worsening the prognosis of COVID-19 patients (Birnbaum, 2020; Cannata et al., 2020; Fosbøl et al., 2020; Mackey et al., 2020; Pirola and Sookoian, 2020; Silhol et al., 2020). In contrast, several recent clinical studies provide evidence that ACE inhibitors and ARBs may be beneficial in COVID-19 patients with hypertension and, hence, recommend continuing ACE inhibitors and ARBs (Kang et al., 2020; Mackey et al., 2020).
During the management of the patients, it is necessary to consider RAS’s role holistically when SARS-CoV-2 occupies the ACE2 receptors. The conventional de novo drug discovery process takes 10 to 15 years for a drug to become available for human use, and the chance of success rate is less than 10% (Gilbert et al., 2003; Reichert, 2003). The drug repurposing/repositioning has emerged as a fast and safe drug discovery strategy to develop effective medication, especially in case of diseases where no viable treatment is available (Ashburn and Thor, 2004; Beck et al., 2020). While attempts are ongoing to target ACE2 for developing new antivirals, the repurposing strategies to consider the use of ACE inhibitors and ARBs are also evolving.
Various drugs evaluated against SARS-CoV and MERS-CoV have been explored in terms of efficacy for repurposing during the SARS-CoV-2 outbreak (Horie et al., 2020; Kalra et al., 2020; Kandimalla et al., 2020; Pizzorno et al., 2020; Santos et al., 2020). Synthetic drugs (example chloroquine and hydroxychloroquine), and antivirals (remdesivir and favipiravir) and herbal phytoconstituents (baicalin, glycyrrhizin, hesperetin, quercetin, and scutellarin), are considered for COVID-19 due to their potential action on host ACE2 or RdRp (Chen et al., 2004; Ekins et al., 2020; Ho et al., 2007; Zhang and Yap, 2004). None of the other antiviral therapeutic interventions like Lopinavir/Ritonavir, and Remdesivir seemed to alter the natural clinical course of the disease based on the available literature (Romagnoli et al., 2020; Rosendaal, 2020), however multiple investigations are ongoing.
Besides the repurposing efforts, in silico molecular docking and simulations studies revealed potential synthetic and naturally occurring compounds in herbal and medicinal plants that interact with both viral and host target proteins (Amin and Abbas, 2020; Gao et al., a; Maurya et al., 2020; da Silva et al., 2020). These compounds targeted the SARS-CoV-2 spike proteins (Abdelli et al., 2020; Kiran et al., 2020; Maurya et al., 2020; Zhang et al., 2005) Mpro (Ibrahim et al., 2020a; Jin et al., 2020; Joshi et al., 2020; Kumar et al., 2020; Narkhede et al., 2020) and 3CLpro proteases (Gyebi et al., 2020; Ul Qamar et al., 2020), host ACE2 (Benarba and Pandiella, 2020; Maurya et al., 2020) and host TMPRSS2 (Benarba and Pandiella, 2020; Rahman et al., 2020). In addition to the docking scores, few of these studies have determined drug-like properties in silico (Narkhede et al., 2020) and have reported activity of compounds in functional assays in vitro (Jin et al., 2020). These investigations open new avenues for identifying naturally occurring novel lead compounds for the treatment of COVID-19, and for designing potential new chemical entities (NCEs) based on lead natural compounds. We focus on host RAS, viral RdRp, and ORF8 proteins as potential therapeutic targets using in silico molecular docking investigations.
The virus is an obligatory intracellular parasite that relies on host biosynthetic factors and its replication and propagation environment. Thus, the concern in developing an antiviral drug is to develop an effective drug with selective toxicity for viruses and not the host. SARS-CoV-2 binds to ACE2 on the host cell membrane and, by endocytosis, enters into the cell. Small molecules targeting the viral binding to the host cells, namely ACE2 in SARS-CoV-2, and nucleoside analogs which interfere with viral replication, are potential therapeutics for treating viral diseases. Current pharmacotherapy for COVID-19 relies upon suppressing viral replication (example remdesivir), inflammation (example dexamethasone), immune response (example azithromycin), and blood coagulation (example enoxaparin). There are no known viral entry blockers that target host ACE2. Blocking viral entry into host cells may be the most effective strategy for prophylactic and therapeutic treatment of COVID-19. We have attempted to identify naturally occurring phytoconstituents that may target ACE2 and block viral entry using molecular docking studies. We also evaluated the molecular interactions of a select number of naturally occurring phytoconstituents at host AT1 and AT2 receptors, and SARS-CoV-2 RdRp and ORF8 proteins using molecular docking.
Five proteins ACE2 (PDB ID: 2AJF), ORF8 (PDBID: 6BB5), AT1 (PDBID:4ZUD), AT2 (PDBID: 4UNF) and RdRp (PDBID: 7BV2) were selected for docking protocols for identifying SARS-COV-2 ligands. The selection of PDB’s was made on the basis of species and resolution. The target proteins were then imported from RCSB Protein Data Bank saved and prepared. The structure was preprocessed by adding hydrogens and checked for missing residues. The selection of water molecules was set at default, and protein was optimized, followed by energy minimization at the default setting. The docking grid was prepared, and docking performed.
Synthetic or phytoconstituents molecules were docked using the GLIDE module of
Maestro 9.6 as per the Glide protocol given in Schrödinger. All the default
parameters were used. For ligand preparation, the pH was 7.0
ACE2 (PDB ID: 2AJF), ORF8 (PDB ID: 6BB5), AT1 (PDB ID: 4ZUD), AT2 (PDB ID: 4UNF) and RdRp (PDB ID: 7BV2) were downloaded from the PDB database, and the potential binding sites were predicted by Prepwizard module of Schrödinger. A total of 30 compounds synthetic or natural origin were docked to the binding site of five proteins ACE2 (PDB ID: 2AJF), ORF8 (PDB ID: 6BB5), AT1 (PD BID: 4ZUD), AT2 (PD BID: 4UNF) and RdRp (PD BID: 7BV2) with Glide docking module of Schrödinger, which provides molecular mechanics algorithm along with shape analysis for accurately docking ligands into active sites. The docked poses were then analyzed using Schrödinger software’s imaging tools.
For ACE2 docking, a selective ACE2 inhibitor GL1001 (MLN4760), was used as
the positive control. GL1001 inhibits ACE2
(IC
At an earlier phase of COVID-19 pandemic, antimalarial drugs hydroxychloroquine and chloroquine were evaluated for potential efficacy in COVID-19 patients (Zhou et al., 2006). We found that hydroxychloroquine, chloroquine, and the active metabolites desethychloroquine, desethyhydroxychloroquine and bisdesethyl-chloroquine bound tightly to the active site of the ACE2 occupying the same binding pocket as GL1001. Hydroxychloroquine (docking score = -8.8 kcal/mol) occupies the ACE2 receptor cavity and makes hydrogen bonding with Phe40, Ala348, Asp350, Phe390, and His401.
Both chloroquine and hydroxychloroquine act by inhibiting glycosylation of host receptors, proteolytic processing, and endosomal acidification (Sanders et al., 2020). A recent in vitro study showed hydroxychloroquine was more potent than chloroquine against SARS-CoV-2 (Yao et al., 2020). Several clinical studies evaluating the action of chloroquine are ongoing, and findings from few clinical studies have been reviewed elsewhere (Chang and Sun, 2020; Pal et al., 2020). A comprehensive meta-analysis of clinical findings reports that the use of hydroxychloroquine was associated with increased mortality rates and increased risk of adverse events in SARS-CoV-2 patients (Chandrasekar et al., 2020). Based on 32 studies involving 29,192 participants, Cortegiani et al. (2020) concluded that chloroquine/ hydroxychloroquine confers no benefit in terms of mortality in hospitalized patients with COVID-19 (Cortegiani et al., 2020). Combination therapy with macrolide may be associated with harm, and postexposure prophylaxis with chloroquine/hydroxychloroquine may not reduce the rate of COVID-19 (Cortegiani et al., 2020). Although the docking of hydroxychloroquine was better than the reference compound GL-1001, the narrow therapeutic index associated with the cardiotoxicity and controversy regarding the efficacy and safety of hydroxychloroquine for prophylactic treatment of COVID-19 tempted us to explore potential natural products-based ACE2 inhibitors.
Herbal drugs are in use for the treatment of various diseases, including respiratory viral infections. The significant advantage of using herbs in viral respiratory infections is due to their potent anti-inflammatory and immunostimulatory action (Patwardhan et al., 2020). Some of the active phytoconstituents in medicinal plants have inhibitory action on coronaviruses (Boukhatem and Setzer, 2020; Mani et al., 2020), however determination of functional actions on ACE2 is critical to elucidate agonist/activation or inhibitor/antagonist. Considering the pathogenesis of COVID-19, we identified some Indian medicinal plants with potent anti-inflammatory, immunomodulatory, and antiviral activities for developing potential treatment of COVID-19. These plants identified were Solanum nigrum, Euphorbia prostrata, Acalyphaindica, Lantana camara and Ricinus communis. A literature search to identify the phytoconstituents of these plants revealed that they contain flavonoids, flavonoid glycosides (Huang et al., 2010; Patil and Limaye, 2017), triterpenoids (Ghisalberti, 2000), steroidal glycoalkaloids (Gu et al., 2018) and other compounds of different classes (Herbert et al., 1991; Nahrstedt et al., 1982; Tai et al., 2018). Furthermore, it has been reported that flavonoids can bind to the SARS-CoV-2 S protein, a viral surface glycoprotein required for initial attachment and internalization within host cells (Rane et al., 2020). Flavonoids, in particularly, luteolin, quercetin, apigenin, amentoflavone, epigallocatechin (EGC), epigallocatechin gallate (EGCG), gallocatechin gallate (GCG) and kaempferol were reported to inhibit the proteolytic activity of SARS-CoV 3CLpro (Nguyen et al., 2012; Ryu et al., 2010) and 3a channel protein of coronavirus (Schwarz et al., 2014). Certain flavonoids viz., luteolin, quercetin, rutin, kaempferol, and apigenin showed ACE inhibitory activity (Guerrero et al., 2012). In addition to flavonoids, we have also docked certain other compounds of phytochemical groups like triterpenoids, steroidal glycoalkaloids, phenolic compounds since these were major chemical constituents of the antiviral plants mentioned above‘. Molecular docking study was also carried out for selected steroidal glycoalkaloids: solamargine, solasonine, solasodine, solanine; flavonoids: rutin, naringenin, luteolin, myricetin, quercetin, apigenin, kaempferol and hesperetin, hesperidin, apigenin-7-O-glucoside, luteolin-7-O-glucoside; triterpenoids: Lantadene A, B, C, oleanolic acid, acalyphin, ricinine, dioscin, and epigallocatechin gallate, verbascoside.
Results of these docking studies revealed that the compounds rutin, verbascoside, and hesperidin bound tightly at the active site of ACE2 (Table 1). The compound rutin (docking score: -11.5 kcal/mol) well occupied the receptor cavity through hydrogen bonding with Ala348, Asp350, Phe390, Ser44, Ser47 Arg393, Asn394 (Fig. 2 and 3) and verbascoside (docking score: -10.2 kcal/mol) showed hydrogen bonding interaction with Ala348, Asp350, Asp382, Phe390, Glu375, His37, Asn394, and Glu402, whereas hesperidin (docking score: -9.5 kcal/mol) showed hydrogen bonding interaction with Ala348, Asp350, Trp69, and Tyr385. The binding free energy of these interactions is strong.
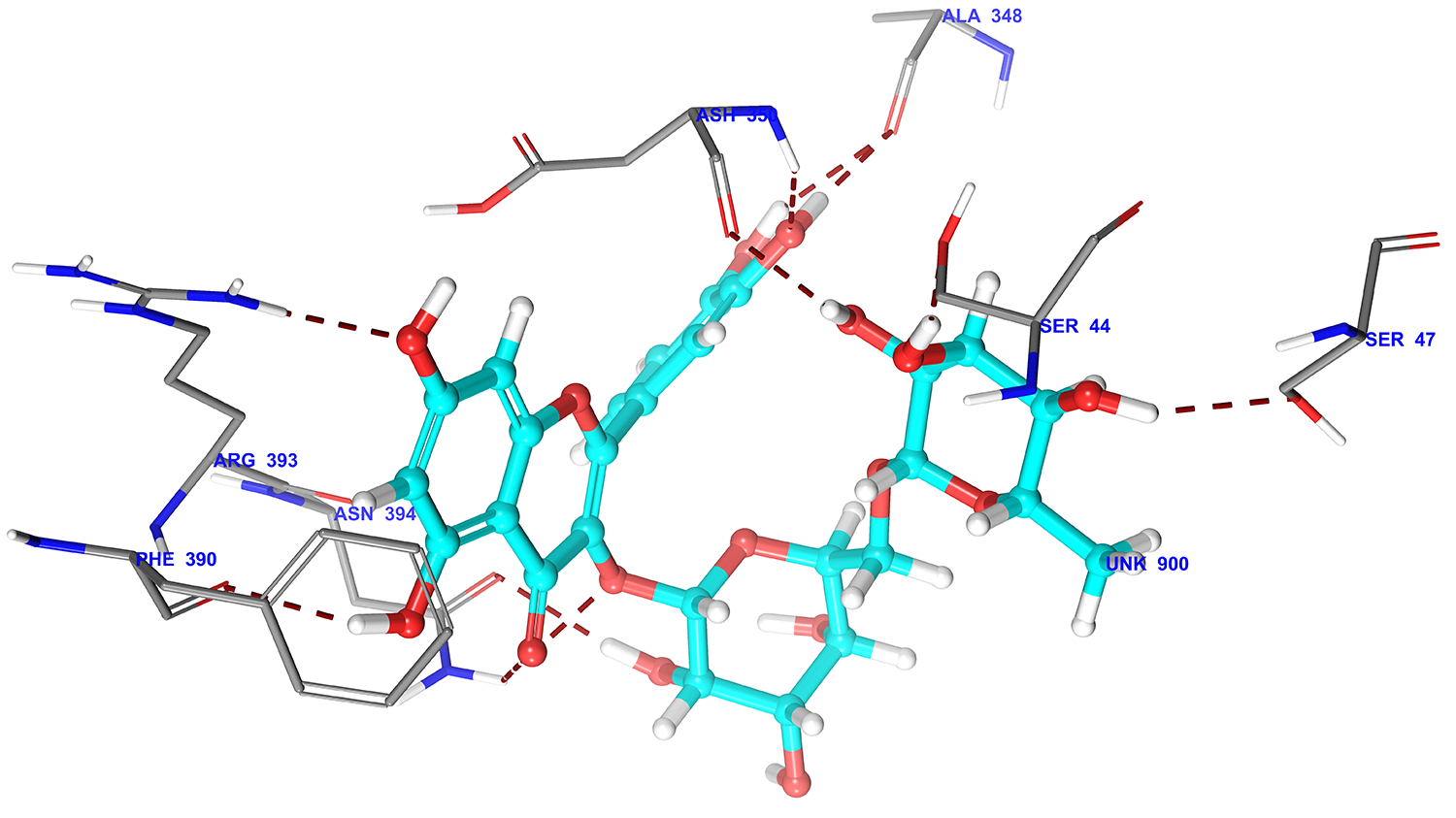 Fig. 2.
Fig. 2.3D docking pose of rutin into the ACE2 receptor catalytic domain (Pdbid: 2ajf) showing binding interaction with key amino acid residues of backbone protein. The backbone amino acid residues highlighted with grey colour. Binding interaction (hydrogen bonding) depicted with red dashed lines.
 Fig. 3.
Fig. 3.D Ligplot interaction of rutin against ACE2 receptor catalytic domain (Pdbid-2ajf). The pose was in 2D form and all the interactions were depicted by red solid lines, hydrophobic amino acid residue denoted by green colour, positive charge residue denoted by blue colour, charged negative residue denoted by red colour, polar residue were denoted by sky blue colour.
| LigandPhytoconstituent | ACE2 Docking scorekcal/mol | ACE2amino acids interactingwith the ligand | Binding free energykcal/mol | |
| Positive control | GL-1001 | -6.8 | Ala348, Trp349, Asp350, Asp382, Arg393 | -60.2 |
| Negative control | Captopril | -3.7 | Tyr385, ASN394 | -25.5 |
| 1 | Rutin | -11.5 | Ala348, Asp350, Arg393Ser44, Ser47, Phe390, Asn394 | -78.2 |
| 2 | Verbascoside | -10.2 | Ala348, Asp350, Asp382Glu375, His378, Phe390, Asn394, Glu402 | -90.4 |
| 3 | Hesperidin | -9.5 | Ala348, Asp350Trp69, Tyr385 | -70.3 |
| 4 | Luteolin 7-O-glucoside | -9.3 | Ala348, Arg393 Glu375, Phe390, Glu402 | -65.3 |
| 5 | Naringin | -9.2 | Ala348, Arg393 | -65.2 |
| 6 | Epigallocatechin gallate | -8.6 | Ala348, Asp350, Asp382, Arg393 Phe390 | -72.0 |
| 7 | Apigenin-7-O-glucoside | -8.1 | Ala348, Arg393 Glu375, Asn394 | -59.2 |
| 8 | Hesperetin | -7.3 | Ala348, Arg393 Phe390, Asn394 | -57.5 |
| 9 | Quercetin | -7.0 | Ala348, Arg393 Phe390 | -57.5 |
| 10 | Naringenin | -6.4 | Ala348 Phe390 | -51.1 |
| 11 | Dioscin | -6.4 | Asp350 | -57.0 |
| 12 | Kaempferol | -6.3 | Ala348, Arg393 Phe390, Asn394 | -52.6 |
| 13 | Apigenin | -6.2 | Ala348 Phe390 | -50.6 |
| 14 | Luteolin | -6.0 | Ala348, Arg393 Phe390 | -65.3 |
| 15 | Solanine | -6.0 | Ala348 Glu402 | -42.2 |
| 16 | Solamargine | -5.6 | Phe390 | -47.4 |
| 17 | Galangin | -5.3 | Arg393 Phe390, Asn394 | -44.8 |
| 18 | Acalyphin | -5.2 | Asp350, Arg393 Asn394 | -46.5 |
| 19 | Solasodine | -4.8 | Phe390 | -42 |
| 20 | Lantadene A | -4.5 | Ala348, Asp350 | -54.8 |
| 21 | Oleanolic acid | -4.4 | Ala348 Asn394 | -29.1 |
| 22 | Lantadene C | -4.0 | Asp350 | -52.2 |
| 23 | Lantadene B | -3.6 | Tyr369 | 50.0 |
| 24 | Ricinine | -3.2 | Tyr385 | -20.7 |
| 25 | Solasonine | -2.6 | None | -30.1 |
| ACE2 amino acids interacting with GL-1001, a selective ACE2 inhibitor, are given in bold. | ||||
ACE2-Phytoconstituent molecular docking studies revealed that Ala348 and Asp350 or Arg393 of ACE2 interact with several flavonoids and may be important in modulating ACE2 activity. Phe390 also interacts many phytoconstituents via hydrogen bonding. Recently, hesperidin has been proposed to be useful in prophylaxis for COVID-19 potentially inhibiting ACE2, cytokine storm and blood coagulation (Haggag et al., 2020; Huang et al., 2020; Wu et al., 2020). Phytoconstituents also interacts with Ala348, Arg393, Phe390, and Asn394. The interaction of Asp382 also observed with verbascoside and epigallocatechin gallate.
ACE2 catalytic site and viral S-protein binding site are reported to be distinct, and there may be an allosteric interaction between these two sites (Gross et al., 2020; Li et al., 2005; Rushworth et al., 2008). Some of the phytoconstituents may interact with both the catalytic and S-protein binding sites, whereas others may interact with one of the two sites. Further investigation of effect of phytoconstituents on the ACE2 catalytic and viral entry into cells using cell-based functional assays may provide insights into phytoconstituents that may be useful for the treatment of COVID-19.
Encouraged by the results of the phytochemicals on ACE2 interaction, we carried out the docking studies with AT1 and AT2 receptors. To confirm our docking approach, we have used AT1 antagonist, Olmesartan, and AT2 antagonist, PD12377, as pharmacological controls. Olmesartan showed a docking score of -8.9 kcal/mol and -6.5 at AT1, and AT2 receptors, respectively. On the other hand, PD12377 showed a docking score of -4.9 and -6.7 kcal/mol at AT2 receptors, respectively, validating our docking approach.
The compounds solanine, rutin, quercetin, hesperetin, epigallocatechin gallate, apigenin-7-O-glucoside, luteolin-7-O-glucoside, verbascoside, hesperidin and naringin showing good binding affinity and docking score better than -6.0 kcal/mol against ACE2, were further selected for docking against AT1and AT2 receptors (Table 2).
| Ligand-Phytoconstituent | AT1 Docking score kcal/mol | Binding free energy against AT1kcal/mol | AT1 amino acids interacting with the ligand | AT2 Docking score kcal/mol | AT2 amino acids interacting with the ligand | Binding free energy against AT2kcal/mol | |
| OlmesartanAT1 antagonist | -8.9 | -84.3 | Tyr35, Trp84, Thr88, Arg167 | -6.5 | Arg182,Lys215, Trp100 | -78.3 | |
| PD12377AT2 antagonist | -4.9 | -85.6 | Tyr87, Tyr92 | -6.7 | Trp100, Phe129, Arg182, Lys215, Phe272 | -82.9 | |
| 1 | Rutin | -12.6 | -95.2 | Tyr35, Thr88, Arg167 Tyr87, Phe182, Tyr184, Asp263, Gln267 | -11.9 | Arg182, Lys215, Phe272 Ser208, Asp279 | -91.5 |
| 2 | Verbascoside | -12.3 | -91.7 | Arg167 Tyr184,Trp253, Gln257 Asp263 | -11.6 | Arg182, Lys215 Tyr108, Thr276, Asp279, Trp283 | -94.8 |
| 3 | Hesperidin | -11.6 | -90.3 | Tyr35 Ser16, Cys18, Ala21, Tyr87 | -10.2 | Arg182, Lys215 Thr276, Asp279 | -91.0 |
| 4 | Luteolin 7-O-glucoside | -10.5 | -76.8 | Tyr35 Cys180, Tyr292 | -9.7 | Arg182, Lys215 Thr125 | -68.0 |
| 5 | Naringin | -10.7 | -84.1 | Tyr35, Thr88, Arg167 Pro19, Ala21, Arg23, Tyr87 | -10 | Lys215 Cys195, Thr276, Asp279 | -83.9 |
| 6 | Epigallocatechin gallate | -7.2 | -72.8 | Arg167 Ala21, Arg23, Tyr87, Tyr92, Asp263 | -9.4 | Lys215, Phe272 Thr178 | -78.2 |
| 7 | Apigenin-7-O-glucoside | -11.1 | -74.9 | Arg167 Ala21, Tyr87, Asp281 | -10.2 | Arg182 Tyr51, Thr125 | -74.3 |
| 8 | Hesperetin | -8.0 | -50.5 | Arg167 Tyr87 | -7 | Lys215, Phe272 Thr178 | -57.7 |
| 9 | Quercetin | -8.4 | -55.3 | Tyr35 Ala21, Tyr87, Tyr92 | -7.5 | Lys215, Phe272 Thr178 | -54.2 |
| 10 | Naringenin | -8.0 | -45.5 | Arg167 Tyr87 | -7.7 | Thr125 Arg182 | -50 |
| 11 | Dioscin | -10.0 | -58.8 | Tyr87 | -8.2 | Thr125 | -54.2 |
| 12 | Kaempferol | -7.0 | -46.5 | Tyr35, Arg167 Tyr87 | -7.3 | Phe272 Thr125, Ile304 | -50.3 |
| 13 | Apigenin | -7.8 | -45.3 | Arg167 | -7.7 | Arg182 Thr125 | -48.5 |
| 14 | Luteolin | -9.0 | -47.4 | Arg167 | -7.3 | Lys215, Phe272 Thr178 | -52.8 |
| 15 | Solanine | -8.4 | -33.5 | None | -8.9 | Arg182 | -50.1 |
After docking against the AT1 and AT2 receptor of selected compounds, we found all compounds showed good binding affinity against both the receptors, which is higher than -7.0 kcal/mol. Moreover, rutin secured a high docking score among all the compounds in docking study against AT1 and AT2 receptors. It forms hydrogen bond interactions with AT1 receptor protein through Tyr35, Thr88, Arg167, Asp263, Tyr184, Gln267, Phe182, and while making pi-pi stacking interaction with Tyr87. The docking score of rutin against the AT1 receptor is -12.60 kcal/mol (Fig. 4 and 5). Interestingly, the docking results of selected compounds at the AT2 receptor also show a high docking score of rutin because of its ability to form hydrogen bonding with amino acid residues like Ser208, Asp279, Lys215, Arg182, and pi-pi stacking with the Phe272. The docking score of the highest-scoring compound rutin against the AT2 receptor was -11.9 kcal/mol (Fig. 6 and 7).
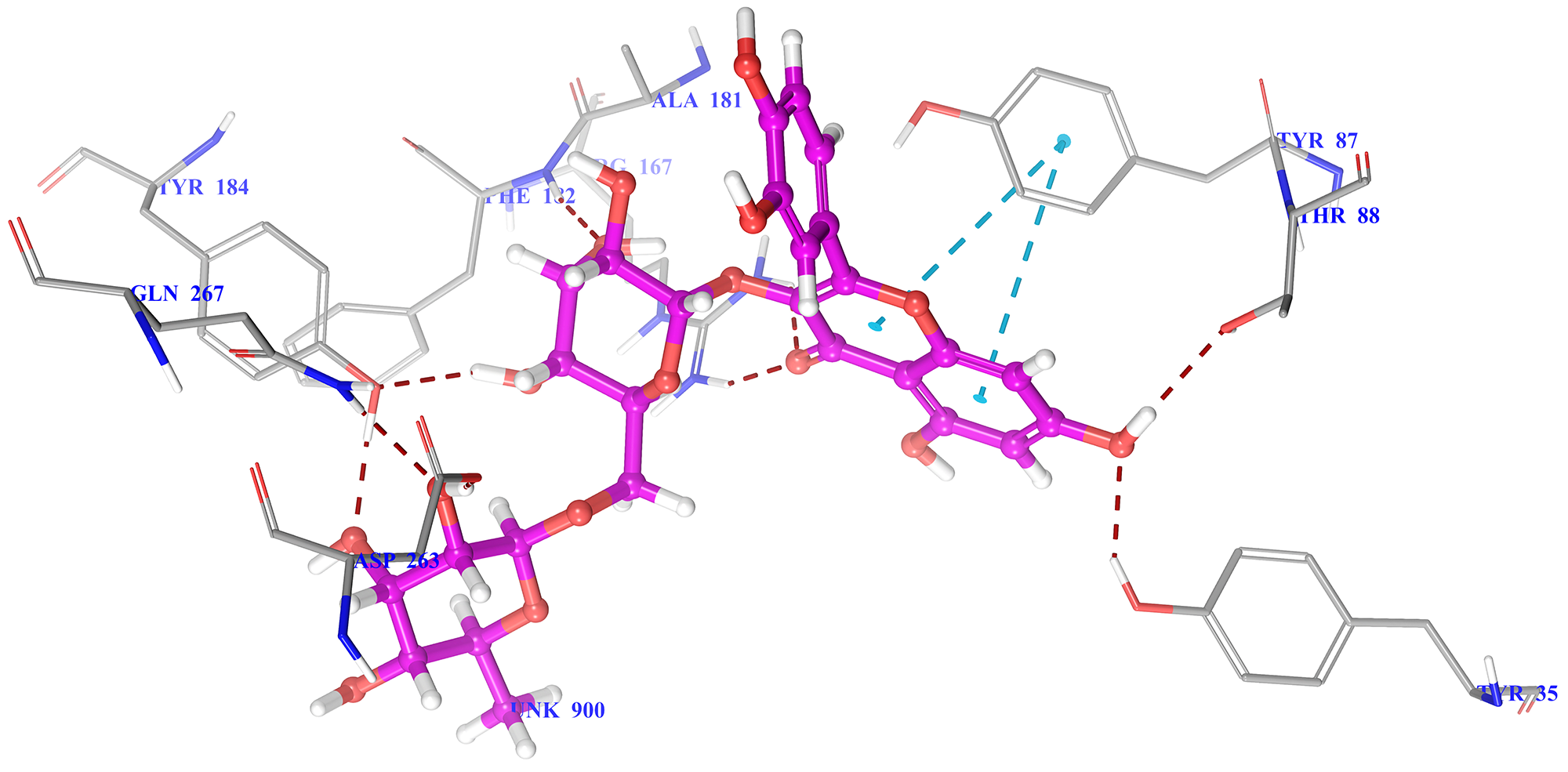 Fig. 4.
Fig. 4.3D docking pose of rutin into the AT1 receptor (Pdbid-4zud) showing binding interaction with key amino acid residues of backbone protein. The backbone amino acid residues highlighted with grey colour while rutin highlighted by green colour. Binding interaction (hydrogen bonding) depicted with red dashed lines. Pi-pi stacking interaction was denoted by blue dashed lines.
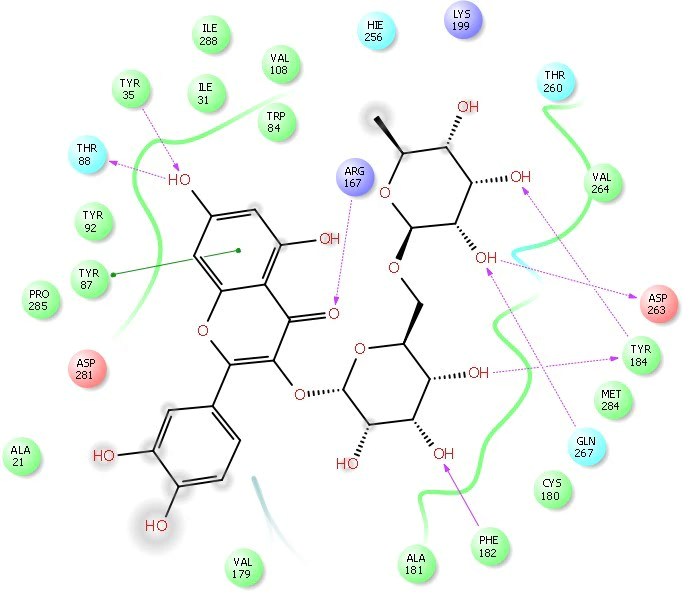 Fig. 5.
Fig. 5.2D Ligplot interaction of rutin docked in to the AT1 receptor catalytic domain (Pdbid-4zud). The pose was in 2D form and all the interactions were depicted by red solid lines, hydrophobic amino acid residue denoted by green colour, positive charge residue denoted by blue colour, charged negative residue denoted by red colour, polar residue were denoted by sky blue colour.
Interestingly, the docking results of selected compounds at the AT2 receptor also show a high docking score of rutin because of its ability to form hydrogen bonding with amino acid residues like Ser208, Asp279, Lys215, Arg182, and pi-pi stacking with the Phe272. The docking score of the highest-scoring compound rutin against the AT2 receptor was -11.9 kcal/mol (Fig. 6 and 7).
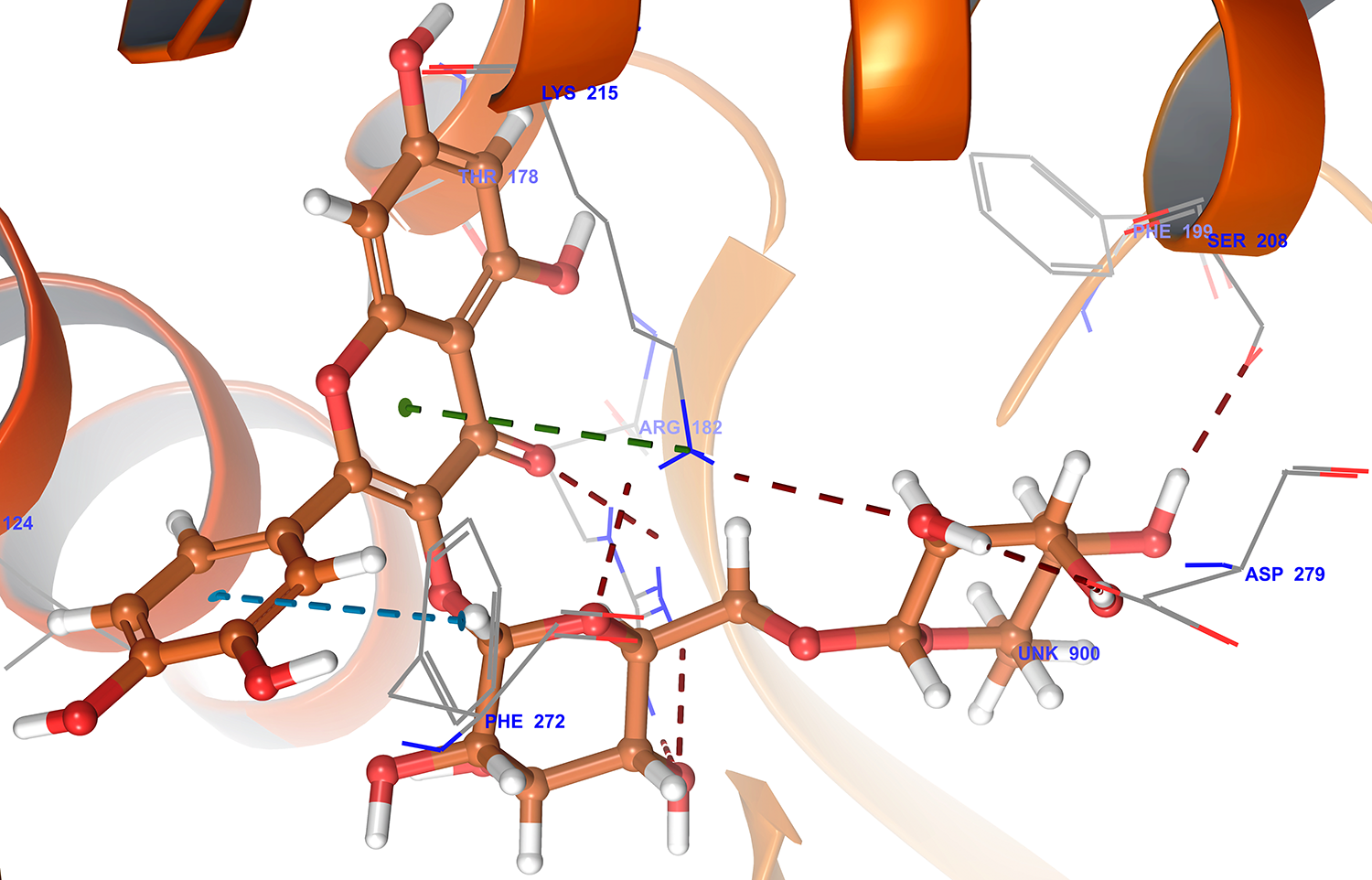 Fig. 6.
Fig. 6.3D docking poses of rutin in to the AT2 receptor catalytic domain (Pdbid-4unf) showing binding interaction with key amino acid residues of backbone protein. The backbone amino acid residues highlighted with grey colour while rutin highlighted by brown colour. Binding interaction (hydrogen bonding) depicted with red dashed lines. Pi-pi stacking interaction was denoted by blue dashed lines.
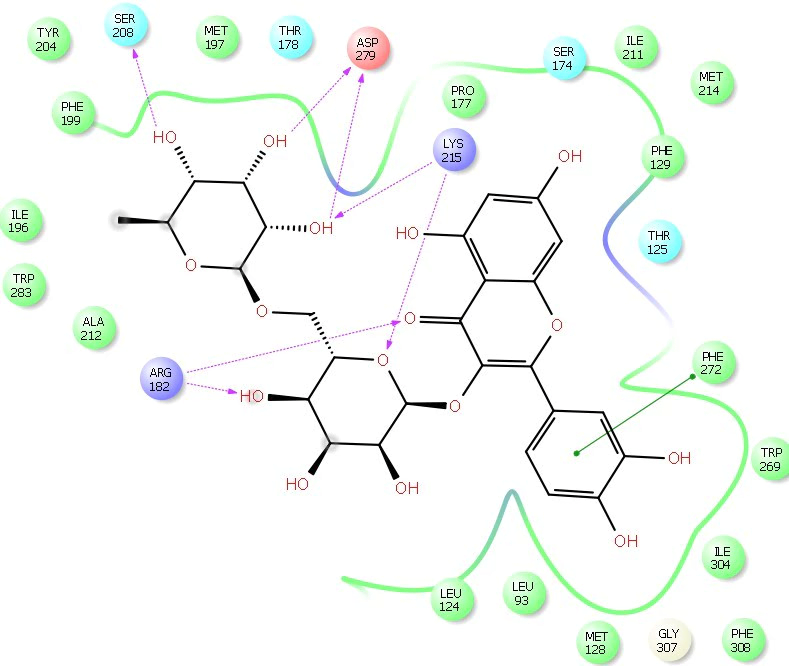 Fig. 7.
Fig. 7.2D Ligplot interaction of rutin docked in to the AT2 receptor catalytic domain (Pdbid-4unf). The pose was in 2D form and all the interactions were depicted by red solid lines, hydrophobic amino acid residue denoted by green colour, positive charge residue denoted by blue colour, charged negative residue denoted by red colour, polar residue were denoted by sky blue colour.
In the docking study results almost all the phytoconstituents showed similar binding interaction with Tyr35 and Arg167 as shown by the standard at AT1 receptor. However, most of the phytoconstituents showed some marked affinity towards Arg182 and Lys215 amino acids at AT2 receptor in a same way as depicted by the standard.
Docking studies revealed that phytoconstituents interacts with ACE2, AT1 and AT2 receptors of the RAS system. Both ACE and ACE2 are involved in the hydrolysis of bradykinin which also contributes to disturbance in RAS homeostasis, inflammatory response and COVID-19 pathology. Inhibition of ACE and ACE2 causes increase in the circulating levels of bradykinin. Moreover, neprilysin mainly due its hydrolytic action on bradykinin and angiotensin 1-7 modulates RAS. Considering that there is no approved bradykinin receptor antagonist and neprilysin inhibitor, evaluating the action of phytoconstituents on bradykinin and neprilysin pathways will provide a more comprehensive understanding of RAS homeostasis, and its modulation in inflammatory diseases and development of therapeutics (Roche and Roche, 2020).
Upon entry into host, SARS-CoV2 triggers both innate and adaptive immune responses with the accumulation of activated monocytes, macrophages, and neutrophils, leading to uncontrolled inflammation due to the release cytokines and chemokines (Del Valle et al., 2020; Nathan, 2020; Sallenave and Guillot, 2020; Vabret et al., 2020). This cytokine storm causes severe lung inflammation accompanied by inflammatory damage to multiple-organs, and vascular thrombosis, which causes death in severely ill COVID-19 adult patients and children (Capone et al., 2020; Crisafulli et al., 2020; Jose and Manuel, 2020; Mahmudpour et al., 2020; Mehta et al., 2020; Rowley, 2020; Soy et al., 2020; Ye et al., 2020). Kinase inhibitors (baricitinib, imatinib, dasatinib, tramitinib, selumetinib) due to their anti-inflammatory actions approved for other inflammatory diseases are being investigated for COVID-19 (Dyall et al., 2014; Richardson et al., 2020a; Schrader et al., 2018; Stebbing et al., 2020; Troiani et al., 2012; Zhang et al., 2020). Anticoagulants (warfarin, dabigatran, betrixaban, and rivaroxaban) are also being evaluated for the treatment of thrombosis in COVID-19 patients (www.clinicaltrials.gov). The main challenge faced by the clinicians while using anticoagulant drugs in treating COVID-19 patients is the possible interaction of anticoagulants with most antiviral drugs.
RNA-dependent RNA polymerase (RdRp) in the viral nonstructural protein 12 (nsp12) has been a target for drug development. Many studies are evaluating nucleoside and nucleotide analogs for the repurposing. Based on the initially promising clinical findings, remdesivir has recently been approved by the FDA for the treatment of COVID-19. It is a broad-spectrum antiviral drug initially developed by Gilead Sciences in 2009 for hepatitis C. It did not work against hepatitis C and then repurposed for Ebola virus (Jacobs et al., 2016). It is effective against many other viruses, including coronavirus (Lo et al., 2017). Remdesivir, an adenosine nucleoside triphosphate analog, is metabolized in the cell, and the active remedisvir metabolite interferes with the viral RdRp that evades proofreading by the viral exoribonuclease (ExoN) in nsp14, ultimately causing a decrease in viral RNA production (Ferner and Aronson, 2020). The RdRp of MERS-CoV, SARS-CoV-1, and SARS-CoV-2 arrest of RNA synthesis occurs after the incorporation of three additional nucleotides (Lo et al., 2017). Hence, remdesivir is a direct-acting antiviral agent that works as a delayed chain terminator (Tchesnokov et al., 2019). Remdesivir treatment produced clinical improvement in two-third of patients, but the mortality rate was similar to the placebo group (Antinori et al., 2020; Beigel et al., 2020; Grein et al., 2020; Wang et al., 2020). The antiviral activity of remdesivir is not only marginal but also variable.
In search of other potential antiviral drugs, a randomized, controlled, open-label trial was conducted among COVID-19 patients with lopinavir-ritonavir (400 mg and 100 mg, respectively) for 14 days (Cao et al., 2020). But no benefit was observed with lopinavir-ritonavir treatment beyond standard care. Nafamostat mesylate, a serine protease inhibitor showed promising effect e against SARS-CoV-2 infection of lung cells (Hoffmann et al., 2020). Ribavirin also supports its potential to significantly impact the treatment of SARS-CoV-2 infections due to low cost and wide availability. An adjuvant therapy of Oseltamivir gave promising results in patients coinfected with novel coronavirus 2019 and influenza virus in Wuhan, China (Ding et al., 2013). Penciclovir, an acyclic nucleoside analog, showed marked efficacy when used in combined antiviral therapy against SARS-CoV-2 infection. In an open-label control study, the favipiravir exerted better treatment effects on COVID-19 patients (Cai et al., 2020). Other antiviral drugs like Ganciclovir, Tenofovir, Dolutegravir, Atazanavir, Asunaprevir, Elbasvir, and Sofosbuvir are being proposed and tested but not found so effective against SARS-CoV-2 (Tan et al., 2004). Hence, it remains an unmet need for an efficacious and safe RdRp inhibitor.
Besides antivirals, clinical investigations are ongoing to repurpose non-antiviral drugs, such as Rapamycin and Tiotropium, which may act by inhibiting the replication of SARS-CoV-2 like in the case of HIV (Nicoletti et al., 2011). The observed hyperactivation of the immune response in COVID-19 involves dysregulation of mTOR molecule. mTOR plays a critical role in immune-regulation and immune-tolerance control pathways. The ability of mTOR inhibitors to suppress early stages of viral replication, reduce mRNA translation, ribosome biogenesis, protein synthesis, mitochondrial metabolism, and viral replication suggest potential clinical benefit from mTOR inhibitors (Ramaiah, 2020).
Targeting cellular mTOR results in antiviral action in HIV infection, specific
mTOR inhibitors, including Rapamycin and Everolimus, may have a beneficial
outcome in COVID-19 subjects. (Fagone et al., 2020; Heredia et al., 2015; Nicoletti et al., 2011; Zheng et al., 2020). Moreover, using the molecular
transformer-drug-target interaction prediction model, (Kang et al., 2020)
showed that SARS-CoV-2 infection strongly activates cytokine and immune system,
including upregulation of TNF and NF
For the identification of phytoconstituents effective at RdRp, docking studies were performed using the Glide module of the Schrodinger-9.6 software on the RdRp (PDB id: 7bv2). We have used nucleoside substrates uridine triphosphate (UTP), and thymidine triphosphate (TTP) as controls. UTP being the natural substrate for RdRp, interacted with RDRp at 5 amino acid residues, namely Lys551, Arg553, Arg555, Lys621 and Asp760 with a docking score - 6.7 kcal/mol. Whereas TTP interacts with RdRp only with Arg 553 and Cys622.
Remdesivir is a prodrug that is hydrolyzed to yield the active metabolite. The
molecular docking of remdesvir metabolite triphosphate (RemTP) into the active
site of the RdRp enzyme reveals that it occupies the same binding pocket with
root mean square deviation (RMSD) of 0.44
| Ligand Phytoconstituent | RdRp Docking score kcal/mol | Binding free energy against RdRp kcal/mol | RdRp amino acids interacting with the ligand | ORF8 Docking score kcal/mol | ORF8 amino acids interacting with the ligand | Binding free energy against ORF8 kcal/mol | |
| Uridine Triphosphate | -6.7 | -56.8 | Lys551, Arg553, Arg555, Lys621, Asp760 | ND | ND | ND | |
| Thymidine derivative | -6.5 | -53.2 | Arg553, Cys622 | ND | ND | ND | |
| Remedisvir | -6.7 | -61.5 | Lys551, Arg555 | ND | ND | ND | |
| metabolite | Ser549 | ||||||
| 1 | Rutin | -9.0 | -90.6 | Arg555 | -10.9 | Tyr42, His45, His58, Lys61, His87 | -68.9 |
| Ile548, Arg553, Asp623, Asp760 | |||||||
| 2 | Verbascoside | -7.4 | -132.7 | Lys551, Arg555, | -9.9 | Lys61, Lys90 | -72.7 |
| Asp760 | |||||||
| Asp618, Cys622 | |||||||
| 3 | Hesperidin | -10.0 | -98.1 | Arg555 | -14.5 | Phe43, His45, His58, Lys61, His87, Asn97 | -97.5 |
| Lys545, Ile548, Asp845 | |||||||
| 4 | Luteolin 7-O-glucoside | -8.5 | -102 | Ile548, Ser759, Arg836 | -13.1 | His58, His87, Tyr42 | -74.4 |
| 5 | Naringin | -9.6 | -90.9 | Asp760 | -11.2 | His58, His87, Lys90 | -63.2 |
| Ser549, Tyr619, Asp623, | |||||||
| 6 | Epigallocatechin gallate | -5.2 | -103.7 | Ile548, Ser814 | -11.6 | Tyr42, His58 | -54.2 |
| 7 | Apigenin-7-O-glucoside | -8.6 | -90.7 | Arg555, Asp760 | -12.6 | Tyr42, His58, His87 | -74.8 |
| Ile548 | |||||||
| 8 | Hesperetin | -6.1 | -61.5 | Arg555 | -9.7 | His87 | -51.5 |
| Ser682 | |||||||
| 9 | Quercetin | -7.8 | -62.6 | Arg555 | -10.5 | His87 | -53.7 |
| Ser682 | |||||||
| 10 | Naringenin | -6.4 | -43.6 | Ser682 | -9.3 | His58, His87 | -46.9 |
| 11 | Dioscin | -5.9 | -100 | Tyr619 | -4.2 | Lys90 | -40.1 |
| 12 | Kaempferol | -5.7 | -53.7 | Arg555 | -9.8 | His87 | -50.1 |
| 13 | Apigenin | -5.5 | -52.1 | Arg555 | -9.0 | His87 | -52.1 |
| Ser682 | |||||||
| 14 | Luteolin | -6.4 | -54.1 | Arg555 | -10.0 | His87 | -52.2 |
| Ser682 | |||||||
| 15 | Solanine | -6.1 | -102.9 | Ser682 | -5.5 | His87 | -51.2 |
| 16 | Solamargine | -10.8 | -72.8 | Tyr619 | -4.7 | Trp42 | -43.2 |
| 17 | Galangin | -5.6 | -52.1 | Arg555 | -9.8 | Tyr42, His58, His87 | -51.2 |
| 18 | Acalyphin | -5.6 | -46.6 | Asp760 | -6.6 | His87 | -37.5 |
| Asp618, Lys621 | |||||||
| 19 | Solasodine | -4.2 | -58.6 | Asn691 | -2.0 | Leu91 | -16.9 |
| 20 | Oleanolic acid | -3.0 | -58.2 | Ser549, Tyr619 | -1.2 | Lys90 | -69.2 |
| SARS-CoV-2 RdRp amino acids interacting with uridine triphosphate, are given in bold. | |||||||
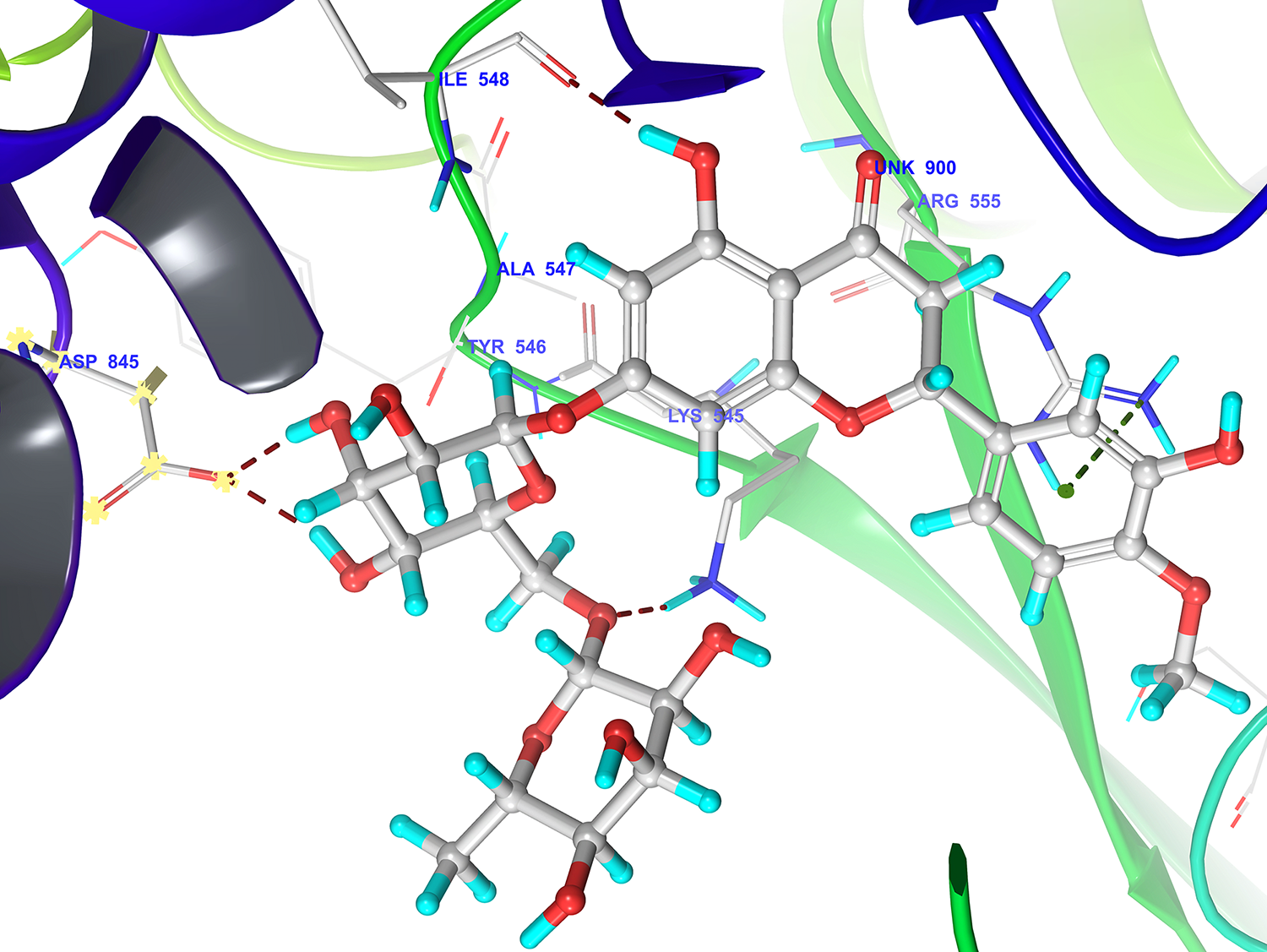 Fig. 8.
Fig. 8.3D docking pose of hesperidin in to the RdRp receptor catalytic domain (Pdbid-7bv2) showing binding interaction with key amino acid residue of backbone protein. The backbone amino acid residues highlighted with grey colour while hesperidin highlighted by grey colour, Binding interaction (hydrogen bonding) depicted with red dashed lines. Ionic interaction was denoted by green dashed lines.
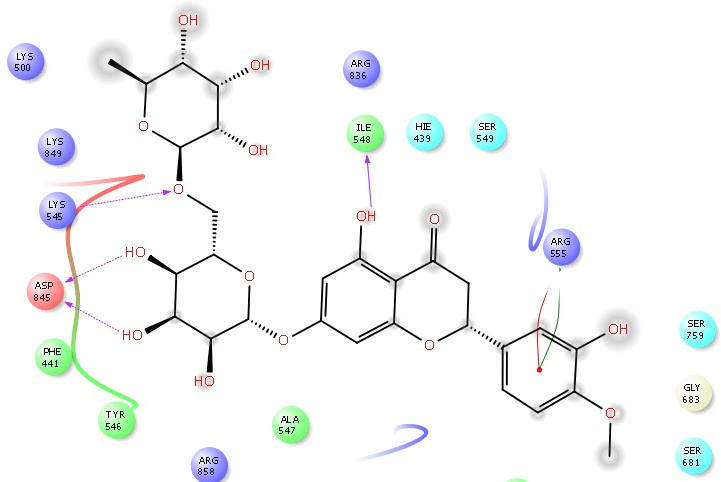 Fig. 9.
Fig. 9.2D Ligplot interaction of hesperidin docked in to the RdRp receptor catalytic domain (Pdbid-7bv2). The pose was in 2D form and all the interactions were depicted by violet solid lines, hydrophobic amino acid residue denoted by green colour, positive charge residue denoted by blue colour, charged negative residue denoted by red colour, polar residue were denoted by sky blue colour.
Open reading frame 8 (ORF8) is one of the accessory proteins present in beta coronaviruses, including SARS-CoV-2. All animal SARS-CoVs and human SARS-CoVs affected by early diseases contain full-length ORF8 (Song et al., 2005). However, in the mid and lately affected patients, there is a deletion of 29-nucleotide, resulting in the splitting of full-length ORF8 into putative ORF8a and ORF8b (Oostra et al., 2007). In SARS-CoV-2, the ORF8 protein does not contain a known region or motif and shares the least homology with ORF8 of SARS-CoV. The viral protein encoded from ORF8 interacts with MHC-I and downregulates their surface expression, a function that is absent with ORF8a and ORF8b of SARS-CoV. Thus, ORF8 of SARS-CoV-2 seems more lethal as it also disrupts antigen presentation and reduction of recognition and elimination of viruses infected cells by cytotoxic T lymphocytes. Hence, in this review, we wish to throw some light concerning potential agents that can inhibit ORF8.
The sequencing of SARS-CoV-2 RNA revealed the coding sequences for structural and nonstructural proteins. Structural and molecular docking studies reported that the nonstructural protein encoded from open reading frame 8 (ORF8) of SARS-CoV-2 could combine with porphyrin to form a complex, facilitating the dissociation of the iron form heme (Liu and Li, 2020), resulting in reduced oxygen-carrying capacity. ORF8 also interacts with MHC-I molecules directly and significantly downregulates the expression of MHC-1 on various cell types. MHC-I molecules selectively target ORF8 expressing cells for lysosomal degradation. Downregulation of MHC-1 reduces the recognition and the elimination of virus-infected cells by CTLs (Zhang et al., 2020). Therefore, the inhibition of ORF8 may enhance immune surveillance and accelerate the removal of SARS-CoV-2 infected cells and enhance the oxygen-carrying capacity of the blood. Currently, there is no approved drug for specific drug targeting of ORF8, and our findings with phytoconstituents may stimulate research efforts on the ORF8 target protein.
Molecular docking studies were performed against ORF8 using structural protein with PDB ID: 2AJF and 6BB5 (Table 3). Docking results reveal that the compounds luteolin-O-glucoside and hesperidin bound tightly to an active site of ORF8. The compounds luteolin-O-glucoside and hesperidin interacts with Tyr42, Phe43, His58, Lys61, His87 and Asn97. These compounds showed docking scores of -13.06 and -14.45kcal/mol, respectively, against ORF8 (Fig. 10 and 11), and provide insights into developing ORF8 inhibitors.
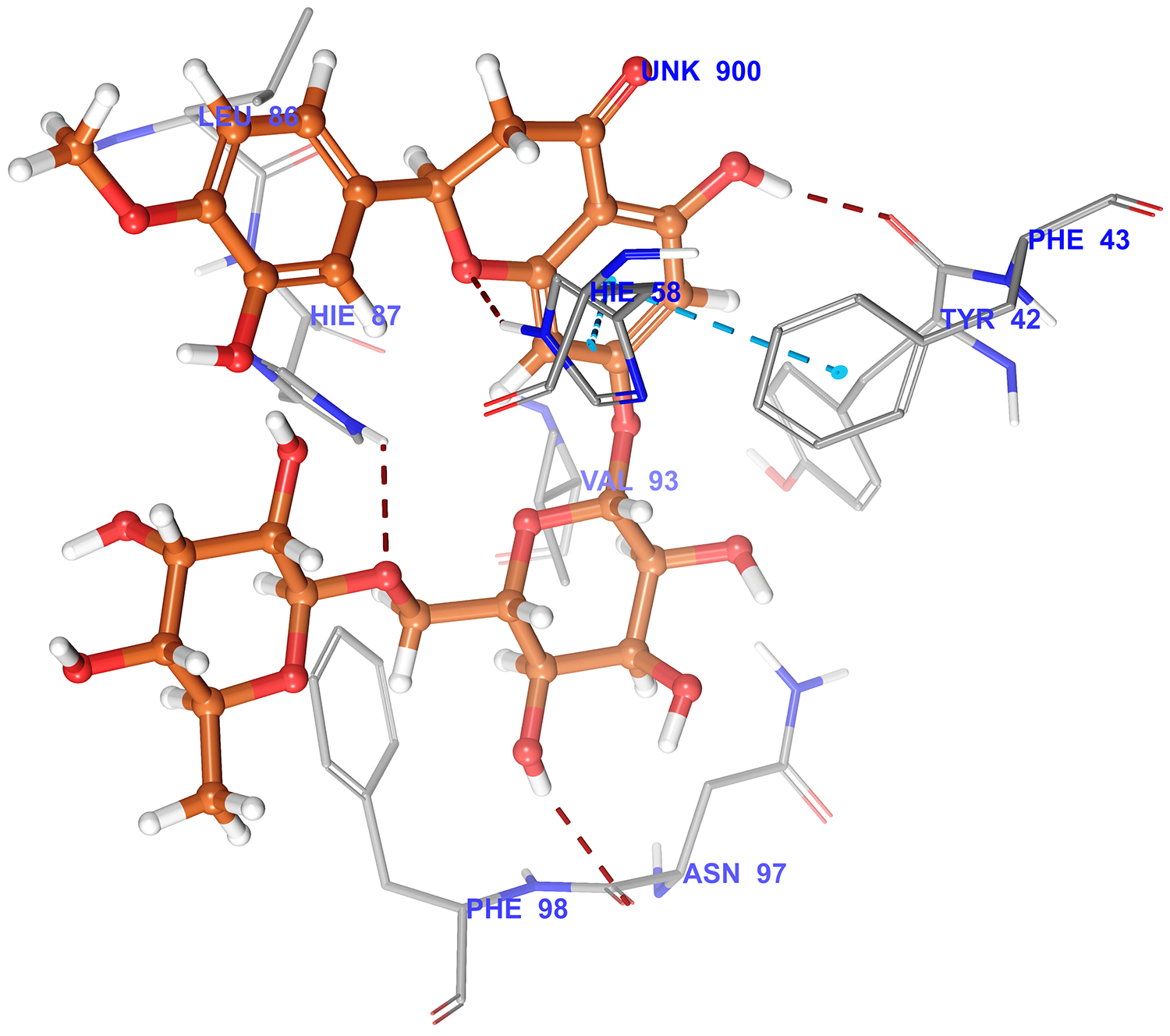 Fig. 10.
Fig. 10.3D docking pose of hesperidin in to the ORF8 receptor catalytic domain (Pdbid-6bb5) showing binding interaction with key amino acid residue of backbone protein. The backbone amino acid residues highlighted with grey colour while hesperidin highlighted by brown colour Binding interaction (hydrogen bonding) depicted with red dashed lines. pi-pi interaction was denoted by blue dashed lines.
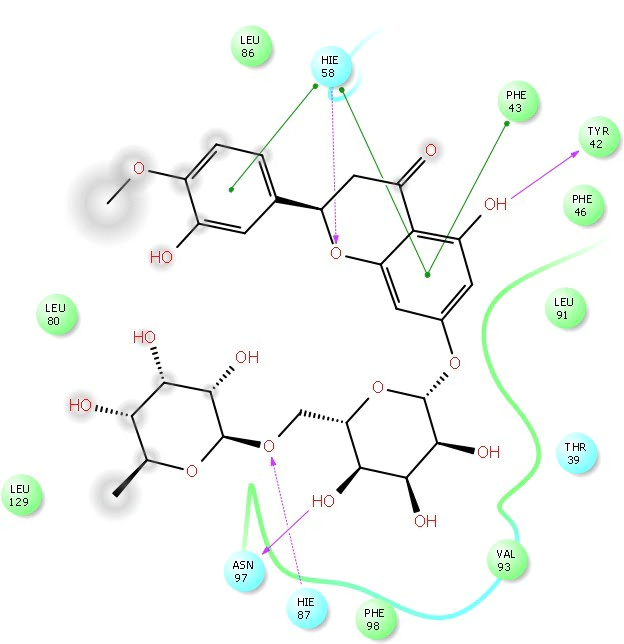 Fig. 11.
Fig. 11.2D Ligplot interaction of hesperidin docked in to the ORF8 receptor catalytic domain (Pdbid-6bb5). The pose was in 2D form and all the interactions were depicted by violet solid lines, hydrophobic amino acid residue denoted by green colour, positive charge residue denoted by blue colour, charged negative residue denoted by red colour, polar residue were denoted by sky blue colour.
Although the efficacy of phytoconstituents and herbal plants in reducing the SARS-CoV-2 severity is under investigation, a systematic meta-analysis of Chinese Herbal Medicines suggests that herbal medicines may improve clinical outcomes when used in conjunction with conventional medical care (Fan et al., 2020). Moreover, a single case study reported a combination of herbal medicines comprising of several herbal plants enabled the patient to recover from COVID-19 (Girija and Sivan, 2020). Herbal medicines may be useful for the prevention, treatment of mildly symptomatic, severe, and recovering COVID-19 patients (Lee et al., 2020). Moreover, over eighty clinical investigations involving herbal medicines (ctri.nic.in; (www.clincialtrials.gov)) are in the registry for investigating the clinical benefits of phytoconstituents, and results from these studies will provide confirmation of clinical efficacy of phytoconstituents in treating COVID-19.
Naturally occurring phytochemicals are a powerful resource of chemical compounds to discover therapeutics for COVID-19 treatment. Molecular docking studies provided in this review and elsewhere offer insights into the interaction of phytoconstituents with the host or viral proteins. However, the studies on the functional implications of the molecular interactions on the target proteins in vitro and in vivo would shed light on the ability of these compounds to activate or inhibit their target proteins. Furthermore, insights into the potency, selectivity, mode of action, and therapeutic index for each phytochemical are required for testing in clinical trials. Nonetheless, these molecular docking studies provide insights for developing phytochemical-based therapeutics or designing new chemical entities (NCEs) suitable for generating safe and efficacious therapeutics from phytochemicals.
The pathogenesis and prognosis of COVID-19 are complex, involving inflammatory damage to multiple organs, including lung, heart, kidney, and vasculature. Besides serving as a cell entry site for SARS-CoV-2, ACE2 plays a very critical role in regulating the balance between the ACE2 driven anti-inflammatory actions and ACE/angiotensin AT1 receptors driven proinflammatory effects. Currently, there are no approved ACE2 inhibitors for preventing the entry of SARS-CoV-2 into host cells. Docking studies revealed phytoconstituents that interact with ACE2. Although the functional implications of these phytoconstituents on ACE2 activity are yet to be confirmed, available evidence suggests that inhibition of ACE2 may be a pivotal step to inhibit SARS-CoV-2 virus infection and consequent inflammatory damage. Phytoconstituents may interact with catalytic and/or S-protein binding sites. Further investigation of effect of phytoconstituents on the ACE2 catalytic and viral entry into cells using cell-based functional assays may provide insights into phytoconstituents. Furthermore, interaction of phytoconstituents at ACE2, AT1, and AT2 receptors suggests that the phytoconstituents may suppress the imbalance between ACE’s counterregulatory functions by interacting with both arms of RAS. Phytoconstituents also may have inhibitory actions on SARS-CoV-2 replication and improving the oxygen-carrying capacity of hemoglobin. Thus, phytoconstituents may have multiple sites of action, protecting and countering the inflammatory damage. Although the use of ACE inhibitors and AT1 receptor blockers during COVID-19 remains controversial, more recent clinical observations suggest that ACE inhibitors and AT1 blockers are not harmful in patients who had been taking the medications. Phytoconstituents may be useful both as prophylactic and treatment for COVID-19. Moreover, COVID-19 patients who are severely affected and admitted to the hospital, particularly those treated in intensive care units, often have baseline comorbidities, including hypertension, diabetes, and obesity. These diseases are associated with strong activation of both the circulating and the local RAS, further complicating COVID-19 pathology. Although phytoconstituents may suppress both arms of RAS, severely ill COVID-19 patients may benefit from the diverse modulatory action of these phytoconstituents on the host RAS, viral RdRp and viral ORF8 targets.
Ramesh K. Goyal: Concept and skeleton of the Review Paper
Jaseela Majeed: Target Identifications
Rajiv Tonk: Over view of repurposed drugs and their meachanisms
Mahaveer Dhobi: Herbal drugs and Phytoconstituents identification
Bhoomika Patel: Pharmacological and mechanism of action
Kalicharan Sharma: Docking studies with Rajiv Tonk and Jaseela Majeed
Subbu Apparsundaram: Clinical aspects and correlation from new drug discovery point of view
Authors acknowledge Nirma University, Ahmedabad and Mankind Research Center, Gurgaon, Haryanam India for their cooperation.
Authors declare that there is no conflict of interest regarding the publication of this article.
