- Academic Editors
Objective: Cervical cancer (CC) tops the list as the most prevalent malignant tumors of the female reproductive system and is one of the leading causes of cancer-related deaths in women worldwide. To improve the prognosis of CC, research has consistently focus on understanding innovative approaches. Biomarker-guided precision medicine is an important direction to improve prognosis. In this paper, we aim to deepen our understanding of CC biomarkers and accelerate their translation into clinical practice. Mechanism: Literature studies on biomarkers for CC were screened from 23 May 2019 to 23 May 2023. A comprehensive search was performed in NIH/NCBI/PubMed databases. Findings in Brief: There are numerous biomarkers that have a significant role in the diagnosis, prognosis, and treatment of CC. They can influence various processes such as tumour proliferation, invasion, infiltration, and apoptosis. They are also related closely to the sensitivity and resistance of radiotherapy and chemotherapy. Conclusions: To date, multiple CC biomarkers have been identified. Few biomarkers have been successfully applied in clinical practice due to the lack of validation in large-scale clinical studies. Therefore, further clinical studies are necessary to realize the great potential in the clinic.
Among the top five causes of cancer deaths in the United States by age and sex in 2020, cervical cancer (CC) had the second-highest death rate among females aged 20–39 years [1]. However, advancements in prevention and screening have contributed to a decline in CC rates. In fact, the incidence of CC has decreased by more than 50% since the mid-1970s, primarily due to widespread screening usage. Furthermore, the Global Cancer Statistics 2023 report also reveals a 65% decline in CC incidence among women in their early 20s who received the human papillomavirus (HPV) vaccine between 2012 and 2019 [1].
Despite the advancements in cancer diagnosis, early detection of CC remains challenging due to its often asymptomatic and nonspecific symptoms during the initial stages. Current clinical practice relies on two main screening methods: cervical cytology (Pap smear) and HPV testing, both of which have their limitations. The Pap smear primarily detects abnormal cell changes through the examination of cell morphology [2]. On the other hand, HPV testing offers several clinical applications, including screening women with ambiguous cytologic results, predicting postoperative treatment outcomes, and primary screening for CC and precancerous lesions [3]. By detecting HPV DNA or RNA, HPV testing provides higher sensitivity and a broader application range compared to the Pap smear. Thus HPV DNA testing has been approved as a primary screening test, either alone or in combination with cytology. A study by Voidăzan et al. [4] highlights the potential of biomarkers, such as p16/Ki-67, to enhance the sensitivity and specificity of CC screening. Rokos et al. [5] conducted a comprehensive review of 25 studies on the role of microRNA-21 (miR-21) in CC. Their findings suggest that miR-21 is associated with tumorigenesis and lymphatic metastasis in CC, and it also has an impact on chemoresistance. Hence, the detection of miR-21 presents a potential avenue for enhancing early detection and treatment of CC [5]. These biomarkers can aid in identifying high-grade lesions that may be overlooked by Pap smear alone. Furthermore, when combined with HPV testing, they can enhance the sensitivity and specificity of the diagnosis of cervical dysplasia. Early detection, facilitated by improved biomarkers or diagnostic methods, can greatly increase the chances of successful treatment and improve the prognosis of CC.
Treatment options for CC vary depending on the stage of the disease. Surgery, radiotherapy, chemotherapy, immunotherapy or combination therapy are commonly used and have a high cure rate for early-stage CC. However, the prognosis for advanced CC is generally unfovorable, with limited treatment effectiveness [6]. Consequently, early detection and precise treatment are crucial. Exploring CC biomarkers is essential for developing new diagnostic, prognostic, and therapeutic strategies. This field of research merits further exploration to improve outcomes for patients.
Searching the National Institutes of Health/National Center for Biotechnology Information (NIH/NCBI/PubMed) database by identified keywords. Keyword formula: ((((((((((((((((((biomarkers [MeSH Terms]) OR (Marker, Biological [Title/Abstract])) OR (Biological Markers [Title/Abstract])) OR (Biomarker [Title/Abstract])) OR (Immune Markers [Title/Abstract])) OR (Immunologic Markers [Title/Abstract])) OR (Immune Marker [Title/Abstract])) OR (Serum Markers [Title/Abstract])) OR (Surrogate End Point [Title/Abstract])) OR (Clinical Marker [Title/Abstract])) OR (Clinical Markers [Title/Abstract])) OR (Clinical Marker [Title/Abstract])) OR (Biochemical Marker [Title/Abstract])) OR (Biochemical Marker [Title/Abstract])) OR (Markers, Laboratory [Title/Abstract])) OR (Marker, Laboratory [Title/Abstract])) OR (Surrogate Markers [Title/Abstract])) OR (Surrogate Marker [Title/Abstract])) AND (((((((((((Uterine Cervical Neoplasms [MeSH Terms]) OR (Uterine Cervical Neoplasm [Title/Abstract])) OR (Cervical Neoplasms [Title/Abstract])) OR (Cervical Neoplasm [Title/Abstract])) OR (Cancer of the Uterine Cervix [Title/Abstract])) OR (Cancer of the Cervix [Title/Abstract])) OR (Cervical Cancer [Title/Abstract])) OR (Cervical Cancers [Title/Abstract])) OR (Uterine Cervical Cancer [Title/Abstract])) OR (Uterine Cervical Cancers [Title/Abstract])) OR (Cancer of Cervix [Title/Abstract])) OR (Cervix Cancer [Title/Abstract]). Timeframe for search: from 23 May 2019 to 23 May 2023. Both Chinese and English literature were accepted, but unpublished studies were not included in the search.
Inclusion criteria: (1) sample size
A total of 1759 abstracts were retrieved based on keywords. Two independent researchers (MC and YW) conducted a first screening based on titles and abstracts, resulting in a total of 550 articles. A second screening was then performed by reading the full text, which totaled 105 articles. Three independent reviewers (HM, LY and ZX) conducted a detailed evaluation of the inclusion and exclusion criteria, ultimately selecting 43 articles for discussion and summary. The screening process is illustrated in Fig. 1. Out of the total, 6 papers were related to diagnostic biomarkers, 36 papers with prognostic biomarkers, and 26 papers with therapeutic biomarkers.
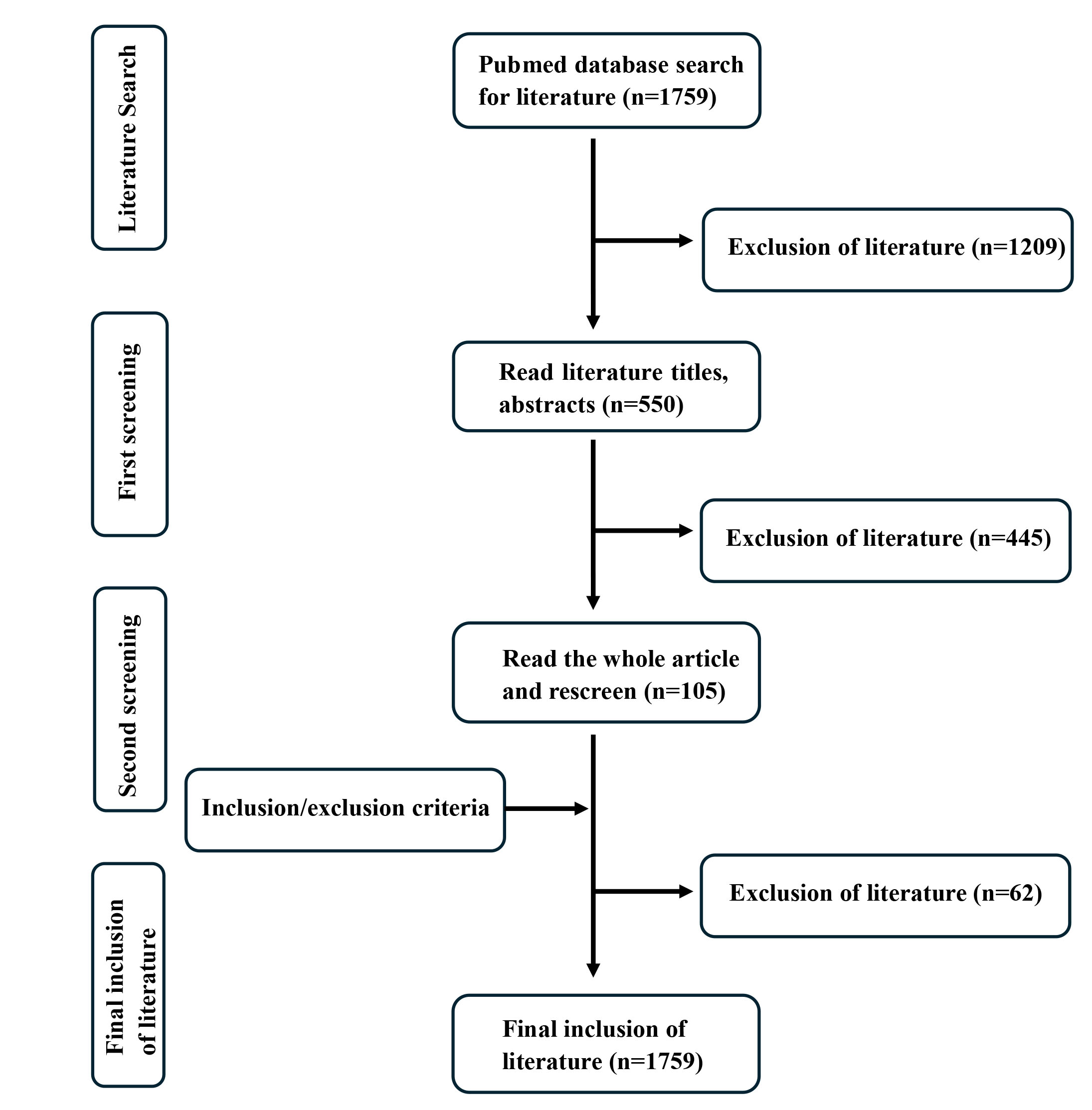 Fig. 1.
Fig. 1.Screening flowchart.
CC is a significant health concern, and the identification of biomarkers for its diagnosis has garnered considerable attention. In this context, Shao et al. [7] conducted a study in 2020 to explore the clinical significance of miR-210-3p in CC. The findings of the study revealed that miR-210-3p was remarkably up-regulated in CC tissues compared to normal and cervical intraepithelial neoplasia (CIN) tissues. This up-regulation was found to be correlated with significant clinical factors, including International Federation of Gynecology and Obstetrics (FIGO) stage, tumor differentiation, and lymphatic metastasis. These correlations suggest that miR-210-3p may be useful in the progression and metastasis of CC. Moreover, the study revealed that miR-210-3p exhibited promising diagnostic potential by effectively distinguishing CC tissues from CIN tissues. This indicates that miR-210-3p could serve as a valuable diagnostic marker for CC. Zheng et al. [8] found that exosomal miR-30d-5p and let-7d-3p were significantly different in plasma samples from CC compared to those from healthy volunteers. This study encompassed a total of 121 plasma samples, marking one of most extensive investigations of plasma miRNAs aimed at identifying cancer biomarkers. The exosomes miR-30d-5p and let-7d-3p discovered in this research are valuable diagnostic biomarkers for the non-invasive screening of CC and precancerous lesions. Yamanaka et al. 2021 [9] found that the relative expression level of miR-100 in serum was significantly lower in CC patients and lymph node-positive metastatic cases by analyzing RNA extracted from serum of 144 healthy volunteers, high-grade CIN, and CC patients. The study concluded that the expression of miR-100 was associated with the diagnosis of CC and was proportional to its favorable prognosis. Tumor suppressor genes, such as RARB, LMX1A, CADM1, DAPK1 and PAX1, play a crucial role in regulating cell growth and preventing tumor formation. Studies have shown that hypermethylation of these genes plays a role in DNA repair and apoptosis, regulation of the cell cycle, among other processes during tumorigenesis and progression [10, 11, 12]. DNA methylation testing is currently used for early screening of various cancers, including bladder and lung cancers [13, 14]. Currently, DNA methylation screening is not widely used in the early stages of CC. Further research is required to explore the potential of DNA methylation as a screening tool for CC and to establish its effectiveness in early detection.
Table 1 (Ref. [7, 8, 9]) provides a summary of studies focusing on biomarkers relevant to the diagnosis of CC.
| Study | Study design (P/R) | Sample size | Source of sample | Platform/assay technique | Application | Biomarkers analyzed | DOI |
| Shao et al. 2020 [7] | P | 435 | Tissue | qRT-PCR | Diagnosis | miRNA-210-3p [7] | 10.26355/eurrev_202006_21643 |
| Zheng et al. 2019 [8] | P | 121 | plasma | qRT-PCR | Diagnosis | miR-30d-5p and let-7d-3p [8] | 10.1186/s12943-019-0999-x |
| Yamanaka et al. 2021 [9] | P | 144 | Serum | qRT-PCR | Diagnosis | miR-100 [9] | 10.3233/CBM-201021 |
| Prognosis |
CC, cervical cancer; P, prospective study; R, retrospective study; qRT-PCR, quantitative real-time polymerase chain reaction; DOI, digital object identifier; miR, microRNA.
miRNAs play a crucial role in the pathogenesis of CC as major epigenetic regulators, controlling various essential processes such as growth, differentiation, angiogenesis, and development. Zhou et al. [15] demonstrated that exosome-coated miR-1468-5p, secreted by CC, has the ability to guide lymphatic endothelial cells to establish an integrated immunosuppressive tumor microenvironment (TME). Moreover, high serum exosome miR-1468-5p levels were associated with immunosuppressive status and poor prognosis of TME in CC patients. This study suggests that exosomal miR-1468-5p may be a prognostic biomarker and therapeutic target for CC. Yucel Polat et al. [16], found that miR-25 exhibited highly expressed in precancerous lesions and invasive carcinomas. Additionally, they observed that inhibiting miR-25 led to an up-regulation of Krüppel-like factor 4 (KLF4) expression, thereby regulating the proliferation and motility of CC cells. Tang et al., Wang et al., and Zhang et al. [17, 18, 19] concluded that circular RNAs (circRNAs) are closely associated with the diagnosis and prognosis of CC, and that circRNAs can regulate the expression of cellular genes, thereby affecting biological activities such as cell proliferation, invasion, and apoptosis.
Long non-coding RNA (lncRNA) is a non-coding RNA that is greater than 200 nucleotides in length [20]. An increasing body of research suggests that lncRNAs contribute to the regulation of cancer development and progression. Shen et al. and Zhou et al. [21, 22] found that lncRNAs are instrumental in the diagnosis, prognosis, and treatment of CC, and that they are closely linked to cancer progression, metastasis, drug resistance, HPV regulation, and metabolic reprogramming. Several targeted lncRNAs, including antisense oligonucleotide (ASO), Clustered Regularly Interspaced Short Palindromic Repeats- associated nuclease 9 (CRISPR/Cas9), and exosomes, have been utilized in clinical therapy. Furthermore, it has been shown that post-transcriptional RNA modifications play a crucial role in carcinogenesis, metastasis, progression, and drug resistance in various cancer tissues. Among them, N6-methyladenosine (m6A) is the most abundant type of RNA modification [23]. Jia et al. [24] investigated the effects of m6A-associated lncRNAs on the prognosis and treatment of CC. They discovered that m6A-associated lncRNAs can serve as key mediators of the tumor immune microenvironment. Thus, these lncRNAs could be potential therapeutic targets to improve the prognosis of CC and serve as indicators of CC prognosis. Condic et al. [25] showed that m6A modifications have an impact on the prognosis of CC, with elevated levels of m6A protein expression being associated with poor clinical prognosis. Additionally, Xu et al. [26] found that lncRNA HOXA cluster antisense RNA 3 (HOXA-AS3) was upregulated in CC cells and tissues. Knocking down HOXA-AS3 inhibited the progression of CC, contributting to better patient survival of. Bioassay analysis confirmed that HOXA-AS3 negatively regulates the development of CC by adsorbing miR-29a-3p. These findings suggest that lncRNA HOXA-AS3 may serve as a potential prognostic target for CC.
Wu et al. [27] found that the expression of signal transducer and activator of transcription-3 (STAT3) and phosphorylated STAT3 (p-STAT3) was significantly higher in CC tissues compared to normal tissues and CIN tissues. Furthermore, the overexpression of STAT3 in tissues was negatively correlated with the expression level of microtubule-associated protein 1 light chain 3B (LC3B). Reducing the level of STAT3 significantly increased the autophagy level both in vitro and in vivo, and inhibited the production of CC cells. Ye et al. [28] demonstrated that lactate dehydrogenase (LDH) can serve as a therapeutic and prognostic biomarker for CC, and that elevated levels of LDH expression were correlated with decreased overall survival (OS) in patients with CC. A study by Park et al. [29] found that the expression of telomeric zinc finger-associated protein (TZAP) may be used as a possible prognostic marker for early-stage CC. According to survival analysis, the prognostic value of TZAP expression was higher in stage N1 CC patients, and viral infection, cancer stage, and age were not related to TZAP expression. Furthermore, Zhao et al. [30] found that the expression of secreted phosphoprotein 1 (SPP1) in CC tissues was higher than that in normal cervical epithelial tissues, which was significantly associated with poor tumor prognosis, immune cell infiltration, and could be used as a prognostic biomarker for CC. Cao et al. [31] identified an association between the expression of colon cancer-associated transcript 2 (CCAT2) and FIGO stage, squamous cell carcinoma antigen (SCC-Ag), and lymph node metastasis (LNM) in CC. Moreover, CCAT2 expression in recurrent or metastatic CC was higher than that in primary CC, CIN, and was associated with poor OS. Therefore, it is believed that CCAT2 can be used as a biomarker for the diagnosis, staging, and prognosis of CC. Zhang et al. [32] found that the methylation level of tyrosine protein kinase ephrin type-A receptor A7 (EphA7) promoter was negatively correlated with the expression of EphA7 and was significantly higher in CC compared to that in normal tissues. Moreover, the methylation level and positivity rate of EphA7 were positively correlated with the severity of tumor progression, which indicated that the high methylation level of EphA7 could potentially serve as a biomarker for the diagnosis of CC. By analyzing 239 samples of CC patients, Li et al. [33] determined that the expression of high-mobility group box-1 protein (HMGB1) was significantly correlated with tumor size, paracervical infiltration, depth of cervical stromal infiltration, FIGO stage, and it can be used as a prognostic biomarker for CC.
Numerous studies have indicated that prognostic factors in CC are closely related to immunity. Scholars have extensively studied genes that have an impact on immune function, particularly in terms of the infiltration of immune cells. Cheng et al. [34] showed that triggering receptor expressed on myeloid cell 2 (TREM2) gene expression was negatively correlated with the level of immune cell infiltration, while it was positively correlated with the levels of M1 and M2 macrophage infiltration. TREM2 may influence the development and progression of CC through its effects on tumorigenesis and immunity. He et al. 2021 [35] discovered that expression of cell division cycle 45 (CDC45) protein correlates with immune-infiltrating cells. Their study, which involved analyzing tumor tissue samples screened from multiple databases, suggests that CDC45 may be useful for immunotherapy in CC. Furthermore, CDC45 may be involved in the development of CC as an independent prognostic factor. Pu et al. [36] explored the prognostic factors related to immunity in CC by analyzing 309 CC samples. Their study demonstrated that the pathway and function of CD79B were mainly correlated with immune activity, and its expression was positively correlated with a favorable prognosis. Furthermore, by exploring CD79B expression-associated immunomodulatory factors, it was confirmed that 10 genes (CD96, LAG3, PDCD1, TIGIT, CD27, KLRK1, LTA, PVR, TNFRSF13C, and TNFRSF17) could serve as promising independent prognostic factors for patients with CC, and could improve the clinical individualized treatment approaches for CC. In addition, Yang et al. [37] conducted a study in terms of the role of immune cells and tumor development. T cells and mast cells were identified among 22 types of immune cells significantly associated with the survival of CC patients, among which there were 9 genes were found to be linked to survival, including BTNL8, LY9, CCR7, GRAP2, CD1E, CD6, CD27, and CD79A. In addition, they found that C-C chemokine receptor type 5 (CCR5) and the chemokine (C-X-C motif) ligand 9 (CXCL9), CXCL10, CXCL11/recombinant chemokine C-X-C-Motif receptor 3 (CXCR3) axis could serve as novel targets for CC therapy. Wang et al. [38] found that the levels of aryl hydrocarbon receptor (AHR) and its related genes were associated with immune infiltration and immunoregulation in CC, and that high expression of AHR, cytochrome P450 1A1 (CYP1A1), heat shock protein HSP 90-alpha (HSP90AA1), and heat shock protein HSP 90-beta (HSP90AB1), and low expression of estrogen receptor 1 (ESR1) were negatively correlated with the prognosis of patients with CC. In addition, a study published by Tuo et al. [39] in 2022 demonstrated that the expression of runt-related transcription factor-1 (Runx1) was associated with patient prognosis and the level of immune infiltration of cancer-associated fibroblasts (CAFs) in a variety of tumors, which suggestes that Runx could serve as a potential prognostic biomarker for CC. Zhang et al. [40] demonstrated that the knockdown of transmembrane protein 33 (TMEM33) was positively associated with various immune-related cells, including dendritic cells (DCs), mast cells, plasma cells, and CD8+ T cells. Therefore, reducing the expression of TMEM33 can significantly inhibit the proliferation and invasion of CC cells, which can be used as a potential biomarker for assessing the prognosis and immune infiltration level of CC.
Zhang et al. [41] found that CircCDKN2B-AS1, a circular isoform of cyclin-dependent kinase inhibitor 2B antisense RNA 1 (CDKN2B-AS1), was highly expressed in CC and some precancerous tissues. This isoform can stabilize Hexokinase 2 (HK2) mRNA by sponging IMP3 (messenger RNA binding protein), thereby promoting the formation of malignant phenotypes of CC, and provide a new therapeutic target for the treatment of CC. Du et al. [42] found that methyltransferase-like 3 (METTL3) could be a potential prognostic biomarker and therapeutic target for CC. METTL3 is a core member of the m6A methyltransferase family, and studies have indicated that the expression of METTL3 promotes the proliferation and metastasis of CC cells. Additionally, METTL3 reduces the death of CC cells induced by endoplasmic reticulum stress by inhibiting the expression of thioredoxin domain containing 5 (TXNDC5). Shen et al. [43] found that recombinant ephrin A1 (EFNA1) was involved in multiple tumorigenic pathways, viruses, and proteins through the analysis of single-cell and RNA sequencing data, in which patients with low expression of EFNA1 had a better OS compared to those with high expression. Table 2 (Ref. [15, 16, 17, 18, 19, 21, 22, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43]) shows a summary of studies addressing biomarkers related to CC prognosis.
| Study | Study design (P/R) | Sample size | Source of sample | Platform/assay technique | Application | Biomarkers analized | DOI |
| Zhou et al. 2021 [15] | R | 169 | Serum | qRT-PCR | Prognosis | miR-1468-5p [15] | 10.1016/j.ymthe.2020.12.034 |
| Tissue | Treatment | ||||||
| Yucel Polat et al. 2021 [16] | P | 64 | Liquid-based cervical cytology samples | qRT-PCR | Prognosis | miR-25 [16] | 10.1016/j.prp.2021.153435 |
| Tang et al. 2020 [17] | P | 103 | Tissue | qRT-PCR | Prognosis | FoxO3a [17] | 10.2147/CMAR.S243329 |
| Wang et al. 2021 [18] | P | 60 | Tissue | qRT-PCR | Prognosis | circular RNA_0000326 [18] | 10.18632/aging.103711 |
| Zhang et al. 2021 [19] | P | NA | Tissue | qRT-PCR | Prognosis | circular RNA hsa_circ_0043280 [19] | 10.1038/s41419-021-04193-7 |
| Treatment | |||||||
| Shen et al. 2019 [21] | P | 50 | Tissue | qRT-PCR | Prognosis | lncRNA CCAT1 [21] | 10.1080/15384101.2019.1609829 |
| Western blot | Treatment | ||||||
| Zhou et al. 2019 [22] | P | 130 | Tissue | qRT-PCR | Prognosis | lncRNA-OIS1 [22] | 10.3892/ol.2019.9891 |
| Jia et al. 2022 [24] | R | 273 | Tissue | Bioinformation analysis | Prognosis | m6A-associated lncRNA [24] | 10.1155/2022/8700372 |
| Treatment | |||||||
| Condic et al. 2022 [25] | R | 118 | Tissue | Immunohistochemical | Prognosis | m6A RNA [25] | 10.3390/ijms23137165 |
| Treatment | |||||||
| Xu et al. 2022 [26] | P | 132 | Tissue | qRT-PCR | Prognosis | HOXA-AS3 [26] | 10.1111/jog.15360 |
| Wu et al. 2022 [27] | P | 46 | Tissue | Immunohistochemistry | Prognosis | STAT3 [27] | 10.1186/s13000-021-01182-4 |
| Treatment | |||||||
| Ye et al. 2022 [28] | P | 408 | Serum | NA | Prognosis | LDH [28] | 10.1089/gtmb.2021.0006 |
| Treatment | |||||||
| Park et al. 2020 [29] | P | 264 | Tissue | Survival analysis | Prognosis | TZAP [29] | 10.3390/medicina56050207 |
| Zhao et al. 2022 [30] | P | 100 | Tissue | TCGA, GEO, and Genotype-Tissue Expression databases | Prognosis | SPP1 [30] | 10.3389/fgene.2021.732822 |
| Cao et al. 2022 [31] | P | 180 | Serum | qRT-PCR | Diagnosis | CCAT2 [31] | 10.1186/s13048-022-00950-0 |
| Prognosis | |||||||
| Zhang et al. 2022 [32] | P | 57 | Tissue | qRT-PCR | Diagnosis | EphA7 [32] | 10.1186/s12885-022-09653-7 |
| MSP and QMSP | |||||||
| Li et al. 2019 [33] | P | 239 | Tissue | RNA isolation and qRT-PCR | Prognosis | HMGB1 [33] | 10.1042/BSR20181016 |
| Cheng et al. 2021 [34] | R | 11069 | Tissue | NA | Prognosis | TREM2 [34] | 10.3389/fimmu.2021.646523 |
| He et al. 2021 [35] | P | 211 | Tissue | PPI network | Prognosis | CDC45 [35] | 10.7717/peerj.12114 |
| Treatment | |||||||
| Pu et al. 2022 [36] | P | 309 | Tissue | qRT-PCR | Prognosis | CD79B [36] | 10.3389/fgene.2022.933798 |
| Yang et al. 2021 [37] | R | 307 | TCGA database | ESTIMATE algorithm | Prognosis | CCR5 and CXCL9, -10, -11/CXCR3 axis [37] | 10.1016/j.gene.2020.145119 |
| Treatment | |||||||
| Wang et al. 2021 [38] | R | 115 | Tissue | Survival analysis and Immunohistochemistry | Prognosis | AHR and its related genes [38] | 10.1080/21655979.2021.2006953 |
| Tuo et al. 2022 [39] | P | 529 | Tissue | NA | Prognosis | RUNX1 [39] | 10.1186/s12885-022-09632-y |
| Treatment | |||||||
| Zhang et al. 2023 [40] | R | 306 | Tissue | qRT-PCR | Prognosis | TMEM33 [40] | 10.1155/2023/5542181 |
| Zhang et al. 2020 [41] | P | 162 | Tissue | qRT-PCR | Prognosis | CircCDKN2B-AS1 [41] | 10.1186/s13046-020-01793-7 |
| Treatment | |||||||
| Du et al. 2022 [42] | P | 124 | Tissue | IF, dot blot assays, and m6A quantitative measurement | Prognosis | METTL3 [42] | 10.1038/s41388-022-02435-2 |
| Treatment | |||||||
| Shen et al. 2022 [43] | P | 306 | Tissue | Single-cell sequencing | Prognosis | EFNA1 [43] | 10.1007/s13577-022-00679-4 |
| Treatment |
P, prospective study; R, retrospective study; qRT-PCR, quantitative real-time polymerase chain reaction; NA, not available; TCGA, The Cancer Genome Atlas; GEO, Gene Expression Omnibus; MSP, methylation-specific PCR; QMSP, quantitative methylation-specific PCR; PPI, Protein-protein interations; IF, immunofluorescence; lncRNA, long non-coding RNA; DOI, digital object identifier; miR, microRNA; ESTIMATE, the Estimation of Stromal and Immune cells in Malignant Tumours using Expression data; CCAT1, colon cancer-associated transcript 1; FoxO3a, Forkhead box O3; lncRNA-OIS1, Long non-coding RNA-oncogene-induced senescence 1; m6A, N6-methyladenosine; HOXA-AS3, HOXA cluster antisense RNA 3; STAT3, signal transducer and activator of transcription-3; LDH, lactate dehydrogenase; TZAP, telomeric zinc finger-associated protein; SPP1, secreted phosphoprotein 1; CCAT2, colon cancer-associated transcript 2; EphA7, ephrin type-A receptor A7; HMGB1, high-mobility group box-1; TREM2, triggering receptor expressed on myeloid cell 2; CDC45, cell division cycle 45; CD79B, B-cell antigen receptor complex-associated protein beta chain; CCR5, C-C chemokine receptor type 5; CXC, chemokine (C-X-C motif); CXCR3, recombinant Chemokine C-X-C-Motif Receptor 3; AHR, aryl hydrocarbon receptor; RUNX1, runt-related transcription factor-1; TMEM33, transmembrane protein 33; METTL3, methyltransferase-like 3; EFNA1, ephrin A1.
Common treatments for CC include surgery, radiotherapy, chemotherapy, and immunotherapy [44]. Considering treatment benefits and complications helps assess the long-term impact of each option and provides a balanced view of associated risks. This information guides clinicians in making informed decisions and effectively managing potential complications.
The current trend in CC therapy is to avoid combined treatment involving radical surgery and chemoradiotherapy. Especially for disease stages T1b1, T1b2, T2a1, the treatment strategy should aim to avoid combining radical surgery and radiotherapy due to the high morbidity induced by the combined treatment [45]. Radiotherapy is a common method for treating CC, employing high-energy rays for either external or internal radiation therapy. Squamous CC is sensitive to radiotherapy. Therefore, radical radiotherapy is important for CC patients who are not suitable for surgical treatment. However, the side effects of radiotherapy, including fatigue, gastrointestinal discomfort, and skin reactions, can seriously affect the quality of a patients life. Intensity Modulated Radiation Therapy (IMRT) can reduce the irradiated dose to normal tissues, such as the bowel and bladder, effectively reducing the side effects of treatment. Chemotherapy helps to control the disease and reduce the risk of recurrence, and is an important treatment option for patients with advanced or recurrent CC. Chemotherapy can also result in side effects, including nausea, vomiting, hair loss, and immune system suppression, which may adversely affect a patient’s quality of life. Surgery, such as radical hysterectomy, stands as another treatment option effectively controlling early-stage CC with low recurrence rates. It is important to note that recent studies have shown that the prognosis of radical laparoscopic surgery is comparable to that of open surgery for patients with low-risk CC. Therefore, laparoscopic surgery is worth trying [46, 47]. While postoperative recovery may impact the patient’s quality of life, advancements in surgical techniques have significantly reduced its negative effects.
Additionally, recent research by Xu et al. [48] demonstrated that never
in NIMA (never in mitosis gene A)-related kinase 2 (NEK2) not only activates the
Wnt1/
In the treatment of CC, different approaches are utilized based on the stage and characteristics of the disease. Surgery and radiotherapy have proven effective for early-stage patients, whereas immunotherapy has emerged as a pivotal option for advanced and recurrent CC caused by HPV [55]. Immunotherapy functions by activating or enhancing the patient’s immune system, enabling it to recognize and attack cancer cells. Clinical studies have shown that immunotherapy can significantly prolong survival in advanced or recurrent CC caused by HPV. Additionally, it can be used alongside other treatments like chemotherapy or radiotherapy to improve overall outcomes [56]. However, it is important to note that not all patients respond favorably to immunotherapy.
Additionally, Wang et al. [57] demonstrated that the expression of oncogenes were associated with immunotherapy of tumors, specificaly the expression of the oncogene recombinant FK506 binding protein 10 (FKBP10) was shown to be negatively correlated with the degree of tumor differentiation and was shown to be positively correlated with tumor stage. On the other hande, the expression of the oncogene sphingosine 1-phosphate receptor 4 (S1PR4) showed a contrary association. Additionally, the low-risk group presented a more favorable immune activation phenotype and a higher enrichment of immunotherapy-related biomarkers. Shi et al. [58] demonstrated that the expression of glycolipid transfer protein (GLTP) was not only associated with CD4+ T cells, macrophages, and DCs, but also correlated with the expression of immune checkpoints, iron-mutagenesis-associated genes, and m6A-associated genes. Notably, these gens were markedly up-regulated in CC specimens. However, the Kaplan-Meier assay showed that CC patients with lower GLTP expression tended to have lower OS. Furthermore, Wang et al. [59] found that CD3-gamma (CD3G) is associated with increased expression of RNA and proteins within tumors, as well as with related immune pathways and HPV defense mechanisms, suggesting its potential as a biomarker for immunotherapy of CC. Xu et al. [60] demonstrated that RHO family interacting cell polarization regulator 2 (RIPOR2) is positively correlated with immunocompetent cells, such as T cells and immune checkpoint proteins (ICPs), and that the cumulative effect of these ICPs enhances immunity, ultimately influencing immunotherapeutic response of tumors. In addition, Li et al. [61] demonstrated that prostate transmembrane protein androgen induced 1 (PMEPA1) serves as a prognostically relevant biomarker for various cancers, particularly CC. Their finding suggests that PMEPA1 is involved in tumor immunity, with increased expression of PMEPA1 being positively correlated with high immune infiltration levels of multiple immune cells, which suggests that PMEPA1 could be a potential immunotherapy target for CC. Furthermore, CC is predominantely characterized by a local invasiveness. Hence, localized anti-PD-1/PD-L1 therapies are likely to be more effective and less toxic, and efforts should be made to identify precise biomarkers for PD1/PD-L1 blocking therapies in order to improve efficacy [62, 63]. The authors also mentioned that neoadjuvant immunotherapy triggers the detection and elimination of micrometastatic tumors, which may be the source of recurrence. Therefore, PD-L1 plays an important role in immunotherapy for CC. Interestingly enough, Zhang et al. [64] concluded that interleukin-33 (IL-33) could be synergistically combined with PD-L1-related antitumor therapies. This conclusion was drawn by analyzing tissue samples collected from 93 cases of CC, and they found that the expression of PD-L1 in tumor tissues was negatively correlated with prognosis, whereas the expression of IL-33 in tumor tissues was positively correlated with prognosis. In the treatment of tumors, while many tumor cells initially respond well to initial chemotherapy, resistance remains a major limitation to the efficacy of chemotherapy in numerous advanced cancers. Pang et al. [65] in 2022 showed that death-associated protein kinase 1 (DRAK1) inhibited the growth of paclitaxel-resistant CC cells both in vivo and in vitro. In CC tumor cells with paclitaxel resistance, cullin-3/speckle-type POZ (CUL3/SPOP) E3 ubiquitin ligase promotes tumor cell growth through degradation of DRAK1. Therefore, DRAK1 has an impact on the chemosensitivity of CC cells. To offer a comprehensive overview of biomarker studies related to CC treatment, Table 3 (Ref. [48, 49, 50, 52, 53, 54, 57, 58, 59, 60, 61, 64, 65]) provides a summary of these findings. By examining the various biomarkers associated with immunotherapy, researchers can gain insights into potential targets and mechanisms for improving treatment outcomes in CC.
| Study | Study design (P/R) | Sample size | Source of sample | Platform/assay technique | Application | Biomarkers analized | DOI |
| Xu et al. 2020 [48] | P | NA | Tissue | qRT-PCR | Treatment | NEK2 [48] | 10.1186/s13046-020-01659-y |
| Han et al. 2021 [49] | P | 692 | Tissue | Immunohistochemistry | Prognosis | TRPV1 [49] | 10.1097/PGP.0000000000000677 |
| Treatment | |||||||
| Nilsen et al. 2022 [50] | P | 200 | Tissue | Powerplex 16 | Diagnosis | miR-200a, miR-200b, and miR-429 [50] | 10.1002/1878-0261.13184 |
| Prognosis | |||||||
| Zhou et al. 2022 [52] | R | 19 | Tissue | Microarray and bioinformatic analyses, qRT-PCR, and western blotting | Prognosis | KLK5 [52] | 10.1016/j.ygyno.2022.05.010 |
| Treatment | |||||||
| Tian et al. 2020 [53] | P | 120 | Tissue | qRT-PCR | Treatment | MAPK4 [53] | 10.1186/s13046-020-01644-5 |
| Western blotting | |||||||
| Li et al. 2021 [54] | P | 55 | Tumor tissues and PBMCs | Immunohistochemistry | Diagnosis | PD-1/PD-L1 [54] | 10.1016/j.ijrobp.2021.03.003 |
| Treatment | |||||||
| Wang et al. 2022 [57] | P | 291 | Tissue | Immunohistochemistry | Prognosis | FKBP10, S1PR4 [57] | 10.3389/fimmu.2022.993118 |
| Treatment | |||||||
| Shi et al. 2022 [58] | R | 300 | Tissue | Survival analysis | Treatment | GLTP [58] | 10.1155/2022/9109365 |
| Wang et al. 2022 [59] | P | 293 | Tissue | Survival analysis and ESTIMATE | Diagnosis | CD3G [59] | 10.3389/fonc.2022.979226 |
| Prognosis | |||||||
| Treatment | |||||||
| Xu et al. 2022 [60] | P | 300 | RNA | High Pure RNA Paraffin Kit protocol - Roche | Prognosis | RIPOR2 [60] | 10.3389/fimmu.2022.930488 |
| Treatment | |||||||
| Li et al. 2022 [61] | P | 309 | RNA | Chip data of RNAs from TCGA datasets | Prognosis | PMEPA1 [61] | 10.1155/2022/4510462 |
| Treatment | |||||||
| Zhang et al. 2022 [64] | P | 93 | Tissue | Immunohistochemistry | Diagnosis | IL-33, PD-L1 [64] | 10.1002/JLB.5MA0322-746R |
| Prognosis | |||||||
| Treatment | |||||||
| Pang et al. 2022 [65] | P | NA | Tissue | qRT-PCR | Prognosis | DRAK1 [65] | 10.1038/s41419-022-04619-w |
| Treatment |
P, prospective study; R, retrospective study; qRT-PCR, quantitative real-time polymerase chain reaction; TCGA, The Cancer Genome Atlas; NA, not available; PBMC, peripheral blood mononuclear cells; DOI, digital object identifier; NEK2, NIMA (never in mitosis gene A)-related kinase 2; TRPV1, transient receptor potential vanilloid 1; KLK5, kallikrein 5; MAPK4, mitogen-activated protein kinase 4; PD-1, programmed cell death protein 1; PD-L1, programmed death ligand 1; FKBP10, recombinant FK506 binding protein 10; S1PR4, sphingosine 1-phosphate receptor 4; GLTP, glycolipid transfer protein; CD3G, CD3-gamma; RIPOR2, RHO family interacting cell polarization regulator 2; PMEPA1, prostate transmembrane protein androgen induced 1; IL-33, interleukin-33; DRAK1, death-associated protein kinase 1.
Currently, the incidence and mortality rates of CC have been decreasing due to the widespread adoption of the CC preventive vaccine and the large-scale implementation of cytology screening [66]. However, CC remains the fourth most common cancer among women worldwide, and more effective prevention and treatment strategies are still required [67]. At this stage, in order to better diagnose and treat CC, researcher continue to explore and update CC-related biomarkers.
Prognostic biomarkers play a crucial role in influencing various processes, such as tumor proliferation, invasion, infiltration, and apoptosis. By utilizing these biomarkers, patient survival can be effectively managed, and precision treatment can be guided. Moreover, therapeutic biomarkers are closely associated with tumor sensitivity and resistance to radiotherapy and chemotherapy. Identifying and targeting these biomarkers can significantly improve therapeutic sensitivity in CC treatment, resulting in improved outcomes and prognosis. Furthermore, the identification of novel biomarkers has expanded the scope of tumor screening. By combining these novel biomarkers with existing ones, the capability to predict tumor survival is enhanced, allowing for stratified diagnosis and treatment. This approach establishes a solid foundation for precision treatment, wherein individualized therapies can be tailored based on specific biomarker profiles.
Despite the potential of cancer biomarkers, their translation into clinical application has progressed relatively slow. The process requires an extensive foundation of clinical samples as the basis and involves repeated design and validation. Translational research has been suggested as a means to bridge the gap between the results of basic research on biomarker discovery and clinical practice [68]. However, strengthening the process of translational research for clinical application remains an area that requires further exploration.
HM, YW, MC contributed to research topic selection, article screening and analysis; HM, LY contributed to paper screening and analysis, manuscript writing and revision; ZX contributed to article screening and analysis, manuscript revision. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
Thanks to all the peer reviewers for their opinions and suggestions.
This work was supported by Shenzhen Science and Technology Program (No.JCYJ20210324114600002 and JCYJ20230807113009019); Shenzhen Key Medical Discipline Construction Fund (No.SZXK014); Shenzhen Key Laboratory for cancer metastasis and personalized therapy (No.ZDSYS20210623091811035) and Sanming Project of Medicine in Shenzhen (No.SZSM202211017).
The authors declare no conflict of interest.
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
