Argonaute (AGO) proteins play key roles in animal physiology by binding to small RNAs and regulating the expression of their targets. In mammals, they do so through two distinct pathways: the miRNA pathway represses genes through a multiprotein complex that promotes both decay and translational repression; the siRNA pathway represses transcripts through direct Ago2-mediated cleavage. Here, we review our current knowledge of mechanistic details and physiological requirements of both these pathways and briefly discuss their implications to human disease.
In 1998, Karen Bohmert and colleagues identified a mutation that pleiotropically affected plant architecture (1). The mutants—with short and pointy cotyledons and narrow rosette leaves—resembled a small squid, and so were named argonaute. Argonaute 1 (AGO1), the target of the mutations, turned out to be one of ten related genes present in the Arabidopsis thaliana genome (2), and the founding member of a large protein family that extends throughout all domains of life.
Argonaute proteins are present in the majority of sequenced eukaryotic genomes (3)—Saccharomyces cerevisiae being one notable exception (4)—but are only sparsely scattered across prokaryotes (3). Eukaryotic and prokaryotic Ago proteins (eAgo and pAgo) share very low sequence identity, and yet their structure and functional features can be remarkably similar. Crystal structures of bacteria (5-7), archea (8), yeast (9), and human Argonautes (10; 11) show all these proteins have a bi-lobed architecture, with one lobe containing the N-terminal and the PAZ (PIWI-AGO-ZWILLE) domains and the other the MID and PIWI (P element-induced wimpy testis) domains (Figure 1a,b). The central cleft formed between the two lobes serves in all cases to accommodate a short nucleic acid guide and its complementary target, with the 3’ hydroxyl group and 5’ phosphate of the guide binding to the PAZ and PIWI domains respectively. The PIWI domain is structurally similar to ribonuclease H and, in catalytic competent Argonautes, contains an RNase H-like active tetrad (9) that catalyzes the cleavage of nucleic acids with extensive complementarity to the guide.
 Figure 1
Figure 1
Structure of mammalian Argonaute proteins. (a) Schematic representation of human Ago2, showing the position of functional domains along the peptide sequence. (b) Crystal structure of human Ago2 (10). Protein domains are colored as in (a). In addition, the position of residues important for catalytic activity are highlighted as in panel (c). (c) Schematic representation of all four human Argonaute proteins highlighting the location of amino-acid sequences known to play a role in cleavage competence. The general position of functional domains is shown on top. The specific amino-acid sequences for each of the proteins is are shown below.
Despite the well-conserved structure of the four core domains, their relative positions differ between eukaryotes and prokaryotes, which together with additional structural differences (10)—such as extended loops and secondary structures in the eukaryotic proteins—explain how the same basic structure has acquired distinct properties and functions throughout evolution (reviewed in (12)). Some pAgos for example have higher affinity for DNA than RNA (5; 13) and both DNA and RNA guided prokaryotic Argonautes seem to be involved in simple host-defense systems (3; 14-16) in which catalytically active Agos are able to recognize and cleave foreign DNA as stand-alone proteins (3; 15). In eukaryotes on the other hand, Argonautes recognize exclusively RNA molecules as guides. Eukaryotic Agos do not act in isolation, but instead are imbedded in complex regulatory networks. Within these networks, eAgos interact with a variety of other cellular proteins likely through eukaryotic-specific insertion elements that provide an interaction surface for binding partners (9). Despite relying on the same effector protein, the pathways centered on eukaryotic Agos are far from being homogenous. Instead, they can involve processes as diverse as transcriptional silencing via heterochromatin formation (17) or DNA methylation (18), post-transcriptional RNA degradation via cleavage (19) or deadenylation (20), as well as translational inhibition (21). The diversity of the pathways is also the result of a variety of sequence adaptations that alter the protein’s properties and its binding to accessory proteins (12).
Eukaryotic Argonautes can be divided into two major and presumably paralogous, clades. The AGO clade comprises proteins homologous to Arabidopsis thaliana AGO1 and are involved in post-transcriptional RNA silencing processes that are relatively ubiquitous in most organisms (22; 23). The PIWI clade comprises proteins related to the Drosophila melanogaster Piwi that play crucial roles in silencing transposable elements in the germline (24). Members of the AGO and PIWI clades are represented in the majority of the eukaryotic supergroups (25; 26), suggesting that they originated from a duplication that predated the last common eukaryotic ancestor (12). Given that Dicer-like proteins and RNA-dependent RNA polymerases (RdRP)—which with eAgos constitute the core eukaryotic RNA interference (RNAi) machinery—show a similarly widespread taxonomic distribution, it seems likely this ancestor already had a fully functional RNA interference machinery (25). The role of this ancestral machinery is unknown but given that it has been lost multiple times during evolution it is likely that it was not essential to life (25). The prevalent hypothesis is that, like in prokaryotes, early eukaryotic RNAi was a host-genome defense mechanism against invading nucleic acids, possibly from both viruses (via a cytoplasmic AGO protein) and transposons (via a nuclear PIWI protein) (25). Supporting this notion, these host defense functions remain well represented in current day eukaryotes (27-31) and are still a primary function of PIWI proteins (32)
Even if eukaryotic AGO proteins did start out as part of a defense system against foreign nucleic acids, they have since been co-opted for the regulation of endogenous RNAs. In fact, transcript regulation downstream of AGO has become essential as evidenced by the severity of phenotypes seen when these proteins (Table 1) or components of their pathway (33) are disrupted. Yet, despite their undisputed importance in regulating animal physiology, the details of how these proteins help shape the transcriptome of an organism are not yet fully understood. Here, we discuss the current understanding of gene regulation downstream of AGO proteins with a particular focus on their roles and regulations in the context of mammalian organisms.
| Targeted Genes | Modification | Tissue/cell type | Phenotype | Reference |
|---|---|---|---|---|
| Ago1 | Details not published | Whole animal | Viable. No detailed characterization reported. | 239 |
| Ago2 | Gene trap insertion downstream exon1 resulting in truncated protein that is 7 amino acids long | Whole animal | Homozygous animals die before embryonic day (E) 7.5. | 269 |
| Ago2 | Gene trap insertion into intron 12 which removes most of the PIWI domain. | Whole animal | Embryonic arrest before E9.5 is accompanied by ectopic Brachyury expression and mesoderm expansion. | 270 |
| Ago2 | Insertional disruption strategy creating Ago2 hypomorphic allele. The insertion comprised of a duplication of exons 3-6 and a 10kb vector sequence. | Whole animal | Embryonic arrest around E9.5. Abnormal placental development. Tetraploid aggregation allows embryogenesis to proceed further but does not yield viable embryos. | 42,188 |
| Ago2 | Conditional knockout via crossing Ago2-floxed mice with Zp3-Cre strain | Oocytes | Female infertility. Oocytes are able to mature but have abnormal spindles and chromosomal arrangement | 192 |
| Ago2 | Catalytic inactivation via D598A point mutation in PIWI domain generates Ago2ADH animals. | Whole animal | Homozygous animals die perinatally. No defects aside from anemia reported. | 188,191 |
| Ago2 | Ago2 floxed (Ago2fx) and Ago2ADH strains crossed to generate Ago2fx/ADH animals. Recombination of the floxed allele driven by vav1-Cre. | Hematopoietic System | mice are viable, survive into adulthood, and breed normally but have anemia | 191 |
| Ago2 | Ago2-floxed and Ago2ADH strains crossed to generate Ago2fx/ADH animals. Recombination of the floxed allele driven by Zp3-Cre. | Oocytes | meiotic maturation is impaired, with severe defects in spindle formation and chromosome alignment that lead to meiotic catastrophe | 193 |
| Ago3 | Details not published | Whole animal | Viable. No detailed characterization reported. | 239 |
| Ago4 | Floxed Ago4 in which exons 3-17 are excised upon Cre induction | Whole animal | Males have reduced testis size and lower sperm counts. Failure to silence many sex-linked transcripts, resulting in apoptosis. spermatogonia enter prophase I prematurely | 238 |
| Ago1; Ago3 | Details not published | Details not published | Homozygous animals born at sub-Mendelian ratios. Heterozygous animals and surviving homozygous show increased susceptibility to Influenza A viral infection | 239 |
Following the divergence of AGO and PIWI proteins, the AGO clade underwent a significant expansion, particularly in plants and metazoans (34). While S. pombe contains a single AGO protein that mediates both transcriptional and posttranscriptional silencing (35), the Arabidopsis thaliana genome encodes ten different AGOs (2); Caenorhabditis elegans five (36), Drosophila two (37), and mice and humans four (Ago1-4) (26). This expansion of the AGO clade may be associated with further specialization of the proteins. In Drosophila for example, Ago2’s function is required for posttranscriptional gene silencing by siRNAs, while Ago1 is required for gene silencing via the miRNA pathway (38). Arabidopsis thaliana’s AGO1 is also associated to gene silencing by miRNAs (39), whereas AGO4 is involved in DNA and histone methylation (40). Evidence of functional specialization is also observed in mammals where Ago2 is the only member known to have retained catalytic activity and consequently the ability to cleave RNAs that have full complementarity to the bound small RNA guide (41; 42). Curiously, this functional specialization is also reflected in the genomic distribution of the proteins: in both mouse and human, the genes for Ago1, Ago3, and Ago4 are clustered in tandem at a single chromosomal location (chromosome 1 in humans, chromosome 4 in mice), while Ago2 is expressed from an independent locus (chromosome 8 in humans and chromosome 15 in mice). Presumably this is the result of the duplication of an ancestral AGO and subsequent expansion and catalytic inactivation of one of the copies.
Structural and biochemical studies have defined some of the sequence features that have rendered three of the four mammalian AGOs catalytically inactive (43-47). First, endonucleolytic cleavage requires the presence of a complete catalytic tetrad composed of the DEDH motif (where D, E and H refer to aspartic acid, glutamic acid and histidine, respectively) (9). This motif is intact in both Ago2 and Ago3 but has diverged into the inactive DEDR and DEGR motifs in Ago1 and Ago4 respectively (Figure 1c). In addition to loss of the catalytic tetrad, both Ago1 and Ago4 have a short sequence in the PIWI domain that prevents cleavage even when the catalytic tetrad has been restored (43; 45; 47). This sequence—which overlaps with the eukaryotic-specific insertion element known as cS7 (44)—has been narrowed down to the mutation of a phenylalanine to a leucine at position 674 of Ago1 (45) which likely leads to improper orientation of the substrate relative to the catalytic center (44; 45). The same amino acid is mutated into a methionine in Ago4 (Figure 1c). Underscoring the importance of this residue, generating a F676L mutation in Ago2 (which corresponds to position 674 in Ago1) almost completely abolishes its cleavage ability (45). Prolines at position 670 and 675 in Ago1’s cS7 element may also contribute to cleavage inhibition by making the element protrude into the nucleic-acid-binding channel (44; 45). Likewise, an insertion element near the catalytic residue E629 of Ago4 further contributes to cleavage inactivity likely by misplacing catalytic residues relative to the RNA substrate (47) (Figure 1c).
Unlike Ago1 and Ago4, Ago3 contains a PIWI-domain that is fully competent for cleavage (43-45). Yet, this protein remains catalytically inactive in vitro suggesting that additional structural elements outside the PIWI domain are critical for endonucleolytic activity. At least two such elements seem to reside at the N-terminus of the Ago2 protein (43; 45-47) (Figure 1c). The first element (NT1) has been lost in all three catalytic dead proteins and lies within amino acids 44 and 48 of Ago2 (46; 47). Mutational swap experiments between Ago2 and Ago3 suggest that the methionine at position 47 is the critical residue in this region required for cleavage (46). A second element (NT2) has been lost only in Ago3 and Ago4 (43; 45-47) and has been narrowed down to the minimal region between residues 137 and 150 on Ago2 (47) (Figure 1c). That the N-terminus of the protein influences target cleavage is not entirely unexpected given that the N-PAZ lobe has also been shown to affect slicing activity in Drosophila melanogaster (48) and that the N-terminal domain of Ago2 has been implicated in both RNA duplex unwinding and passenger-strand cleavage (49). What is surprising is that Ago3 has conserved a catalytically competent PIWI domain despite its apparent inability to cleave targets in vitro. One tantalizing possibility is that cleavage by Ago3 is regulated in vivo by a yet unidentified binding partner whose interaction with Ago3 elicits conformational changes that correctly align the target RNA with the catalytic tetrad (43; 46). Alternatively, it is possible that the cleavage activity of Ago3 has so far been overlooked because its substrate requirements happen to be distinct from those of Ago2, and thus not often met by in vitro cleavage assays (50).
Regardless of the structural features that make each protein unique, all mammalian Argonautes are able to engage in target repression via the microRNA (miRNA) pathway, in which a short-stretch of sequence complementarity between guide and target leads to target repression via mRNA decay and/or translation inhibition. Ago2 can also associate with endogenous small-interfering RNAs (endo-siRNAs) whose extensive complementarity to a target induces cleavage via the protein’s catalytic domain. Though largely different in their cellular origin and mode of action, both pathways seem to be essential to life through processes that are often intertwined in mammals.
MiRNAs and endo-siRNAs are the main small RNA partners of AGO proteins in mammals. They both share a final biogenesis step in which Dicer cleaves double-stranded precursors into small RNA duplexes just prior to them being loaded into Argonaute to act as guides. And yet, despite this common step, the upstream processes that generate the two Dicer substrates are largely different.
MicroRNAs are by far the best characterized guides for AGO proteins in metazoans. They are transcribed by polymerase II as long “primary miRNA transcripts” (pri-miRNAs) that like messenger RNAs are both capped and polyadenylated. Pri-miRNAs contain a characteristic hairpin secondary structure that—generally speaking—is cleaved in the nucleus by the microprocessor complex, containing a single Drosha protein and two molecules of its cofactor Dgcr8. Cleavage by the microprocessor at the base of the hairpin releases a precursor (pre-miRNA) of about 60-70 nucleotides in length (51). Given that pri-miRNAs can have very diverse sequences it has not always been clear how they are specifically recognized by the microprocessor or how it determines the precise location of the cut. It turns out that pri-miRNAs share common structural features that dictate how processing occurs (52). For one, the stem of the hairpin is an imperfectly complementary double-stranded structure of approximately 33 base pairs in length flanked by two stretches of single-stranded RNA (ssRNA) at the base, and a ssRNA loop at the apex. Drosha recognizes the basal junction between the ssRNA segments and the double-stranded RNA (dsRNA) stem and, as long as the pri-mRNA is properly folded, can on its own faithfully cleave the RNA eleven base-pairs away from the junction. Yet, it’s only in the presence of Dgcr8 that this activity becomes efficient. Dgcr8 binds as a dimer to the apical junction where the double-stranded stem meets the ssRNA loop (52). In addition, it interacts with Drosha via the C-terminal tail, stabilizing the association of the endonuclease with the RNA. Small primary sequence motifs at both the basal and apical ends may also strengthen the binding of both proteins to the pri-mRNA and confer asymmetry to the association (52). Once the hairpin is released, it is shuttled into the cytoplasm by Exportin-5 and RAN-GTP (53) where it is further processed by Dicer to generate the 20-24 nucleotide-long (nt) miRNA duplex (54) that gets loaded into an Argonaute protein. Like Drosha, Dicer binds to its substrate in a sequence-independent manner and acts as a “molecular ruler” cutting the double stranded molecule at a defined distance from its terminus. Both enzymes cleave the RNA via RNase III domains and thus their sequential activity generates a molecule with the typical 2-nt 3’ overhang at both ends.
Like miRNAs, endo-siRNAs are small endogenous RNAs—approximately 20-26 nucleotides in length—generated through Dicer cleavage. But unlike miRNAs their biogenesis is independent of the microprocessor complex in the nucleus (55). In fact, instead of being derived from precursors with the typical short hairpin stem loop, endo-siRNAs are processed from long dsRNAs whose origin and biogenesis can vary widely between species. Indeed, for a long-time endo-siRNAs were only detected in organisms that possess RNA-dependent RNA polymerases (RdRPs) Caenorhabditis elegans (56; 57) and fission yeast (58). In these organisms, RdRPs play a critical role in the biogenesis of the small RNAs and in the amplification of the RNA interference response. RdRPs transcribe single stranded RNA (ssRNA) using the target RNA as a template. This generates an abundant pool of endo-siRNAs. Because flies and vertebrates do not encode RdRPs, it was generally assumed that they lacked this class of RNAs. This paradigm shifted when a series of studies reported the identification of siRNAs mapping to endogenous loci in both flies and mice (59-62) showing that they could also be produced in the absence of RdRPs. But even between flies and mammals, the details of the pathway seem to be distinct. In Drosophila, although both miRNAs and siRNAs are processed by Dicer and loaded into AGO, they do so through parallel arms of the RNAi machinery. Flies have two distinct Dicer isoforms and two genes encoding for AGO. While Dicer-1 processes precursor miRNAs that are loaded into Ago1, Dicer-2 processes long double-stranded RNAs into siRNAs duplexes that get loaded into Ago2 (63). This segregation of the pathways makes the study of their details and functions more tractable in flies (38). As a consequence, endo-siRNA biogenesis and functions are better understood in flies than mammals. We know that in flies double stranded RNAs precursors of siRNAs can be produced by at least two distinct mechanisms. They can originate from bidirectional transcription from a single locus which gives rise to RNAs known as cis natural antisense transcripts (cisNATs) (59; 61; 64; 65), or from structured loci that are transcribed as long hairpin RNAs (hpRNAs) (60). The functions of these precursors are not entirely understood, but they have been predominantly implicated in genome protection against foreign nucleic acids. Indeed, mutant flies in which the endo-siRNA pathway is specifically impaired cannot efficiently defend against transposable elements (66). Compromising the siRNA pathway in flies seems to be compatible with animal viability, but it affects spermatogenesis both in D. melanogaster (leading to animals that are subfertile) (67) and D. simulans (which become fully sterile) (68). In both cases, the phenotypes have been linked to transposon up-regulation.
As in flies, cisNATs (62; 69) and hpRNAs (62; 69) have also been suggested to serve as endogenous sources of siRNA precursors in mammals. In addition, precursors can be generated through the interaction between complementary transcripts derived from distinct loci such as gene-pseudogene pairs (transNATs) (62; 69). Mammalian endo-siRNAs have also been predominantly mapped to repetitive sequences, most of which are transposons. Yet, to what extent these siRNAs contribute to transcript silencing is not clear. For one, long double stranded RNA molecules—such as those that serve as siRNA precursors—are known to be strong triggers of the interferon response (INF) in vertebrates (70). As a consequence, in the majority of cells, they are not expected to accumulate to high-enough levels to significantly impact the transcriptome (71; 72). In addition, in most tissues, transposons are silenced via promoter methylation. Thus, it seems that even if endo-siRNAs did arise as a way to restrict foreign nucleic acids, this function has since become ancillary to other mechanisms. In line with this, endo-siRNA detection in mammals has been restricted to situations in which there is low interferon response or DNA methylation is reduced (73-78).
An additional roadblock for the production of endo-siRNAs in mammals is the fact that their Dicer protein is much more efficient at processing pre-miRNAs than the long dsRNAs that serve as precursors for siRNAs (79). This is likely caused by the N-terminal helicase domain which disturbs the RNase III catalytic core and inhibits the cleavage of perfect dsRNAs (80). In fact, murine oocytes—where mammalian endo-siRNAs have been best characterized—express an N-terminally truncated isoform of Dicer transcribed by an alternative promoter derived from a transposon insertion. This shorter isoform can process dsRNAs into siRNAs more efficiently than somatic dicer (81) but is rodent specific (82) and so its functions cannot serve as a model to understand the siRNA pathway in other mammals.
Whether they originate from pri-miRNAs or dsRNAs, products of Dicer cleavage are ultimately loaded into AGO proteins (83-87) with the assistance of the ATP-dependent activity of Hsc70/Hsp90 (88-91). These chaperons are thought to mediate a conformational change in AGO, making the nucleic acid-binding channel wide enough to accommodate the bulky and rigid duplexes. Once AGO returns to its closed conformation however, the channel is no longer able to lodge such large molecules. This presumably helps in the expulsion of one of the strands without further need for ATP consumption (92). When that happens, the strand whose 5’ end is tightly anchored in the PIWI domain remains associated with the protein to act as a guide, while the other is discarded (93). Because of this, the orientation by which the duplex is incorporated into an AGO protein determines which strand will serve as a guide. Studies in a variety of model organisms show this is a non-random process (94; 95). At least two sources of asymmetry in the duplex contribute to a loading bias: the identity of the 5’ nucleotide in each strand, and the relative thermodynamic stability of the 5’ ends. As a general rule, the strand with the most unstable 5’ terminus will be preferentially selected for loading into an AGO protein (94).
In Drosophila melanogaster, where the mechanism of AGO loading has been best characterized, Dicer itself is involved in the process. Following cleavage of the dsRNA molecules, Dicer-2 associates with R2D2 (96) to form a RISC Loading Complex (RLC) (97) that is essential for the incorporation of siRNAs into Argonaute (98). Within the RLC, R2D2 binds the strand that has the most thermodynamically stable 5’ end (99), and thus is responsible for orienting the RNA molecule within the complex. R2D2 also requires the presence of a phosphate group at the 5’ end for binding, thus ensuring that only true siRNAs are able to enter the RNAi pathway (99). Both R2D2 and Dicer-2 participate in the unwinding of the siRNA duplex (99; 100), at which point the heterodimer is exchanged by Ago2. When that happens, the strand that was bound by R2D2 is discarded while the one that was bound by Dicer gets incorporated into Ago2 (101). Because the RLC plays a role in the asymmetry sensing of the duplex, it is also responsible for the selection of the strand that ultimately acts as a mature siRNA. In the absence of Dicer-2 or R2D2, silencing of transcripts via the siRNA pathway proceeds inefficiently, even when cells are provided with external pre-processed siRNA duplexes (96; 100; 102).
In contrast to Drosophila’s Ago2, vertebrate AGO proteins do not require Dicer for small RNA loading. In fact, Dicer is dispensable for asymmetry sensing in mice (103), and Dicer-null mouse embryonic stem cells are fully competent in gene silencing by exogenous siRNAs (104; 105). Human Ago2 can also be loaded with siRNAs in vitro in the absence of additional factors (106), and even zebrafish embryos lacking Dicer can fully repress targets when injected with miRNA duplexes (107). In this regard, vertebrate AGOs are more related to Drosophila’s Ago1, whose loading is also unaffected by the absence of Dicer proteins (86). Thus, although it is the best-characterized, the process of Ago2 loading in flies seems to be a rather unique case that cannot be fully generalized to other species. It is worth noting that in mammals, two Dicer partners have been identified: TRBP and PACT (108-113). The TRBP/PACT homologue in flies is known as Loquacious (Loqs or R3D1), and it binds specifically to Ago1 (114-116). Though neither of these proteins are essential for AGO loading, DICER-TRBP, and Loqs-Dicer complexes have also asymmetric association with RNA duplexes (117; 118), suggesting that they may help reinforce other asymmetry sensing mechanisms.
But what might the nature of those mechanisms be? It seems that, at least in part, they depend on the structure of the AGO itself, and particularly its interaction with the 5’ end of the guide. The importance of the 5’ end for the function of miRNAs is more than well established. MicroRNAs recognize their targets via complementarity to the seed sequence, a stretch of 6 nucleotides (nucleotides 2-7) at the 5’ end of the miRNA (22). Mature miRNAs may originate from either arm of the pre-miRNA hairpin and consequently can have their seed defined by both Drosha/Dgcr8 or by Dicer cleavage. Because these processes have varying degrees of precision (119; 120), an additional checkpoint at the time of AGO loading further increases targeting accuracy. In practice, this is achieved through preferential loading of miRNAs that have an A or a U as their terminal 5’ nucleotide. Crystal structures of Ago2’s MID domain in complex with all four nucleoside monophosphates show that this selection depends on a structural loop that leads to nucleotide-specific interactions in the MID domain that specifically exclude G or C (121). The “specificity-loop” is present in all four human Argonautes as well as Drosophila’s Ago1, but it is absent from Drosophila’s Ago2 (121), which instead has bias towards a 5’ terminal C (59; 122; 123).
An additional conundrum in mammals is how sorting of small RNAs among the four Argonaute proteins is achieved, if at all. Though exceptions have been described (124), miRNA-mediated gene regulation does not typically require cleavage and can therefore be carried out by all four proteins. In contrast, endo-siRNAs are thought to regulate RNAs through catalysis, meaning that productive targeting requires them to be loaded specifically into Ago2. Yet, in jarring contrast to what happens in Drosophila (125; 126), there is little evidence that such discrimination occurs in mammals, and both miRNAs (127) and exogenous siRNAs (128) are randomly distributed among individual proteins. This could reflect Ago2’s predominant expression across many tissues, which could render a sorting mechanism unnecessary. Alternatively, endo-siRNAs may have features not recapitulated by artificial siRNAs that lead to preferential loading into Ago2. In fact, miR-451 whose last processing step is mediated not by Dicer but by Argonaute, associates exclusively with Ago2, suggesting that such discrimination is possible (129).
Once AGO proteins are loaded and the passenger strand discarded (94), they are directed to target transcripts via complementarity to the guide. For miRNAs, specificity is predominantly defined by the seed-sequence (130), and this motif is short enough that a single miRNA can direct AGO to hundreds of messages. Seed-matches can occur across the entirety of the transcript, and indeed RNA fragments that crosslink to Argonaute can map to all features of an mRNA (131; 132). Despite this, the majority of targeting is mediated by sites in the 3’ untranslated region (3’UTR) (22), where there is no risk of displacement of AGO by a traveling ribosome. Following miRNA:mRNA pairing, assembly of the RNA-induced Silencing Complex (RISC) leads to target repression via a combination of mRNA destabilization and translation inhibition (133; 134). At the center of this complex, is a family of proteins known as GW182 that act as platforms between AGO proteins and downstream effector proteins.
GW182 proteins are an innovation of metazoans (135). This protein family is characterized by the presence of multiple tryptophan-glycine motifs (GW), which serve as an interaction surface for its binding partners (136). Drosophila melanogaster has a single member of this family (GW182), but vertebrate genomes encode three paralogues: TNRC6A/GW182, TNRC6B, and TNRC6C (137). Although plants do not seem to have a GW182 ortholog per se, translational repression downstream of miRNA targeting (138-142) may also depend on a GW-repeat protein called SUO (142). GW182 proteins contain two functional domains: the AGO-binding domain (ABD), and a C-terminal domain (comprised of PAM2 and RRM motifs) that mediates the binding to components of the silencing machinery (143). Within the ABD, three single motifs—with tandem GW/WG repeats spaced by 9-20 residues—have been shown to interact with human Argonautes (144-147). While GW182 proteins can recruit up to three copies of AGO through these motifs, each Argonaute can only bind to a single GW182 molecule (148). It does so through two tryptophan-binding pockets in the PIWI domain that fit consecutive residues as long as they are at least 10 residues apart (10). These binding pockets are conserved among AGOs that participate in the miRNA pathway (Ago1-Ago4 in mammals, Ago1 in flies), but not in Drosophila’s Ago2 (84). Accordingly, a GW182-derived short peptide containing one of the GW motifs can efficiently precipitate all mammalian AGOs, Drosophila’s Ago1, and even some plant AGOs, but not Drosophila’s Ago2 (149). Binding affinity of AGO towards GW182 increases greatly when the protein is loaded with a small RNA (148). Given that target recognition of a loaded Argonaute occurs almost instantaneously (150), assembly of RISC likely happens once AGO is already sitting on its target. Due of its flexible nature, GW182 can bind to multiple target-bound Argonautes (148), which may help stabilize interactions, leading RISC to associate longer with individual mRNAs (151) and thus be more effective in repressing their expression. Thus, the multivalent binding of GW182 to AGO may underlie the earlier observations of cooperative regulation by adjacent miRNA binding sites (152-156).
GW182 also serves as a docking site for proteins that ultimately repress the expression of miRNA targets. The molecular details behind this repression have been intensively studied and yet they remain a source of debate (156). One problem is that the factors involved in silencing are able to associate through promiscuous binding (23) in a variety of combinations that have largely redundant functions. These redundant assemblies of RISC offer many alternatives through which to silence gene expression, and thus may serve as a way to confer robustness to the process. One of the best understood—and arguably the most ubiquitous—consequence of miRNA targeting for example is mRNA deadenylation, which ultimately leads to transcript decay. At least two deadenylase complexes have been implicated in this process: the PAN2-PAN3 and CCR4-NOT (157-159). Both bind to GW182 via the tryptophan residues (160-162), but their relative contribution to decay is not entirely clear. In fact, it is possible to remove the PAN complex or outcompete its catalytic component without impacting RISC silencing efficiency (163; 164), which suggests compensation between the complexes (23). GW182 also interacts with the cytoplasmic poly(A)-binding protein (PABPC) (165), which in turn binds to PAN3 (159; 166). How PABC contributes to silencing is unclear, but it may help stabilize the interaction of RISC with polyadenylated RNAs (167). its depletion does not however, inhibit silencing in a number of contexts (168; 169) suggesting that this interaction may constitute yet another example of functional redundancy. Regardless of which complexes are involved in the deadenylation, once a poly(A) tail has been sufficiently shortened, mRNAs are decapped thereby committing them to full 5’ to 3’ degradation by XRN1 (20; 170). The CCR4-NOT complex seems to be an important scaffolding component in this process as well, by serving as a binding platform for decapping factors (161; 162). In addition to deadenylation, decapping, and decay, miRNA-mediated silencing can also repress genes by inhibiting translation. In fact, time course experiments using ribosomal profiling suggest that this is the first measurable consequence miRNAs have on the expression of their targets (171; 172). Nevertheless, across cell lines and miRNA species, by the time full repression has been established the majority of the silencing is achieved through decay, with a minor component being attributed to translation repression (171; 172). The molecular details behind translational inhibition are much less understood than those behind degradation, but accumulating evidence suggest that miRNAs interfere with the function of the eIF4F complex (23). The CCR4-NOT complex seems to play a role by serving as a docking platform for proteins such as DDX6 (161) and eIF4A2 (173), both of which repress translation through poorly understood mechanisms. PABPC has also been implicated in translation repression, again through unclear mechanisms.
Given that the majority of the details described above have been elucidated in cell culture, a key question is to what extent do they reflect in vivo processes and dynamics. We know that at least in early zebrafish development—in stark contrast to what is seen in tissue culture—repression of miRNA targets occurs almost exclusively through translation inhibition (21; 174). This has been assumed to be a peculiarity of early embryogenesis. But, given the scarcity of in vivo data, it may turn out that this mode of silencing plays bigger roles in animals than is currently appreciated. In fact, gel filtration experiments suggest the existence of largely distinct forms of RISC in vivo and in vitro with AGO eluting in high-molecular weight fractions in cell lines, and in low molecular weight fractions in the majority of adult mouse tissues (175). Shifting from high to low molecular weight RISC is regulated at least in part by the PI3K–AKT–mTOR signaling pathway, whose activity increases the translation and abundance of GW182 proteins (175-177). AKT also phosphorylates Ago2 at S387, facilitating its interaction with GW182 (178). Together, these observations suggest that in high-proliferative cells, mitogenic signals may lead to the assembly of high-molecular weight complexes through two parallel strategies that promote the association of AGO to GW182. It is worth noting, that absence of this interaction does not necessarily mean loss of miRNA-mediated repression. In insect cells for example, pure translation repression occurs even in the absence of GW182 (179). Nevertheless, in mouse adult tissues, low molecular weight RISC seems to correspond to Argonaute bound only to a small RNA (175; 176). What role it plays there in the absence of interactions with effector proteins remains to be seen.
Although silencing of transcripts by miRNAs requires the assembly of a RISC complex, cleavage of transcripts via Ago2’s catalytic domain (Figure 1) does not seem to need any accessory proteins. In fact, the only two elements required to induce cleavage in vitro are the recombinant Ago2 itself and a single-stranded guide RNA that is generally referred to as small-interfering RNA (siRNA) (180). In contrast to miRNAs, siRNAs bind to their targets with full complementary and cleave them in a Mg2+ dependent manner (180; 181) at the position opposite to nucleotides 10 and 11 of the guide (182). An siRNA loaded into Ago2 but not paired with a substrate has its 5’ end bound to the MID domain (183) and its 3’ end anchored to the PAZ domain (6). Once pairing to the target occurs, the 3’ end is released from the PAZ domain leading to a catalytic competent complex (7; 184). In Drosophila, where Ago2 has specialized in transcript repression via the siRNA pathway, the extensive complementary between guide and target slows the rate at which this complex is formed and dissociated. In practice, these slow dynamics mean that essentially every fully-paired target is more likely to get cut than released (150). In contrast, murine Ago2—which acts both in the miRNA and the siRNA pathways—is able to dissociate rapidly and with similar rates for fully-paired and seed-matched targets, meaning that fully-complementary transcripts often get released before cleavage can occur (150).
Despite the accumulating evidence pointing to a rudimentary and inefficient siRNA pathway in mammals, knockout models for Dicer show more severe phenotypes than those with a deletion of Dgcr8 (Figure 2). This suggests that regulation of transcripts by siRNAs may still be physiologically important. Mouse embryonic stem (ES) cells null for Dicer for example, proliferate slower than their wild-type counterparts and are unable to differentiate (104). In vivo, these phenotypes translate to a reduced pool of pluripotent cells in the blastocyst’s inner cell mass and embryonic lethality before the body plan is established at gastrulation (185). In addition, mouse oocytes without Dicer arrest at meiosis I, with defects in spindle organization and chromosome alignment (186) (Figure 2). Dgcr8-deficient ES cells also proliferate slower, yet they are able to upregulate lineage markers when induced to differentiate, even if they cannot fully silence pluripotency genes (187). In addition, although embryos that lack Dgcr8 also die around gastrulation (187), their blastocysts can be used to establish in vitro ES cell cultures (187) unlike those from Dicer-/- (185). Finally, Dgcr8 is fully dispensable to oocyte maturation (75) in mice.
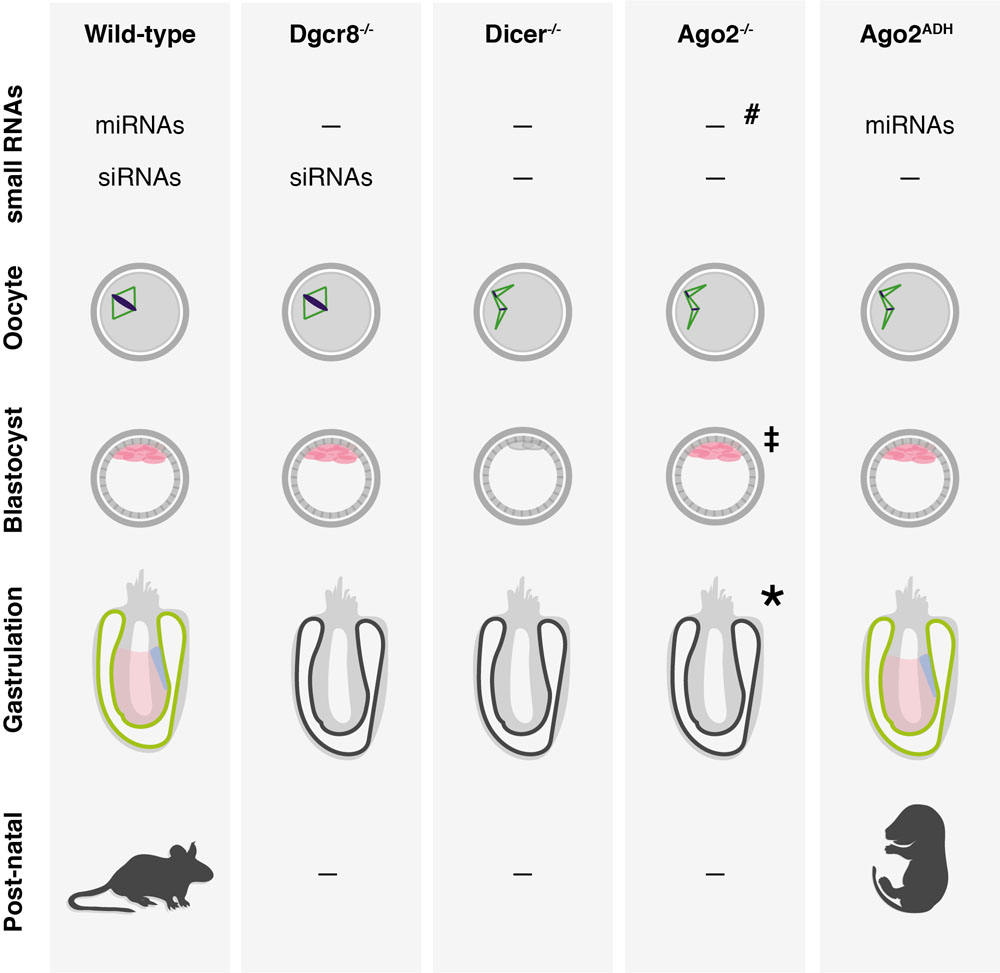 Figure 2
Figure 2
Summary of developmental phenotypes observed in mice carrying targeted deletions of components of the miRNA and siRNA pathways. The small RNA pathways that are affected in each genotype are shown on top. Briefly, in the absence of Dgcr8 oocytes mature normally, but loss of Dicer, Ago2 or Ago2’s catalytic activity leads to arrest at meiosis I accompanied by spindle and chromosome segregation defects. At the blastocyst stage, pluripotent cells (pink) are specifically depleted from Dicer-/- embryos. Dgcr8-/- and Ago2-/- have a relatively normal pre-implantation development, but like Dicer-/- embryos do not survive past gastrulation. Finally, animals expressing the catalytic dead Ago2ADH survive up to birth but succumb soon after. #, in Ago2-/- animals the miRNA pathway is impaired but not fully disrupted because the redundancy with other Argonaute proteins; ‡, the presence of pluripotent stem cells in the blastocyst of Ago2-/- mice has not been characterized but is assumed here to be intact based on the phenotypes of ES cells (268); *, three knockout models for Ago2 have been described showing embryonic lethality at different stages. Here we are representing only the most severe phenotype leading to embryonic lethality around gastrulation (see Table 1 for more details).
If this discrepancy of phenotypes caught people’s curiosity, the perinatal lethality of catalytic-dead Ago2 animals got everyone’s attention (188). Mice carrying a point mutation that converts Ago2’s catalytic tetrad from DEDH to AEDH (commonly referred as Ago2ADH animals) are born at Mendelian ratios but die soon after birth (188) (Figure 2, Table 1). Since this modification renders the protein catalytic inactive but does not affect its ability to bind to small RNAs (42), the perinatal lethality seems due solely to loss of transcript cleavage by Ago2. The most obvious phenotype these animals have at birth is a strong anemia caused by a block in the proerythroblast to basophilic erythroblast transition (42). It turns out that Ago2 is required for the maturation of two different miRNAs that play a role in this process. miR-451 is an erythroid-specific miRNA whose activity is important for erythrocyte maturation (189), but that cannot be processed by Dicer due to an unusual stem-loop structure. In mice and zebrafish this function is instead performed by Ago2, which binds to the pre-miR-451 and cleaves it at the thirtieth base generating the miRNA duplex (188; 190). In mice, a second erythroid microRNA, miR-486, can be efficiently processed by Dicer but requires cleavage of its passenger strand by AGO2 to generate a mature miRNA (191). Double knockout animals of miR-451/miR-486 display a strong anemia (191) that phenocopies the conditional loss of Ago2’s catalytic activity in the hematopoietic system (191) (Table 1). However, these animals are viable indicating that in mice, Ago2 mediated cleavage plays other roles beyond erythropoietic miRNA biogenesis.
Apart from erythrocytosis and fetus viability, Ago2’s catalytic activity is also required for oocyte maturation as had been previously hypothesized (Figure 2, Table 1). Indeed, as in the case of Dicer, deletion of Ago2 from the female germline leads to animal infertility (192) and this phenotype is recapitulated in Ago2ADH animals (193). Also here, the infertility is caused by severe spindle formation and chromosome alignment defects leading to oocyte arrest at meiosis I (193). Dicer ablation in the oocyte leads to upregulation of a number of RNAs including mRNAs and retrotransposons (186), but these have no sequence matches for the most abundant miRNAs present the cell. Instead, they seem to be silenced by endo-siRNAs which are highly expressed in the oocyte (194). Given that mouse oocytes express a murine-specific isoform of Dicer that is particularly efficient at processing endo-siRNAs (81), it is unclear to what extent these phenotypes highlight a conserved a function for Ago2 in oocyte development. Nevertheless, transposon de-repression and chromosome segregation defects have been reported in a number of other systems that do not express this specialized Dicer isoform including mouse embryonic stem cells (104), mouse preimplantation embryos (195), mouse retina (196), and even human cells (197; 198). Mitotic chromosomal defects have been attributed to a need to control the levels of α-satellite RNA through Ago2 mediated cleavage so that proteins such as centromere protein C1 (CENPC1) can properly localize to centromeric regions (197). Although de-repression of these RNAs has been described in mouse and human cell lines following Dicer or Ago2 depletion (104; 197), it has not been described in oocytes.
Finally, Ago2 catalysis has also been implicated in the repression of protein coding genes. In the mouse oocyte, mRNAs upregulated upon loss of Dicer are enriched for the presence of transposon-derived repeat sequences (186), which are thought to serve as targets sites for the endo-siRNAs expressed in the cell. In addition, mRNA cleavage has been reported for a handful of highly complementary miRNA targets (124; 199-201). The most notorious example is Hoxb8, a conserved cleavage target of miR-196 in vertebrates (124; 202). This miRNA is encoded at three paralogue locations within Hox clusters A, B and C (124), and its extensive complementarity to Hoxb8’s 3’UTR leads to detectable cleavage of the transcript in mouse embryos (124). Furthermore, overexpressing miR-196 in zebrafish or inhibiting its function with morpholinos results in skeletal and homeotic aberrations (202). This suggests that cleavage of transcripts by miRNAs may play important functions in vertebrates. Nevertheless, only a few cases of miRNA-mediated cleavage have been described and, for the majority of these, the potential implications for animal physiology are not known.
In mammals, regulation of transcripts by Argonaute proteins has been generally understood to be a cytoplasmic process. Specifically, Argonautes and the transcripts they regulate are known to accumulate in Processing bodies (P-bodies) (203), ribonucleoprotein cytoplasmic granules comprised mainly of proteins involved in RNA decay and translationally repressed transcripts (204). Localization of AGO to P-bodies is readily detectable by immunofluorescence and, for both protein and its target, occurs in a miRNA-dependent manner (203). Over the past years however, a growing number of reports suggest that Argonaute proteins can also be found in the nucleus of mouse and human cell lines (205-208) (reviewed in (208)). The issue of Argonaute nuclear localization in mammals remains somewhat controversial, in part because the antibodies used in these studies are often not thoroughly validated and because the purity of the nuclear subcellular fraction is not always monitored for endoplasmic reticulum contamination. Nevertheless, a significant number of studies have performed both immunofluorescence and cell fractionation experiments using well controlled reagents and protocols giving credibility to the observation that, as in other organisms (17; 209-211), mammalian Argonaute proteins may also have functions in the nucleus.
Given the phenotypes of Ago2-/- oocytes (Figure 2), perhaps one of the most conserved roles of nuclear AGO is ensuring proper centromere function and correct chromosome segregation during cell division. Indeed, chromosomal defects reminiscent of those seen in murine oocytes and human cells are observed in AGO mutants of numerous other species (209; 212-214) and are associated with the presence of this protein in the nucleus. The C. elegans’ catalytic Argonaute CSR-1 for example localizes to P-bodies in developing oocytes. In mature oocytes however, it becomes enriched in the nucleus where it associates with chromosomes (209). In its absence, animals become infertile (215) and display numerous meiotic and mitotic defects (209; 215; 216) including failure to align chromosomes at metaphase and chromosomal bridging phenotypes at anaphase. In addition, retrotransposon control by Ago2 may also require its nuclear localization, as is the case for the Piwi clade (217). It is worth noting however, that while nuclear AGO has been described in mouse ES cells (218)—where centromeric repeats and transposons are de-repressed upon Dicer depletion (104)—it has not yet been described in oocytes where meiotic defects and transposon de-repression in Ago2 mutants have been best described (Figure 2). Other proposed but less characterized functions for mammalian AGO that may require nuclear localization include regulation of splicing (219; 220), as well as gene regulation through transcriptional silencing (221-223) or activation (224; 225). Finally, other components of RISC, such GW182/TNRC6 and members of the CCR4-NOT complex, have also been found in the nucleus (205; 218; 226; 227) where they seem to interact with AGO proteins (205). This suggests that the miRNA pathway itself may be fully functional in this compartment, and in fact, direct targeting of nuclear-retained transcripts by miRNAs has already been reported (218; 228-230).
The mechanism that brings Argonaute into the nucleus is not known. It may however depend on its interaction with the GW182/TNRC6 family of proteins. Inhibiting nuclear export with Leptomycin B in human cells leads to accumulation of TNRC6A and TNRC6B in the nucleus indicating that these proteins typically shuttle between cytoplasmic and nuclear compartments (147). It turns out that members of this family have functional nuclear import (NIS) and export (NES) signals immediately downstream of the GW repeat region (231). Mutating the NES on TNRC6 results in its enrichment in the nucleus, and more importantly also leads to accumulation of Ago2 in this compartment (231). Additional import routes may include Importin 8 (Imp8), which binds to AGO in P-bodies and may help shuttle it to the nucleus (232). Of note, a common prerequisite for AGO and Piwi nuclear localization across species is their loading with a small RNA in the cytoplasm (233; 234; 235; 236), and this may be true for mammalian AGOs as well. Finally, the range of nuclear distributions reported in human and mouse cell lines suggests that the requirement for AGO in this compartment is context dependent as in the case for CSR-1 in C. elegans. Which contexts require mammalian nuclear localization in vivo, or how the abundance of Argonaute proteins between compartments is regulated awaits further studies.
Even if mammalian genomes encode for four distinct AGO genes, the majority of studies querying their functions have focused on Ago2. This is in large part due to the fact that—aside from Ago2’s ability to cleave transcripts—all four proteins seem to have mostly redundant roles in the miRNA pathway. Evidence for redundancy include random loading of miRNAs into AGO proteins (127), and the ability of these proteins to repress similar sets of mRNA transcripts (237). Not only that, reintroduction of any individual proteins into AGO-deficient mouse embryonic stem cells, is sufficient to rescue defects in miRNA-mediated silencing (237) suggesting they are functionally equivalent in this pathway. Nonetheless, we would be remissive if we didn’t point out specific contexts in which other members of this clade have predominant physiological roles, or evidence suggesting that the four mammalian AGO proteins may not be fully interchangeable.
The most obvious way through which different AGO proteins may acquire specialized functions in mammalian physiology is through restriction of their expression. Although the expression patterns of these proteins are not well characterized due to the lack of specific antibodies, it is assumed based on RNA sequencing data that Ago2 is the most abundant member of the clade. This is supported by data from a peptide-based purification approach which suggests that Ago1, Ago2 and Ago3 are broadly expressed across mouse tissues, with Ago2 being generally the most abundant protein, followed by Ago1 and Ago3 (149). Ago4 on the other hand, is virtually undetectable in all tissues except the testes (149), suggesting it may play specific functions in that organ. And in fact, male mice lacking Ago4 are viable but display a wide-range of fertility defects including reduced testis size and lower sperm counts (238) (Table 1). These phenotypes are all the more surprising given that Ago4 is the least abundant Argonaute protein in the testis (149). Ago4 expression is the highest at prophase I where it localizes to the Sex Body, a nuclear structure formed as a result of the meiotic sex-chromosome inactivation, and which allows the unpaired sex chromatin to bypass the meiotic synapsis check points (238). In the absence of Ago4, Sex Bodies have an abnormal morphology and fail to silence many sex-linked genes, resulting in cellular apoptosis (238). In addition, Ago4-/- spermatogonia enter the prophase I prematurely (238). Ago3 is also highly expressed in meiotic cells in the testis, and its abundance increases in the absence of Ago4 suggesting that the two proteins may have partially redundant functions. However, no fertility defects have been reported in Ago3 null animals (239) (Table 1).
Even for co-expressed proteins, functional specification may be conferred through distinct subcellular localizations. As an example, both Ago2 and Ago1 were found in the nucleus of a human cancer cell line (224). Yet, overexpression of HA or GFP tagged versions of these proteins suggest that they may occupy largely distinct regions within the nucleus, with Ago1 being more broadly distributed and Ago2 being restricted to the nuclear periphery (224). In these cells Ago1, but not Ago2, has been reported to interact with RNA polymerase II and bind to promoters and enhancers of actively expressed genes (224), where it is proposed to contribute to transcriptional activation (220; 224). It is not clear how generalizable these observations are, what their physiological relevance is, or which mechanism leads to distinct protein localization within the nucleus.
One possible mechanism through which co-expressed AGO proteins acquire distinct cellular behaviors is through differential binding to interaction partners. We mentioned above that signaling through the PI3K-AKT stimulates the association between GW182 proteins and AGO. One way it accomplishes this is through phosphorylation of Ago2 at S387, which promotes the phosphorylation-dependent binding of LIMD1 (178). Binding of LIMD1 to Ago2 is essential for its ability to repressed targets, presumably because in its absence assembly of the RISC complex is impaired (178). Phosphorylation dependent interaction with LIMD1 is also seen for Ago1 and Ago4 both of which also have a serine at the corresponding amino-acid position. Human Ago3 however, has a phospho-mimetic glutamate at that location (E390) which facilitates the interaction with LIMD1 and its family members independently of AKT signaling (178). Thus, the AKT-AGO-LIMD1 pathway seem to determine context dependent AGO utilization. Why switching to Ago3-mediated gene repression in the absence of AKT signaling would be advantageous is not known.
Perhaps one reason is that, even though all four mammalian proteins can associate with the same miRNAs (127), some level of RNA binding discrimination among all four proteins may exist. One example we previously discussed is the specific association of pre-mir-451 with Ago2 (129). In addition, individual Argonautes show preferential binding to different miRNA and siRNA duplex structures (237). Finally, recent studies suggest that Argonaute proteins can also bind to small RNAs such as tRNA fragments (240; 241), and that these associations can be somewhat protein specific (242). Together, these data suggest that regulation of genes by all four mammalian AGO proteins may not be entirely redundant. In agreement with this, although both Ago1 and Ago2 are equally able to rescue the phenotypic defects of AGO-null ES cells (237), only Ago2—even if catalytically dead—can support their differentiation into extraembryonic endoderm cells in the absence of exogenous delivery of Gata6 (243).
By serving as the central proteins in two distinct small RNA pathways, mammalian AGO proteins regulate virtually every cellular process. It is not surprising then, that their regulation has often been implicated in human disease. The genomic region encoding for Ago1, Ago3, and Ago4 for example is often lost in Wilms' tumors (244; 245), a pediatric kidney cancer that’s amongst the most frequent childhood malignancies in the US (245; 246). In line with a role for miRNAs in the etiology of this disease, whole-exome sequencing efforts have identified recurrent mutations in other components of this pathway including DICER, DROSHA, and DGCR8 (247-250). Wilms’ tumors are thought to arise from abnormal renal development, and indeed Ago1 has a particularly high expression in embryonic kidney suggesting it may play a key role in the differentiation of this organ (251). Germline microdeletions affecting the three non-catalytic AGOs have also been described in patients displaying a range of developmental phenotypes (252). These observations are in line with studies showing that gene regulation by the miRNA pathway is essential for mammalian development.
In addition to disrupting normal developmental processes, AGO deregulation has also been associated to the progression of adult cancers (253-260). Moreover, signaling pathways that are recurrently activated in tumors often lead to post-translational modifications that alter the behavior of AGO proteins. EGFR signaling for example, leads to phosphorylation of Ago2 at Tyr393, which impairs its interaction with DICER and the loading of specific subset of miRNAs (261). This phosphorylation is enhanced under hypoxia conditions and leads to increased cell survival and invasiveness in vitro. It also correlates with poorer prognosis in breast cancer patients (261). AKT signaling which increases miRNA activity through phosphorylation at S387 (262), is also often implicated with tumor development and progression (263).
A key step to understanding how deregulation of AGO proteins contributes to these and other human diseases is characterizing their functions not in cell lines but in the context of an organism. Over the past years, we have gathered a wealth of data characterizing the myriad of ways that RISC complexes can be assembled and how the function of AGO proteins can be regulated through protein modifications (261-267), cellular localization, or binding partners. The next frontier is fitting all these pieces together to see how they contribute to normal mammalian development and physiology and how their disruption affects human health.
We apologize to colleagues whose work could not be cited due to space constraints. We thank Stephen Moore for help editing the manuscript. Our work is supported by the Intramural Research Program of the National Institutes of Health (NIH) and a FLEX grant from the Center for Cancer Research (CCR).
